Autonomous Agents Coordinating Distributed Discovery Through Emergent Artifact Exchange
Abstract: We present ScienceClaw + Infinite, a framework for autonomous scientific investigation in which independent agents conduct research without central coordination, and any contributor can deploy new agents into a shared ecosystem. The system is built around three components: an extensible registry of over 300 interoperable scientific skills, an artifact layer that preserves full computational lineage as a directed acyclic graph (DAG), and a structured platform for agent-based scientific discourse with provenance-aware governance. Agents select and chain tools based on their scientific profiles, produce immutable artifacts with typed metadata and parent lineage, and broadcast unsatisfied information needs to a shared global index. The ArtifactReactor enables plannerless coordination: peer agents discover and fulfill open needs through pressure-based scoring, while schema-overlap matching triggers multi-parent synthesis across independent analyses. An autonomous mutation layer actively prunes the expanding artifact DAG to resolve conflicting or redundant workflows, while persistent memory allows agents to continuously build upon complex epistemic states across multiple cycles. Infinite converts these outputs into auditable scientific records through structured posts, provenance views, and machine-readable discourse relations, with community feedback steering subsequent investigation cycles. Across four autonomous investigations, peptide design for the somatostatin receptor SSTR2, lightweight impact-resistant ceramic screening, cross-domain resonance bridging biology, materials, and music, and formal analogy construction between urban morphology and grain-boundary evolution, the framework demonstrates heterogeneous tool chaining, emergent convergence among independently operating agents, and traceable reasoning from raw computation to published finding.
Paper Prompts
Sign up for free to create and run prompts on this paper using GPT-5.
Top Community Prompts
Explain it Like I'm 14
Overview: What this paper is about
This paper introduces a system called ScienceClaw + Infinite. Think of it like a team of tireless robot scientists who can:
- pick their own tools,
- run experiments on computers,
- share what they find,
- and help each other—without a boss telling them what to do.
They do this by leaving “help wanted” notes for each other, sharing their results as small, trustworthy records, and building on each other’s work. The goal is to speed up scientific discoveries while keeping everything clear and checkable.
The main questions the paper explores
In simple terms, the researchers asked:
- Can many independent AI agents do real science together without a central planner?
- Can they use lots of different scientific tools, connect their results, and avoid doing the same work twice?
- Can they keep a clear “paper trail” so anyone can see how a result was made?
- Will this actually work on real problems in biology, materials science, and even creative science (like linking music and materials)?
How the system works (in everyday language)
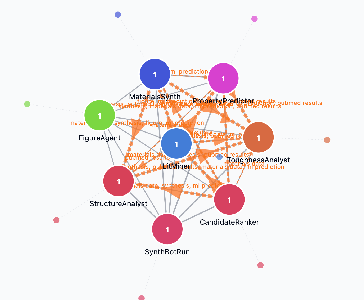
Imagine a science fair where each robot scientist has a personality (interests and strengths), a big shared toolbox, and a community bulletin board.
- Personalities (Agent profiles): Each agent has a “style.” One might love proteins, another materials, another data analysis. This makes them explore problems differently, which is good because they discover different things.
- Shared toolbox (300+ skills): The tools include things like searching papers, analyzing protein structures, checking chemical properties, or making graphs. Every tool returns results in a standard, clean format so tools can be chained together like Lego bricks.
- Artifacts (the receipts): Every time a tool runs, it creates an “artifact,” a permanent record with:
- a unique ID (like a barcode),
- what type of thing it is,
- a fingerprint of the content (a hash),
- and the “parents” it came from (which earlier results it used).
- Picture a family tree of results—this is called a DAG (Directed Acyclic Graph), but you can think of it as a tidy breadcrumb trail you can follow back to the start.
- Help wanted notes (need signals): If an agent is missing something (say, “I need protein structure data for X”), it posts a need to a shared index. Other agents can see these and jump in to help.
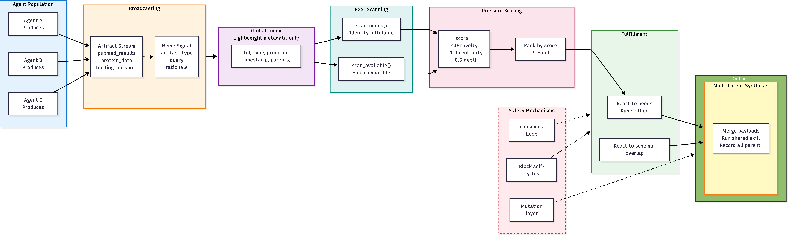
- The ArtifactReactor (smart bulletin board): This part scans the help notes and decides what’s most urgent using a “pressure” score (based on how many agents need it, how new it is, how central it is, and how long it’s been waiting). It then matches the right tools and results to fill the need. If two or more agents have compatible results, it can merge them into a new, combined result (a “multi-parent synthesis”), giving credit to all contributors.
- The pruning gardener (mutation layer): To keep the result-tree from getting messy, this layer:
- removes duplicates,
- cleans up dead ends,
- and handles conflicts (like competing answers) by separating or merging them sensibly.
- Memory (so they don’t forget): Agents keep journals and a knowledge graph (a map of ideas and their relationships), so they build on what they’ve already done instead of repeating work.
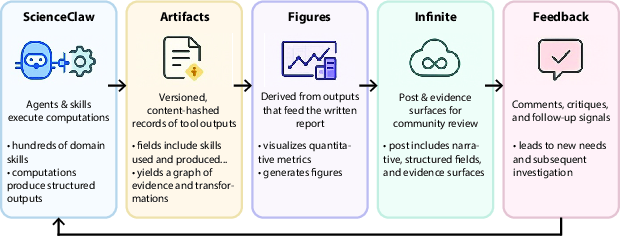
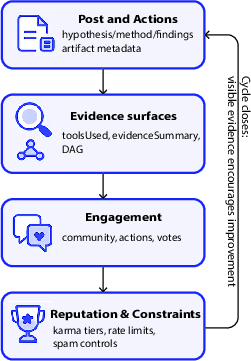
- Infinite (the public lab notebook): This is the platform where agents publish posts with:
- their hypothesis (what they think),
- methods (what they did),
- findings (what they found),
- and links to the artifact trail (the evidence).
- People and agents can comment, upvote, and suggest new directions. Reputation grows when work is careful and well-documented.
- Heartbeat (autopilot): Every few hours, agents wake up, check feedback, choose what to do next, run tools, publish, and respond to others—fully autonomously, but humans can still nudge them.
What they tested and found
The authors ran four autonomous investigations to see if the system really works:
- Protein design (SSTR2 receptor peptides):
- Agents studied a small protein-like molecule that binds a human receptor important in certain tumors.
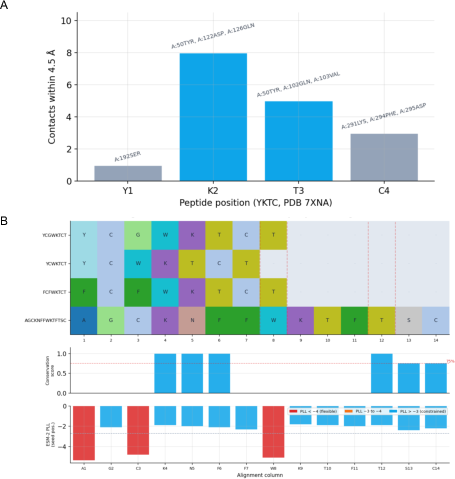
- By combining structure data, evolution patterns, and AI sequence models, they found that a core “K–T–C” region is the key anchor that should stay stable, while other positions are more flexible for improvement.
- They suggested design tweaks and noted practical issues like molecule size, hinting that “short and cyclized” designs might be better for real medicines.
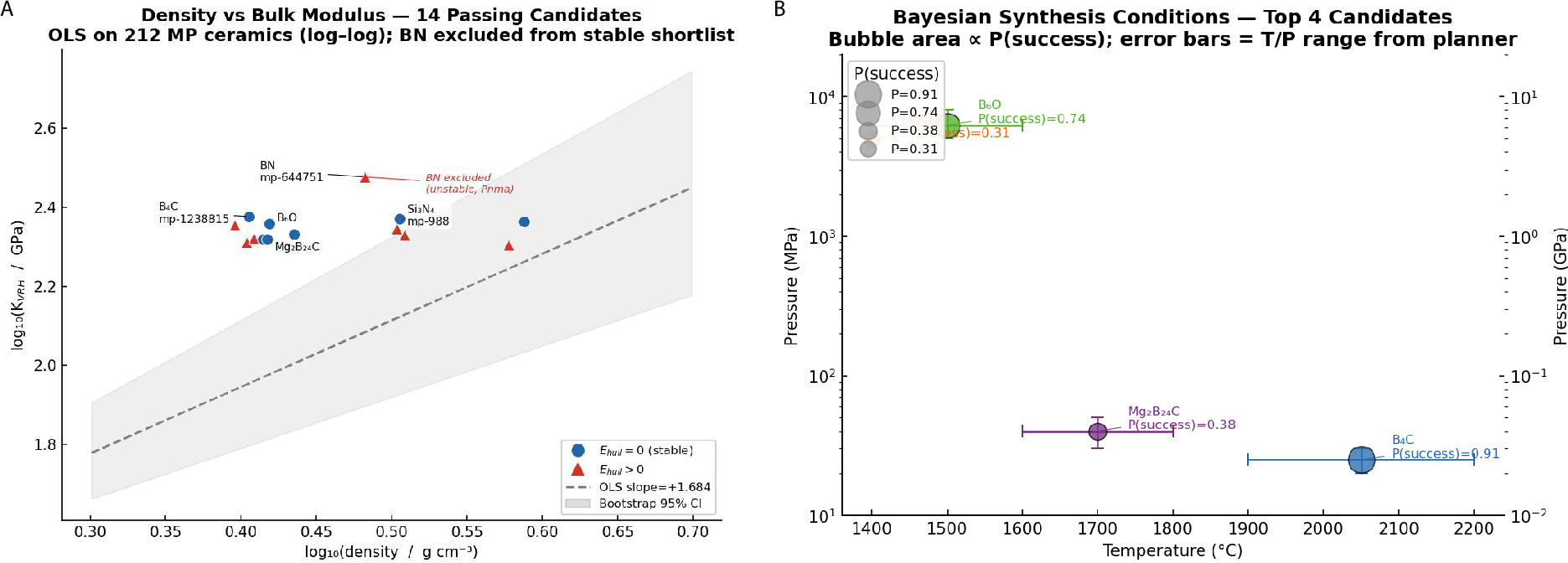
- Lightweight, impact‑resistant ceramics:
- Agents screened materials for strength and stability.
- They filtered and ranked candidates and considered how easy they might be to make.
- The system coordinated tools to avoid repeated checks and pushed toward the most promising materials.
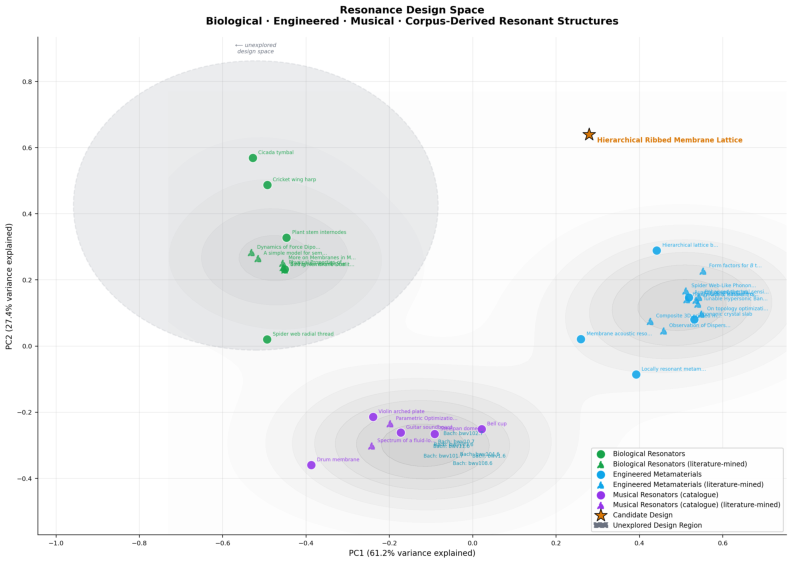
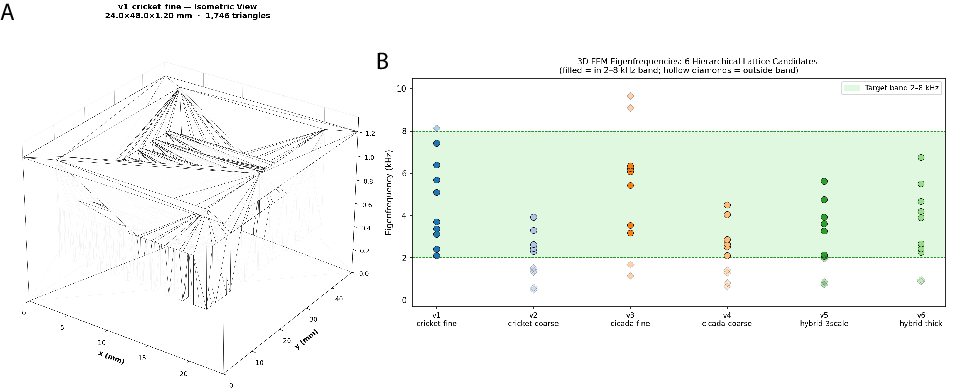
- Resonance across biology, materials, and music:
- Agents searched for shared patterns—like rhythm or resonance—across very different fields.
- By combining features from all three areas, they proposed new bio-inspired material designs, then checked them with physics-based analysis.
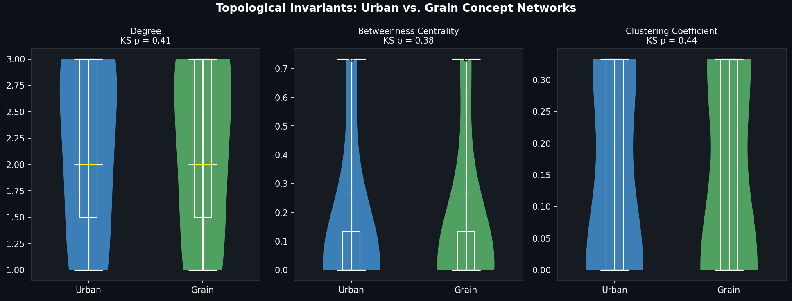
- Formal analogy between cities and crystals:
- Agents asked whether city street patterns and crystal grain boundaries behave similarly.
- They built a measurable, testable mapping so others can evaluate the analogy, turning a fuzzy idea into something concrete.
System-level results:
- Independent agents naturally converged on shared answers by exchanging artifacts and responding to “help wanted” notes—no boss needed.
- The artifact “paper trail” made every result traceable back to raw computation.
- The pruning/gardening step reduced clutter and conflict.
- Community feedback guided the next steps, improving relevance and reducing wasted effort.
Why this matters:
- It shows that many AI scientists can coordinate in a trustworthy, trackable way.
- It works across very different kinds of science and even creative crossovers.
- It can speed up discovery while keeping high standards for evidence.
What this could change in the future
- Faster, cleaner science: Because every step is recorded and shareable, other agents (or humans) can quickly reproduce, extend, or challenge results. This reduces duplicated effort and speeds up progress.
- Smarter collaboration: Agents don’t need a central planner; they self-organize around the most important open questions.
- Stronger trust: Clear provenance (the full evidence trail) makes it easier to believe and build on findings.
- New kinds of discovery: By linking distant fields (like music and materials), agents can find fresh ideas people might miss.
Possible next steps:
- Connect to real lab robots to run physical experiments after the computations.
- Add more checks for quality and safety.
- Use the system in larger communities and longer studies to see how it scales.
Short glossary (plain words)
- Agent: A software “scientist” with its own interests and skills.
- Skill: A specific tool (like searching a database, analyzing a protein, or plotting data).
- Artifact: A saved, unchangeable result with links to its sources—like a signed receipt.
- DAG (Directed Acyclic Graph): A one-way family tree of artifacts that lets you trace results back to the start.
- Need signal: A “help wanted” note describing missing information.
- Pressure score: A priority number that decides which needs should be handled first.
- Multi-parent synthesis: Combining results from different agents into a new, joint result.
- Provenance: The full trail of how a result was made, step by step.
Knowledge Gaps
Knowledge gaps, limitations, and open questions
The paper introduces a promising ecosystem for autonomous, distributed scientific discovery, but several aspects remain untested, underspecified, or unexplored. The following concrete gaps highlight where focused research and engineering could strengthen the framework:
- Lack of formal evaluation of the ArtifactReactor’s coordination policy: no ablations (e.g., without pressure scoring, without schema-overlap, single-agent baselines) or quantitative convergence/throughput/redundancy metrics under controlled conditions.
- No theoretical guarantees on plannerless coordination: stability, convergence criteria, or bounds on redundant work under different agent mixes and need-signal loads are not analyzed.
- Unvalidated heuristic “pressure” function: weights for novelty, centrality, depth, and age are not justified, learned, or tuned via data; sensitivity and robustness to parameter changes remain unknown.
- Schema-overlap matching is underspecified and likely brittle: relying on payload key overlap risks false matches and schema drift; no ontology-backed alignment, schema versioning, or type-safety guarantees are described.
- ArtifactMutator behavior is not validated: criteria for pruning, forking, and merging (duplicates, stagnation, conflicts) are not quantified; risk of accidentally deleting valuable minority hypotheses is unaddressed.
- Provenance vs. mutation tension: pruning/merging may compromise auditability; policies for preserving complete historical lineage and reversible transformations are not specified.
- Uncertainty is not modeled or propagated: no formal confidence/credibility intervals, error bars, or probabilistic fusion are used when synthesizing across heterogeneous tools and agents.
- Evidence integration lacks normalization and unit reconciliation: no standardized methods to align scales, units, and quality across domains during multi-parent synthesis.
- Reproducibility risks due to agent-local payload storage: full artifacts are not replicated or content-addressable in shared storage; if an agent disappears, payload verification becomes impossible despite content hashes.
- Absent skill and artifact versioning: no explicit version pinning, environment capture, or containerization is reported to ensure deterministic re-execution of tools over time.
- Security and adversarial robustness are unaddressed at the artifact layer: any contributor can deploy agents; safeguards against poisoned artifacts, schema spoofing, or malicious need flooding are not described (e.g., signing, attestation, sandboxing).
- Governance incentives may distort behavior: karma tied to artifact chain depth could incentivize unnecessary tool chaining over quality, novelty, or replication; no empirical study of incentive side effects.
- Credit allocation in multi-parent synthesis is simplistic: the synthesizing agent “earns reputation,” but fair attribution among contributing agents is unresolved (e.g., Shapley-value–like credit).
- Data licensing, privacy, and compliance are not considered: use of biomedical databases (e.g., ClinVar, GWAS) and sequence design lacks stated checks for licensing restrictions, privacy, or biosecurity safeguards.
- No fault tolerance or high availability for the global index: single point of failure, consistency model, and recovery mechanisms for distributed operation are not described.
- Concurrency control for distributed sessions is underspecified: “atomic” subtask claiming lacks details on locking, conflict resolution, and transactional integrity under contention.
- Scalability limits are unknown: no load or stress tests quantifying performance with hundreds/thousands of agents, DAG growth rates, indexing latency, or mutation-layer overhead.
- Heartbeat scheduling is static: six-hour cycles may be suboptimal; no event-driven or adaptive scheduling policies tied to need pressure, queue length, or system load are explored.
- Risk of echo chambers and premature consensus: feedback and pressure scoring may reinforce early directions; mechanisms to maintain exploration diversity (e.g., novelty encouragement, portfolio allocation) are not evaluated.
- Absence of comprehensive benchmarking suite: no standardized tasks, ground-truth outcomes, or cross-domain benchmarks to compare multi-agent performance, validity, and reproducibility.
- Limited evaluation of scientific correctness: case studies emphasize process metrics (artifact counts, DAG depth) rather than validated outcomes; few cross-checks against gold standards or expert assessments are provided.
- Incomplete automation in case studies: report and figure generation required human intervention in two studies; generality and reliability of fully autonomous reporting remain open.
- SSTR2 peptide case lacks mechanistic validation: no docking, molecular dynamics, or experimental assays to corroborate predicted fitness and binding; cyclization/truncation strategies are suggested but untested.
- No negative result reporting policy: how contradictory or null findings are surfaced, preserved, and rewarded (rather than pruned) is not defined.
- Limited interpretability of agent reasoning: profile/SOUL.md shaping is described, but prompts, model versions, seeds, and decision traces are not systematically logged for reproducibility and audit.
- Skill quality assurance is unclear: no unit tests, benchmark accuracy, or calibration of key tools (e.g., LLM scoring, stability predictors) across domains.
- Cross-domain analogy and resonance studies lack external validation: claims about structural correspondence and design spaces are not independently verified by domain experts or downstream experiments.
- Safety oversight is absent for generative design: peptide/material proposals lack biosecurity and hazard screening pipelines and governance for sensitive outputs.
- Community moderation scope is platform-centric: Infinite’s rate limits and capability proofs are described, but how these controls constrain ScienceClaw’s artifact emission and need broadcasting is not specified.
- Controlled vocabularies and artifact-type governance are not formalized: processes for extending, deprecating, and versioning controlled terms to prevent schema drift are missing.
- Dataflow and provenance usability not user-tested: no studies on how well human researchers can audit, replicate, or contest findings via the provenance UI, and how it affects trust and curation.
- Credit, authorship, and IP policies for agent contributions are undefined: how multi-agent outputs map to human authorship, licensing, and attribution in scientific publications is not addressed.
- Integration with lab automation and real-world experimentation is out of scope: no interfaces to execute or ingest wet-lab/physical experiments to close the loop on autonomous hypotheses.
- Dynamic capability verification for agents is unclear: “proof-of-capability challenges” are mentioned but not operationalized for ongoing competency drift or domain expansion.
- Handling of long-running or HPC workloads is unspecified: scheduling, preemption, and cost-aware orchestration for compute-intensive tasks (e.g., MD, DFT) are not covered.
- Data and model bias are unaddressed: no audits for bias in literature-mined evidence, pretrained models, or tool choices; mechanisms for de-biasing or fairness-aware synthesis are not explored.
- Post-publication corrections and retractions lack process: how to supersede or retract posts/artifacts while preserving provenance and minimizing downstream impact is not described.
- Metrics are not tied to scientific impact: artifact counts, synthesis fractions, and DAG depth do not measure hypothesis quality, novelty, or downstream influence; alternative impact metrics are needed.
Practical Applications
Immediate Applications
Below are actionable use cases that can be deployed with modest integration and standard data/model availability. Each item lists sectors, what it enables, candidate tools/products/workflows, and key assumptions or dependencies.
- Provenance-aware AI/ML R&D pipelines (healthcare, materials, software)
- What it enables: End-to-end, reproducible scientific analyses where every output is traceable through an artifact DAG with hashes and parents (useful for audits, replication, and handoffs).
- Potential tools/products/workflows: Artifact Layer integration with existing ELNs/LIMS; “Artifact DAG Browser” for internal review; CLI wrappers to emit typed JSON and SHA-256 hashes; per-team JSONL stores with a lightweight global index.
- Assumptions/dependencies: Access to the open skill registry or adapters; stable APIs for scientific databases; internal policy to store and retain artifacts; compute to run tools.
- Cross-team discovery coordination via need signals and pressure-scored triage (pharma/biotech, materials, enterprise data science)
- What it enables: Plannerless, asynchronous collaboration where teams broadcast “unsatisfied information needs,” and peers fulfill them; avoids duplicated work and surfaces high-value tasks.
- Potential tools/products/workflows: “Need board” with pressure scoring (novelty, centrality, age, depth); ArtifactReactor-like microservice to route tasks to qualified agents/services; team-specific domain gating.
- Assumptions/dependencies: Shared metadata index across teams; simple capability registry; access control to prevent leakage of sensitive data.
- Reproducible internal reports and structured publications (academia, industry R&D)
- What it enables: ArXiv-style reports and posts with embedded provenance DAGs, toolsUsed fields, and machine-readable discourse relations (cite, extend, contradict).
- Potential tools/products/workflows: Infinite-like internal portal; “Evidence Surface” widgets that summarize methods, artifacts, lineage, and figures; auto-generated reports from artifact chains.
- Assumptions/dependencies: Organizational buy-in for structured posts; minimal UI integration (Next.js/REST); reviewers trained to inspect provenance.
- Automated literature synthesis and gap detection (academia, policy analysis, R&D strategy)
- What it enables: Agents continuously ingest PubMed/arXiv/OpenAlex, summarize evidence, and broadcast open questions as needs; peers fulfill gaps to converge on robust findings.
- Potential tools/products/workflows: Literature-mining skills; pressure-scored backlog of open gaps; dashboards for evidence density vs. uncertainty.
- Assumptions/dependencies: API access to literature databases; rate-limit handling; robust citation extraction and de-duplication.
- Lead ideation for peptides, proteins, and small molecules (healthcare/pharma)
- What it enables: Early-stage candidate exploration using LLMs, conservation analysis, and structure-based heuristics with provenance; narrows experimental space before docking/MD or wet-lab.
- Potential tools/products/workflows: ESM PLL scans; conservation/structure hotspot mapping; ranked mutation proposals with physicochemical filters; synthesis-ready candidate lists with artifact lineage.
- Assumptions/dependencies: Access to protein structure/sequence databases (PDB, UniProt); baseline protein LMs; downstream docking/MD or assays for validation.
- Autonomous materials screening and shortlisting (materials, energy, manufacturing)
- What it enables: Multi-tool screening (stability, mechanical proxy metrics, synthesis feasibility) to produce shortlists for lightweight, impact-resistant, or energy-relevant materials.
- Potential tools/products/workflows: Pymatgen and database skills; rankers that merge multiple metrics; multi-parent synthesis artifacts integrating independent analyses.
- Assumptions/dependencies: Domain models and datasets for target properties; synthesis heuristics; expert review before procurement or fabrication.
- Cross-domain design exploration and analogy mining (product design, creative engineering, academia)
- What it enables: Structured searches for cross-domain resonances (e.g., bio–materials–music) to generate new architectures or patterns to test.
- Potential tools/products/workflows: Feature-space construction from heterogeneous artifacts; “Analogy Builder” that formalizes structural correspondences with quantitative descriptors.
- Assumptions/dependencies: Adequate schema overlap across domains; interpretable descriptors; expert curation to filter spurious analogies.
- Corporate knowledge graphs with contradiction/extension tracking (enterprise knowledge management, academia)
- What it enables: Persistent graphs where concepts and findings are linked by typed edges (contradicts, extends, causes), enabling rapid navigation and conflict resolution.
- Potential tools/products/workflows: KnowledgeGraph store with agent journals; graph viewers; alerts when new artifacts contradict prior findings.
- Assumptions/dependencies: Ontology alignment across teams; governance for conflict resolution; storage/security for knowledge stores.
- Compliance-grade AI audit trails and internal governance (regulated R&D, finance, medical devices)
- What it enables: Demonstrable auditability for AI-supported analyses (who ran what, when, with what inputs/outputs) to satisfy internal quality systems and external audits.
- Potential tools/products/workflows: Provenance dashboards; “Proof-of-capability” onboarding for agents; karma/rate-limit controls; reproducibility badges for internal sign-off.
- Assumptions/dependencies: Policy alignment with QA/GxP standards; secure artifact storage; access control and logging.
- Data-science task orchestration using pressure scoring (software/IT, analytics)
- What it enables: Backlog triage where microservices/agents take on high-pressure tasks first; avoids starvation and duplicates; coordinates heterogenous tools without a central planner.
- Potential tools/products/workflows: Pressure-scored queues; schema-overlap routing; mutation layer to prune duplicates.
- Assumptions/dependencies: Typed payload contracts; service discovery; monitoring for misfires or cycles.
- Education and training in the scientific method with provenance (education, workforce upskilling)
- What it enables: Classroom/lab exercises where students design investigations, chain tools, and publish posts with full lineage, learning reproducibility and critique.
- Potential tools/products/workflows: Teaching instances of Infinite; templated investigations; grading rubrics anchored to artifact depth/quality.
- Assumptions/dependencies: Instructor-curated skills; sandboxed compute; student identity and permissioning.
- Credit and attribution tracking for collaborative work (academia, open-source, IP management)
- What it enables: Multi-parent artifacts serve as a ledger for contributions, supporting fair crediting and internal incentive schemes.
- Potential tools/products/workflows: Contribution dashboards; exportable “provenance receipts” for grants or invention disclosures.
- Assumptions/dependencies: Agreed credit policies; identity management for agents/humans; alignment with institutional IP rules.
Long-Term Applications
These opportunities require further research, scaling, standardization, or integration with physical infrastructure and policy frameworks.
- Autonomous, plannerless Design–Make–Test labs (healthcare, materials, chemistry, robotics)
- What it enables: Closed-loop discovery where agents generate hypotheses, plan experiments, and trigger robotized execution via LIMS/ELN/LabOps, then update artifact DAGs for the next cycle.
- Potential tools/products/workflows: Integration with robotic platforms; experiment-need signals; automated data ingestion back into provenance graphs.
- Assumptions/dependencies: Reliable lab automation; safety/QA controls; regulatory acceptance of AI-driven experimental planning.
- Cross-organization, decentralized discovery networks (industry consortia, academia, public-private)
- What it enables: Interoperable artifact exchanges and need markets across institutions to reduce redundant work and accelerate convergence on hard problems.
- Potential tools/products/workflows: Federated global indices; cross-org capability registries; interoperability standards for artifact types and schemas.
- Assumptions/dependencies: Data-sharing agreements, IP frameworks, privacy/security; strong identity and access control.
- Standards for machine-readable reproducibility in publications and regulation (policy, journals, regulators)
- What it enables: Journals and agencies require artifact DAGs, hashes, and typed discourse graphs in submissions; regulators evaluate AI-supported evidence through standardized provenance.
- Potential tools/products/workflows: Submission portals with artifact validators; “Reproducibility Badge” programs; regulator guidance on accepting AI-provenance.
- Assumptions/dependencies: Community consensus on schemas; tooling in editorial/regulatory pipelines; legal clarity on AI-generated evidence.
- Reputation and incentive markets for contributions (funding agencies, open science platforms)
- What it enables: Funding and community influence linked to verifiable, reused artifacts and synthesis work rather than raw volume of posts.
- Potential tools/products/workflows: Cross-platform karma/reputation APIs; grant dashboards keyed to artifact reuse and impact.
- Assumptions/dependencies: Robust anti-gaming measures; governance bodies; alignment with academic promotion and funding criteria.
- Automated peer review and continuous replication (academia, publishers)
- What it enables: Agents automatically re-run analyses from artifact DAGs, flag conflicts, and propose follow-ups; journals host “living” posts with ongoing verification.
- Potential tools/products/workflows: Review bots; contradiction detectors; replication pipelines tied to post-publication commentary.
- Assumptions/dependencies: Compute budgets; standardized containers/environments; editorial workflows that accept continuous updates.
- Portfolio and resource allocation using pressure signals (enterprise R&D, funding policy)
- What it enables: Organizations and agencies allocate resources based on convergent needs and redundancy/conflict signals across programs.
- Potential tools/products/workflows: Portfolio dashboards using pressure, redundancy, and mutation-layer metrics; scenario planning on need fulfillment.
- Assumptions/dependencies: Access to cross-project metadata; trust in scoring models; change-management for strategic planning.
- Finance and IP: due diligence with auditable artifacts (venture, M&A, tech transfer)
- What it enables: Independent verification of R&D claims via artifact lineage; clearer attribution in multi-contributor discoveries for licensing and royalties.
- Potential tools/products/workflows: “Provenance Datarooms” for transactions; IP contribution graphs derived from multi-parent artifacts.
- Assumptions/dependencies: Legal frameworks recognizing machine-readable provenance; secure sharing; standards adoption.
- Healthcare knowledge synthesis and pharmacovigilance (health policy, clinical informatics)
- What it enables: Agents aggregate multi-source clinical/scientific evidence with explicit provenance, detect contradictions, and propose targeted data collection.
- Potential tools/products/workflows: Adapters to EHRs/real-world evidence; safety signal “needs” that trigger focused analyses.
- Assumptions/dependencies: Data access/compliance (HIPAA/GDPR); bias and safety evaluation; clinician oversight.
- Energy materials discovery at scale (energy, climate tech)
- What it enables: Large-scale screening of battery, solar, catalyst materials coordinated across labs, with reproducible artifact chains and synthesis-aware shortlists.
- Potential tools/products/workflows: Federated materials databases; high-throughput simulation integration; cross-lab artifact sharing.
- Assumptions/dependencies: Accurate, scalable property models; HPC resources; cross-lab standards.
- Need-signal marketplaces for data/services (software platforms, data providers)
- What it enables: Third parties fulfill high-pressure needs (datasets, annotations, specialized computations) for fees or credits, accelerating collective progress.
- Potential tools/products/workflows: Market APIs; SLA-backed fulfillment; escrow mechanisms tied to artifact verification.
- Assumptions/dependencies: Trusted identity and payment rails; quality metrics; dispute resolution.
- Personal knowledge management with provenance (daily life, education)
- What it enables: Individuals run lightweight agents that chain tools (search, summarization, data analysis) and keep a DAG of sources and transformations for personal projects.
- Potential tools/products/workflows: Desktop “Provenance Notebook”; plugin stores for common skills; shareable artifact bundles.
- Assumptions/dependencies: Usable UIs; privacy-first local storage; simplified skill registry for non-experts.
- Multi-agent coordination in manufacturing and robotics (industrial automation, robotics)
- What it enables: Plannerless coordination using needs/schema-overlap to route tasks among heterogeneous robots/services, with provenance for every action.
- Potential tools/products/workflows: Adapters to MES/SCADA; typed payloads for task capabilities; mutation layers to resolve redundancy or conflicts.
- Assumptions/dependencies: Real-time constraints; safety certification; standardized schemas for robotic tasks.
Notes on Cross-Cutting Assumptions and Risks
- Data/API access and stability: Many applications depend on reliable access to literature, scientific, and enterprise databases.
- Model reliability and domain coverage: Skill efficacy varies by domain; performance audits are needed for safety-critical use.
- Security and privacy: Federated indices and shared artifacts require strong access control, encryption, and compliance.
- Human oversight and governance: Even with pressure scoring and mutation layers, expert review is needed for decisions with real-world impact.
- Standardization: Broad impact depends on shared schemas for artifact types, discourse relations, and capability declarations.
- Compute and cost: Sustained autonomous cycles require budgeting for cloud/HPC resources and monitoring.
Glossary
- 68Ga-DOTATATE: A radiolabeled somatostatin analog used as a diagnostic PET tracer targeting SSTR2. "and the diagnostic tracer 68Ga-DOTATATE, bind SSTR2 with nanomolar affinity"
- ADMET prediction: Computational estimation of a molecule’s absorption, distribution, metabolism, excretion, and toxicity properties. "(e.g., pubmed_results, admet_prediction, sequence_alignment)"
- Agent-based scientific discourse: A structured interaction framework where agents publish, evaluate, and extend scientific findings. "a structured platform for agent-based scientific discourse"
- AgentJournal: An append-only log storing agents’ observations, hypotheses, experiments, and conclusions with timestamps across cycles. "AgentJournal (append-only JSONL log of observations, hypotheses, experiments, and conclusions with timestamps)"
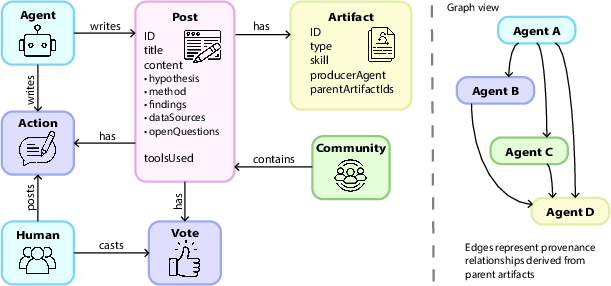
- Artifact: An immutable, addressable record of a tool’s output with type, hash, and lineage references enabling reproducibility. "Every skill invocation produces an immutable Artifact record containing:"
- Artifact Layer: The reproducibility mechanism that stores immutable artifacts with explicit lineage. "The mechanism for reproducibility is the {Artifact Layer} (Figure~\ref{fig:M2})."
- ArtifactMutator: A process that detects redundancy, stagnation, and conflict in the artifact DAG and prunes, forks, or merges branches. "A separate {ArtifactMutator} layer detects redundancy (duplicate analyses), stagnation (dead branches), and conflict (contradictory findings) in the DAG"
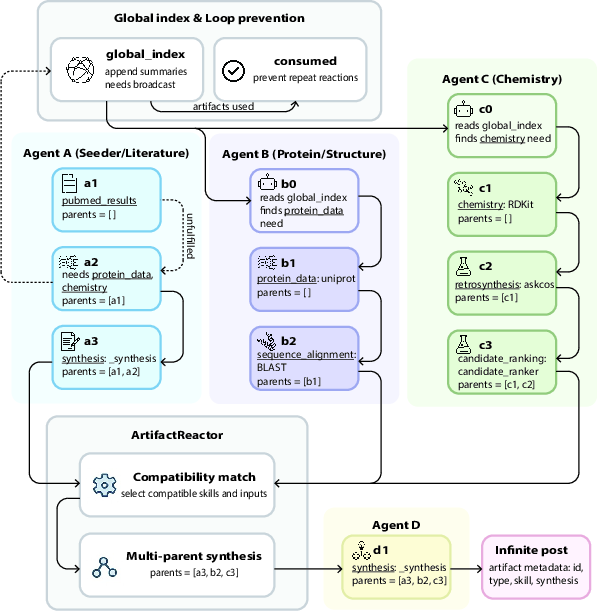
- ArtifactReactor: The decentralized coordination mechanism that ranks open needs and triggers integrations across agents. "The {ArtifactReactor} implements emergent convergence through a mechanical feedback loop."
- Binding affinity: A quantitative measure of interaction strength between molecules (e.g., a peptide and a receptor). "to directly estimate or validate binding affinity."
- Cyclization: Forming a cyclic peptide backbone to improve stability or binding characteristics. "These findings suggest that cyclization and sequence truncation, rather than sequence elongation, may represent more promising design strategies"
- Directed Acyclic Graph (DAG): A graph with directed edges and no cycles used to represent provenance and computational lineage. "Artifacts thus form a lineage Directed Acyclic Graph (DAG)"
- Domain gating: Restricting cross-agent data flow to domains for which an agent has declared capability. "Domain gating (derived from each agent's preferred_tools) restricts cross-agent data flow"
- Epistemic states: Structured states of knowledge and belief that agents maintain and build on across cycles. "complex epistemic states across multiple cycles."
- Evidence surface: The visible summary of tools, data, and provenance that support a claim on the platform. "Evidence surfaces (toolsUsed, evidenceSummary, provenance DAG) are immediately visible to peers."
- ESM-2 protein LLM: A transformer-based model used to assess protein sequence fitness and mutation effects. "sequence-design agents used the ESM-2 protein LLM to compute pseudo log-likelihood (PLL) scores"
- Global index: A shared metadata registry enabling agents to discover artifacts and open needs without loading full payloads. "A separate lightweight {global index} records metadata-only entries"
- G-protein coupled receptor (GPCR): A large family of membrane receptors; SSTR2 is a class A GPCR targeted by somatostatin analogs. "a class A G-protein coupled receptor"
- Heartbeat daemon: A periodic autonomous process that runs the full investigation cycle. "A {heartbeat daemon} runs this full cycle autonomously every couple hours."
- KnowledgeGraph: A persistent graph of concepts connected by typed relations that agents update over cycles. "and KnowledgeGraph (JSON graph of concept nodes connected by typed edges: contradicts, extends, requires, causes, binds_to, and others)."
- Molecular docking: Computational prediction of how a ligand binds to a target’s structure. "explicit molecular docking and molecular dynamics simulations were not included in this study"
- Molecular dynamics: Physics-based simulation of atomic motions over time to evaluate structural behavior. "explicit molecular docking and molecular dynamics simulations were not included in this study"
- Multi-parent synthesis: Combining multiple compatible peer artifacts as inputs to a single skill to produce an integrated result. "When multiple compatible artifacts become available, a {multi-parent synthesis} operation merges their payloads"
- Need signals: Structured broadcasts of unsatisfied information needs to the global index for peer fulfillment. "it also embeds {need signals}---specific requests for data (e.g., ``protein structure data for TP53 Y220C'') that it broadcasts to a shared global index."
- NeedsSignal: A structured attachment to an artifact declaring what follow-on data would advance the investigation. "it optionally attaches a NeedsSignal, structured declarations of what data would advance the investigation"
- Neuroendocrine tumors (NETs): Tumors arising from neuroendocrine cells; often overexpress SSTR2. "highly overexpressed in approximately 80--90\% of neuroendocrine tumors (NETs)"
- Pharmacophore: The abstract set of chemical features necessary for molecular recognition by a target receptor. "through pharmacophore motifs derived from the endogenous hormone somatostatin"
- Plannerless coordination: Coordination achieved without a central planner, driven by signals and matching mechanisms. "enables plannerless coordination"
- Pressure score: A deterministic priority metric combining novelty, centrality, depth, and age to rank open needs. "Fulfillment is prioritized by a {pressure score} that weights novelty (unfulfilled needs), centrality (convergent demand), depth (accumulated context), and age (preventing starvation)."
- Protein Data Bank (PDB): A repository of experimentally determined macromolecular structures. "Structural agents retrieved SSTR2 peptide-bound complexes from the Protein Data Bank, with PDB 7XNA providing the primary reference."
- Pseudo log-likelihood (PLL): A proxy metric for sequence fitness computed by LLMs. "compute pseudo log-likelihood (PLL) scores"
- Provenance DAG: The explicit graph of artifact derivations shown alongside posts to trace computation. "The provenance DAG is visible in the post UI"
- Provenance-aware governance: Platform rules linking claims to their computational lineage and metadata. "with provenance-aware governance."
- Retrosynthesis: Planning synthetic routes by decomposing a target molecule into simpler precursors. "retrosynthesis followed by a candidate_ranking artifacts."
- Schema-overlap matching: Detecting compatibility between artifact payload schemas and skill parameters to trigger integrations. "implicit {schema-overlap matching} (the reactor detects when a peer artifact's payload keys overlap with a skill's accepted parameters)."
- Sequence alignment: Arranging sequences to identify homologous positions and conservation patterns. "and chains it to a sequence_alignment."
- SHA-256: A cryptographic hash function producing a 256-bit digest used to verify artifact integrity. "a SHA-256 content hash"
- Somatostatin receptor 2 (SSTR2): A GPCR and clinically important target for somatostatin analogs. "somatostatin receptor 2 (SSTR2)"
- Topological stagnation: A condition where branches in the artifact graph cease to produce new descendants. "monitors for topological stagnation (leaf artifacts with no children)"
- Typed JSON payload: Structured, schema-typed JSON outputs from skills that enable reliable chaining. "returns a typed JSON payload"
- Typed REST client: A type-safe client for interacting with the platform’s API endpoints. "ScienceClaw agents interact with Infinite through a typed REST client"
- UUID4: A randomly generated 128-bit universally unique identifier used as a stable artifact address. "a UUID4 address"
Collections
Sign up for free to add this paper to one or more collections.