- The paper presents RSD as a novel framework that employs local greedy updates to significantly reduce Pauli weights in fermion-to-qubit mappings.
- It demonstrates substantial reductions, such as >40% in 2D lattice models and >30% in the Hubbard model, outperforming traditional methods and simulated annealing.
- The approach leverages interchangeable local solvers and varied sampling strategies (uniform, Hamming-weight, ML-guided) to achieve robust and scalable optimization.
Randomized Subsystem Descent for Fermion-to-Qubit Mapping
Introduction and Motivation
The design and optimization of fermion-to-qubit mappings constitute a core bottleneck in Hamiltonian simulation on quantum hardware, directly affecting simulation resource costs such as Pauli weight and operator locality. Existing global search algorithms, notably simulated annealing for Clifford-circuit optimization, yield effective reductions but are severely limited by computational overhead, typically failing to scale beyond O(100) modes even with multi-day CPU runtimes. This paper introduces Randomized Subsystem Descent (RSD), an algorithmic framework inspired by block coordinate descent, to enable scalable and efficient optimization for large-scale fermion-to-qubit mappings. RSD reformulates the global non-smooth, combinatorial Clifford optimization problem as a stochastic greedy search, iteratively targeting small subsystems to minimize cost metrics such as Pauli weight.
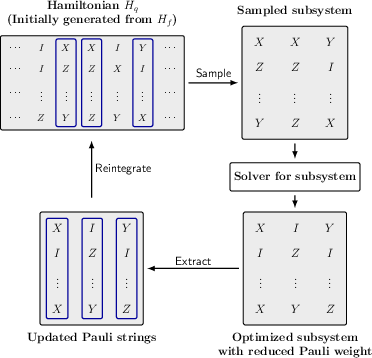
Figure 1: Workflow for the Randomized Subsystem Descent, depicting iterative selection, optimization, and reintegration of tractable subsystems.
Algorithmic Framework
The core technical contribution is the RSD procedure, which, at each iteration, samples a k-qubit subsystem, applies an exact or efficient Clifford solver to minimize a target cost (e.g., Pauli weight), and greedily reintegrates the optimized subsystem into the global Hamiltonian only if it improves the cost. The flexibility to substitute alternative local solvers and sampling strategies is inherent.
Subsystem Sampling Strategies
Three sampling strategies are discussed:
- Uniform random sampling (agnostic to system structure),
- Hamming-weight informed sampling (favoring high-contributing qubits),
- Machine Learning (ML)-guided policies (predicting high-impact subsystems),
with Hamming-weight-based sampling yielding the optimal balance of simplicity, computational tractability, and result quality for most tested regimes. ML-based policies offer superior early convergence but require additional tuning and data/model overhead.
Numerical Results: Lattice and Molecular Benchmarks
1D and 2D Lattice Hopping Models
RSD is benchmarked on extensive 1D and 2D lattice hopping models, showing clear and consistent advantage over all conventional mappings (Jordan-Wigner, Bravyi-Kitaev, ternary tree) and competitiveness or superiority over simulated annealing approaches, especially as system sizes increase.
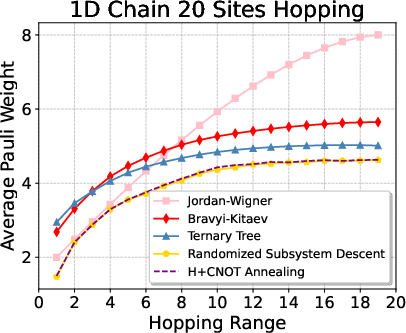
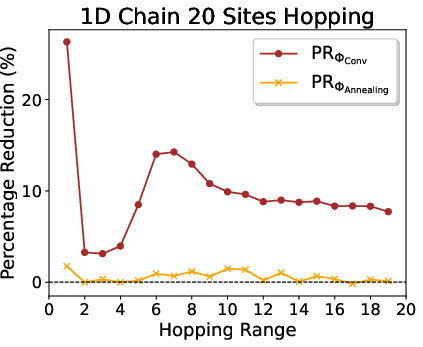
Figure 2: Average Pauli weights and reductions for 1D lattice hopping with varying interaction ranges for N=20.
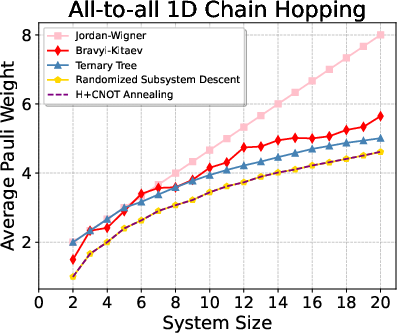
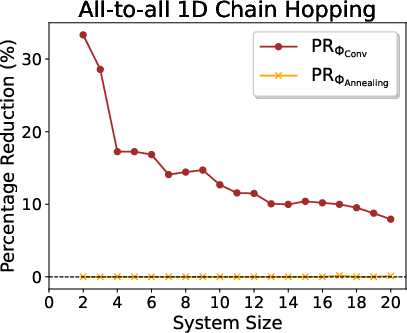
Figure 3: Benchmark results for all-to-all 1D hopping as a function of system size—RSD consistently yields lower final Pauli weights.
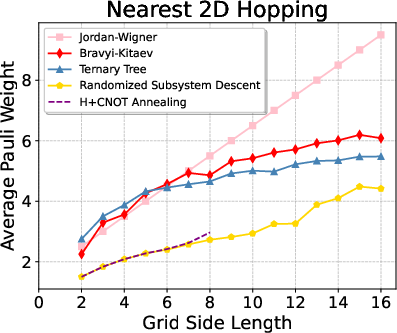
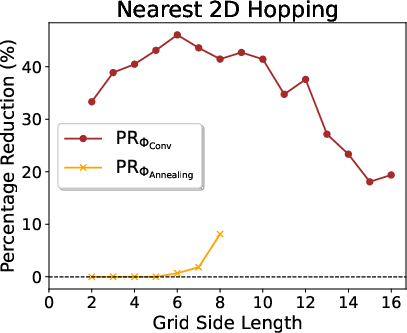
Figure 4: Pauli weight trends and reduction percentages for 2D nearest-neighbor hopping models (up to 16×16 systems).
Notably, for 2D models, RSD achieves >40% reduction in Pauli weight for modest system sizes (side up to 10), and maintains ~20% savings up to 16×16 systems, with rapid empirical convergence.
Hubbard Model
On the highly nontrivial 2D Hubbard model, RSD demonstrates strong empirical scaling, achieving more than 30% reduction in Pauli weight for intermediate grids, and sustaining >10% savings in the 16×16 regime, which is out of reach for global heuristics such as simulated annealing.
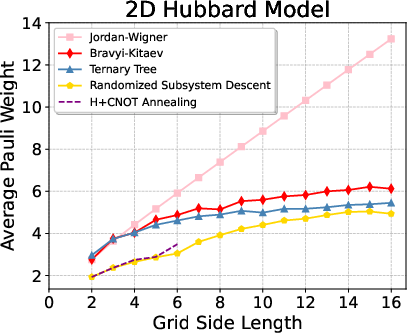
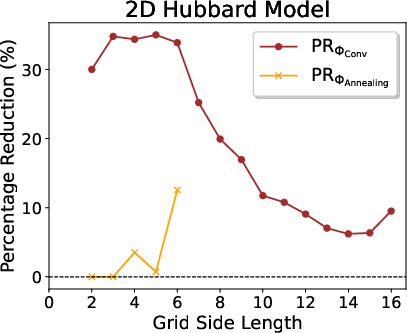
Figure 5: Pauli weight and percentage reductions for the 2D Hubbard model across grid sizes, with RSD outperforming all other approaches.
Analysis of width and depth hyperparameters for the subsystem solver shows that RSD rapidly plateaus to a near-optimal solution with modest local solver resources, indicating limited marginal returns for further local optimization depth/width.
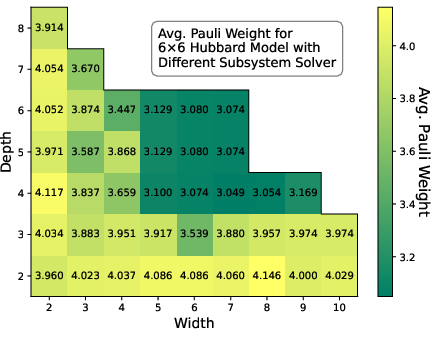
Figure 6: Heatmap of average Pauli weight for 6×6 Hubbard using different local solver depths and widths shows RSD closely approaches the optimal regime.
Molecular Electronic Structure Hamiltonians
On molecular systems (up to 54 modes / 180,000+ Pauli strings in the 6-31G basis), RSD is benchmarked against state-of-the-art methods, including the hardware-aware HATT mapping, using both total and weighted Pauli weight objectives. Substantial reductions (10–25%) are achieved for all systems, with up to 25% reduction in weighted Pauli weight for challenging cases such as NaF (STO-3G) and O2 (6-31G).
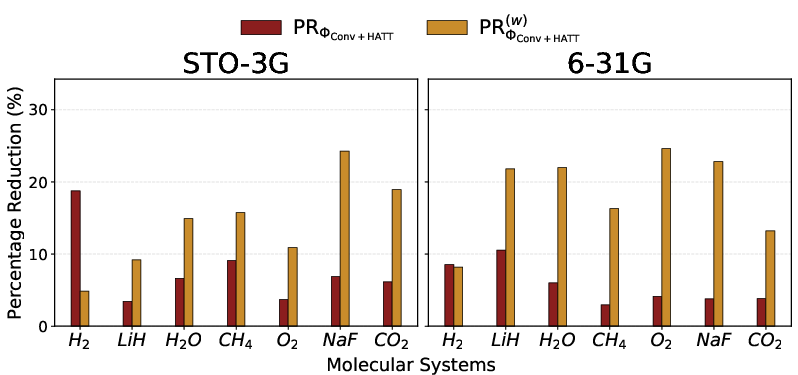
Figure 7: Percentage reduction in total and weighted Pauli weight achieved by RSD for a suite of molecular systems compared to the best conventional and HATT mappers.
The modest per-instance computational cost (typically capped at 2000 iterations for molecular benchmarks) indicates significant untapped potential for deeper optimization by increasing runtime.
Analysis of Sampling and Convergence
A detailed comparison of sampling strategies shows that ML-based heuristics yield accelerated initial descent in Pauli weight, but plateau due to training data and model limitations. Hamming-weight-driven sampling provides robust and steady convergence, balancing simplicity and state-awareness.
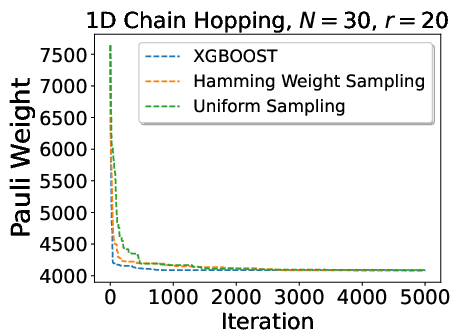
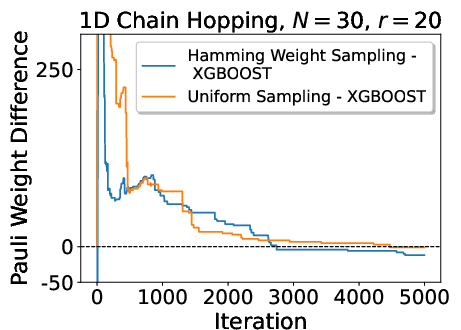
Figure 8: Comparison of Pauli weight minimization as a function of sampling strategy—ML-guided sampling accelerates initial convergence but does not always achieve the lowest final cost.
RSD's monotonic greedy acceptance and local structure yield reliable convergence in practice. Empirical results show that larger models saturate less quickly, suggesting room for improvement with additional iterations; rigorous convergence bounds in the discrete setting remain an open theoretical question.
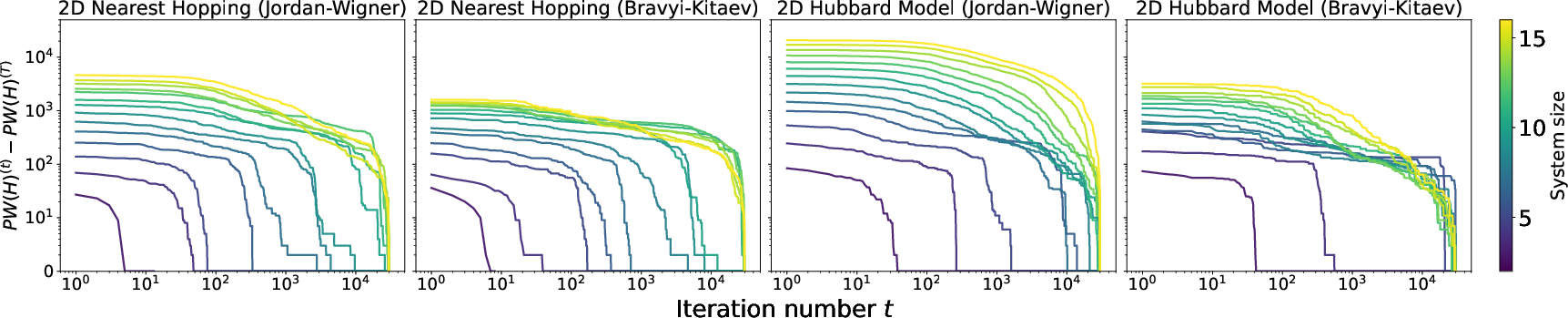
Figure 9: Pauli weight difference PW(t)−PW(T) across 2D lattice models, showcasing convergence behavior with fixed iterations.
Implications and Future Directions
RSD demonstrates that greedy, local, and scalable optimization can systematically and robustly outperform both conventional mappings and global heuristics (simulated annealing) for Hamiltonian encoding problems of realistic size and complexity. This directly enables further quantum simulations with reduced Trotter, qDRIFT, and circuit synthesis resource overheads.
From a theoretical perspective, extending RSD with more powerful subsystem solvers (including non-Clifford operations or continuous parameterizations), ML-accelerated or RL-based sampling strategies, and rigorous guarantees on monotonicity and convergence are natural next steps. The cost-agnostic framework admits alternative objectives, allowing for tailored, hardware- and algorithm-aware encodings—critical for both NISQ and FT quantum computing architectures.
Conclusion
Randomized Subsystem Descent provides a scalable, practical, and conceptually simple framework for fermion-to-qubit mapping optimization, leveraging local greedy updates to systematically suppress simulation overhead at scale. Its versatility, modest computational requirements, and robust empirical performance underscore its promise as a general approach for optimizing Hamiltonian encodings and related quantum compilation primitives.













