- The paper introduces TRIBE, a transformer-based model that integrates text, audio, and video features to predict whole-brain fMRI responses.
- The paper details a methodology using pretrained embeddings and specialized linear layers to standardize dimensions before transformer encoding, capturing 54% of explainable variance.
- The paper demonstrates significant performance improvements in multisensory integration and out-of-distribution scenarios, validated through competition results and modality ablation studies.
TRIBE: TRImodal Brain Encoder for Whole-Brain fMRI Response Prediction
Introduction
The paper "TRIBE: TRImodal Brain Encoder for whole-brain fMRI response prediction" presents a neural network architecture that aims to predict fMRI brain responses to video stimuli across various modalities, cortical regions, and individuals. Historically, neuroscience has tended to focus on specialized domains; this has led to fragmented modeling efforts that overlook integrative and unified representations of cognition. TRIBE addresses this gap by integrating text, audio, and video modalities using a transformer-based architecture to achieve high-precision predictions of spatial and temporal fMRI responses.
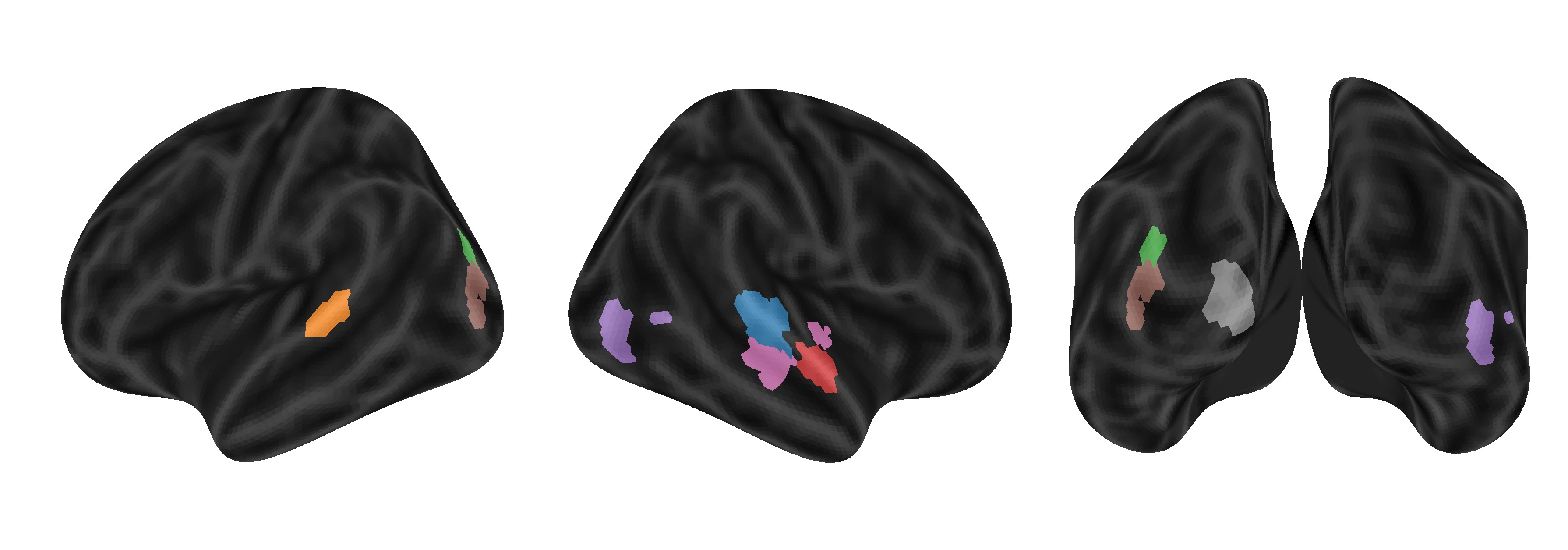
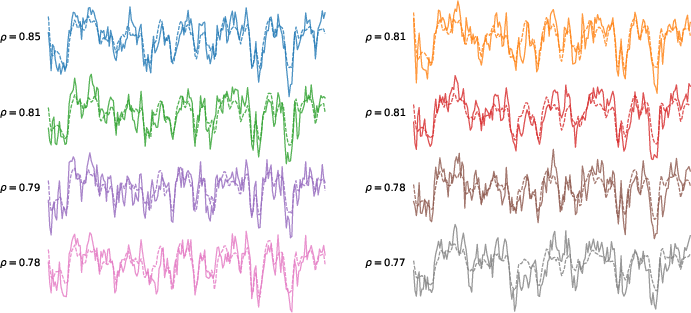
Figure 1: TRIBE predicts brain responses to videos across diverse regions. For eight brain parcels color-coded in the brain images, we report the BOLD response of the first participant to the first 5 minutes of a held-out movie in solid lines and our model's predictions in dashed lines, with the Pearson correlation of the two curves reported on the left.
Methods
Model Architecture
TRIBE leverages pretrained embedding layers from state-of-the-art foundational models for text (Llama-3.2-3B), audio (Wav2Vec-Bert-2.0), and video (Video-JEPA 2). Each modality generates high-dimensional features, which are aggregated through specialized linear layers to standardize their dimensions. Subsequently, these embeddings feed into a transformer encoder that models interactions between these modalities across temporal sequences.
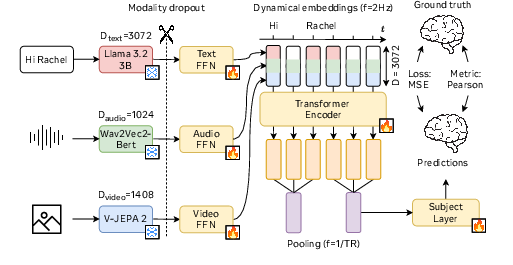
Figure 2: Visual summary of our method.
Dataset and Preprocessing
The model is trained on the Courtois NeuroMod dataset, which features over 80 hours of fMRI recordings from four subjects viewing episodes from TV series and movies. Preprocessing involves projecting BOLD signals to the MNI standard space and parcellating these signals into 1000 parcels using the Schaefer atlas. This greatly reduces computational complexity while preserving functional mapping.
Training Protocol
Optimization is performed using AdamW, incorporating learning rate schedules and modality dropout to ensure robust multimodal integration. The authors employ stochastic weight averaging and ensembling across hyperparameter variations to enhance generalization. Feature extraction is further optimized by using high-throughput GPU resources.
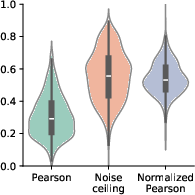
Figure 3: Distribution across parcels.
Results
TRIBE rose to prominence by securing first place in the Algonauts 2025 competition in multimodal brain encoding, demonstrating substantial performance enhancements over competing models. It excelled in predicting responses in associative cortices, crucial for multisensory integration.
Notably, TRIBE maintained robust performance even in out-of-distribution testing scenarios, as depicted through various held-out datasets. Such adaptability illustrates its potential utility across diverse futuristic applications.
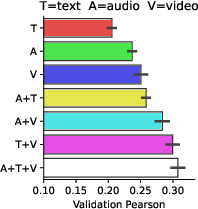
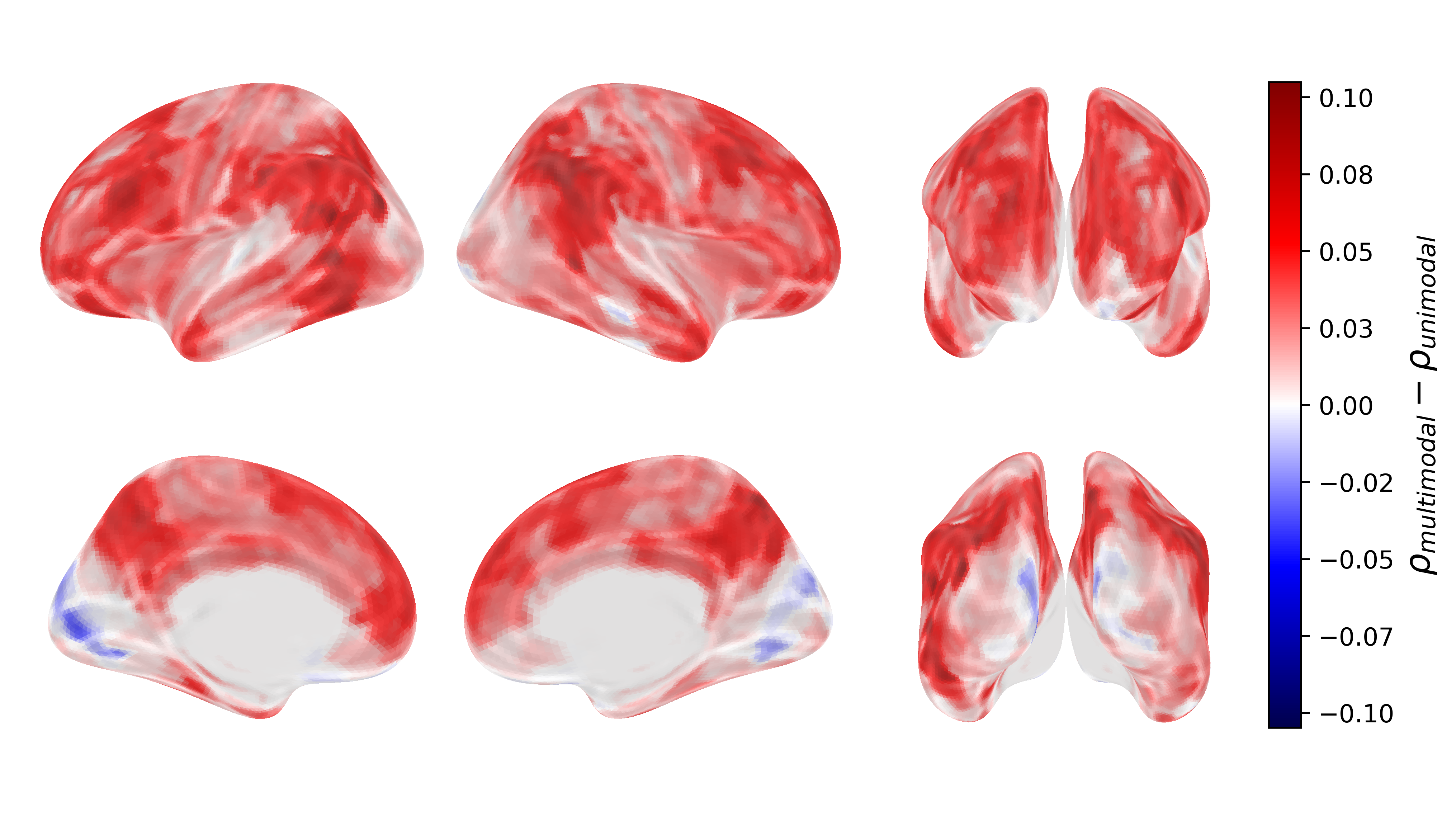
Figure 4: Modality ablations.
Ablation Studies
Ablation analyses reveal that TRIBE's multimodality confers significant predictive advantages compared to unimodal models. Specifically, cross-modal information processing in associative brain regions results in notable performance gains.
Additionally, the noise ceiling analyses elucidate that the model captures 54% of explainable variance within the dataset averages, attesting to its effective modeling capability across brain areas.

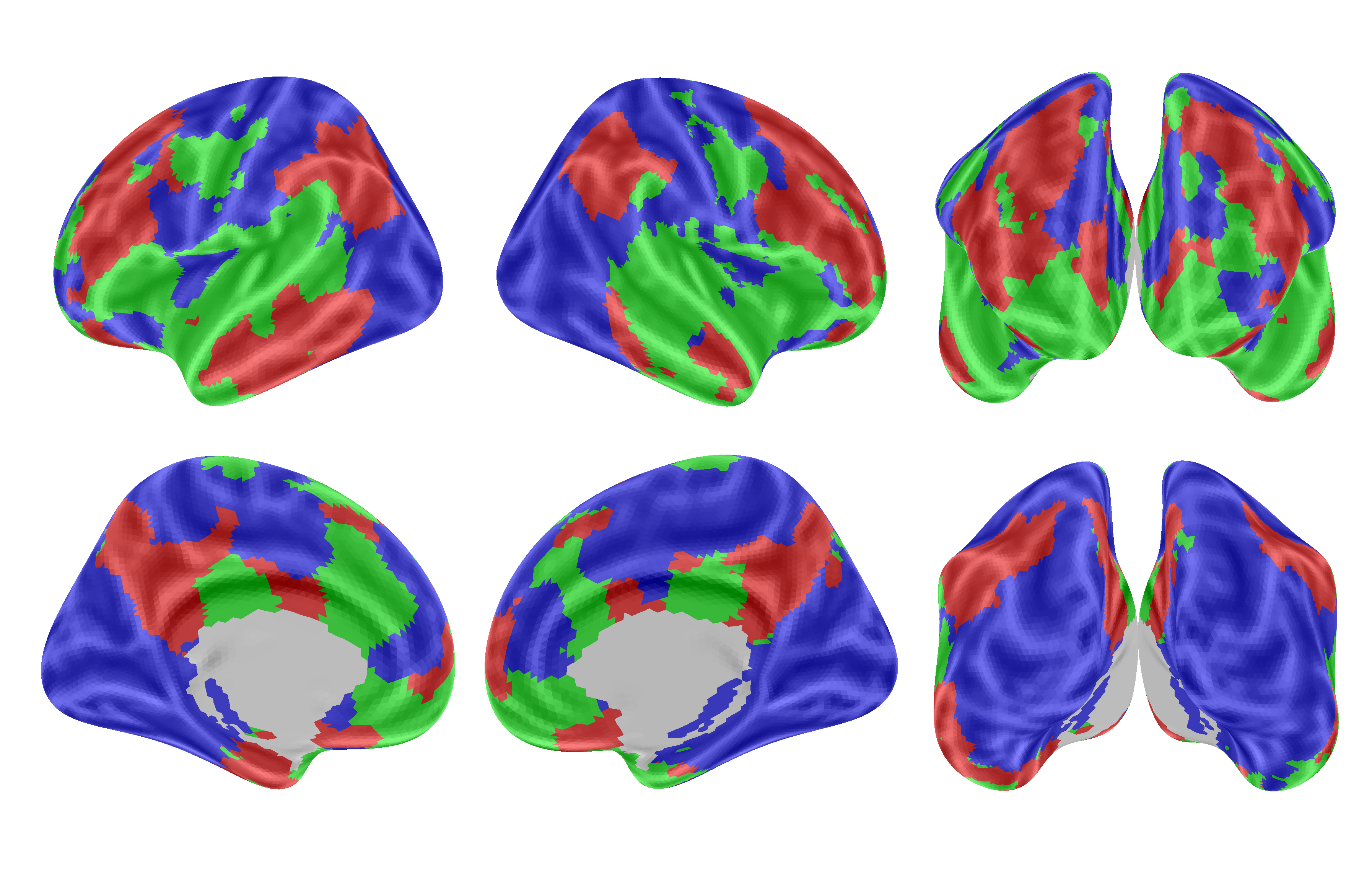
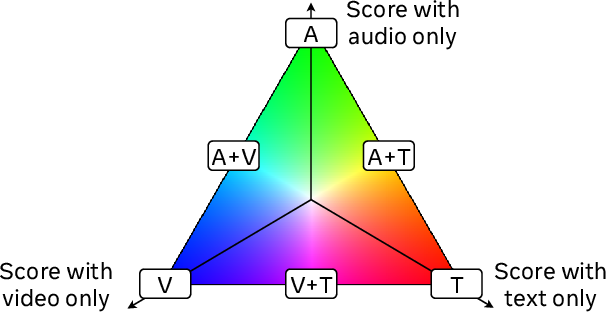
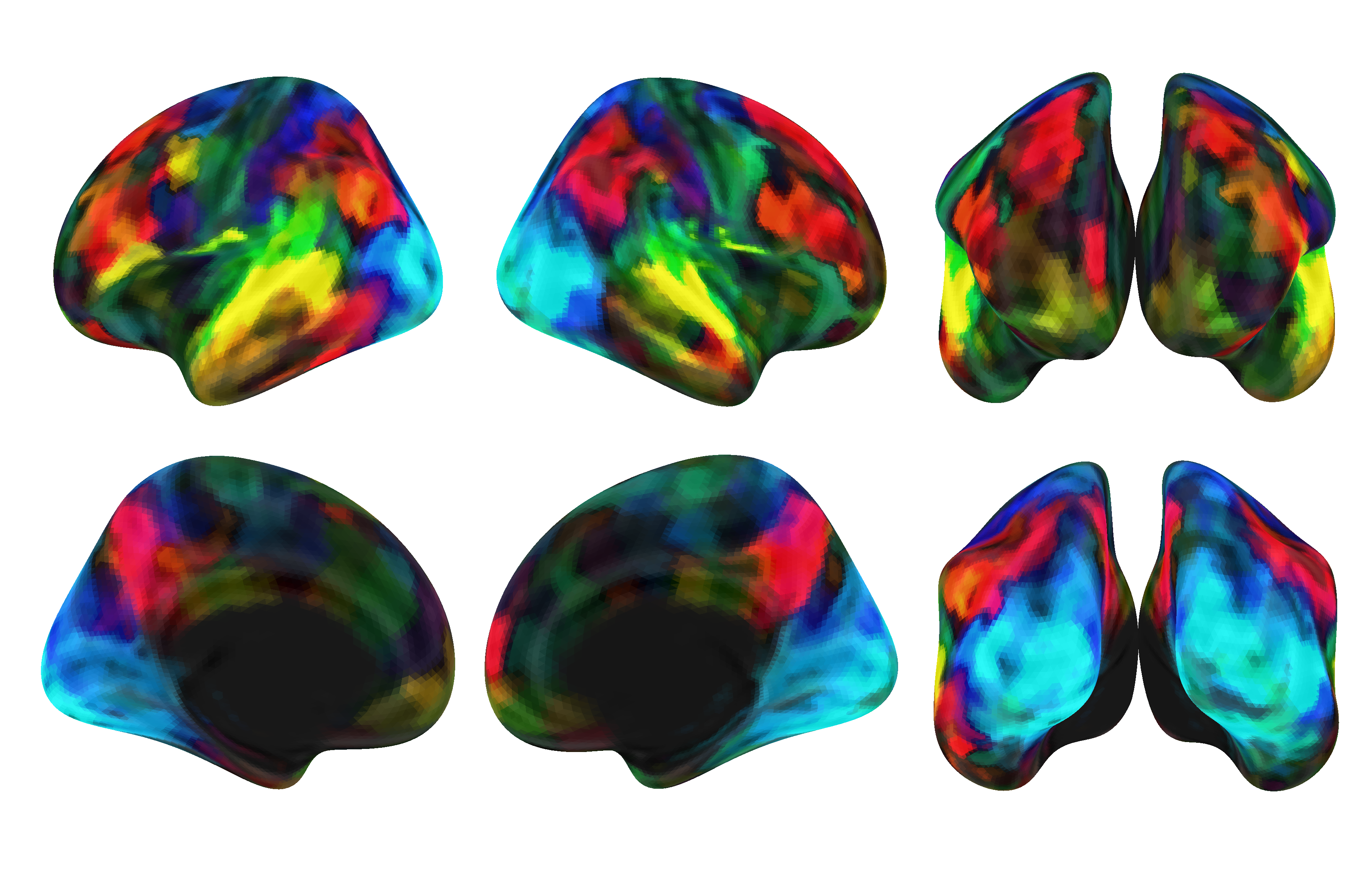
Figure 5: Highest scoring modalities.
Scaling Laws
The study demonstrates promising scaling laws correlated with dataset size and model depth, hinting at further improvements in encoding accuracy with increased data availability and model complexity. These trends stress the importance of scaling models in predicting complex biological systems.
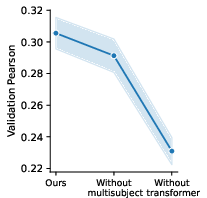
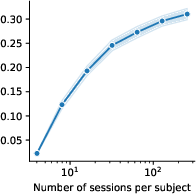
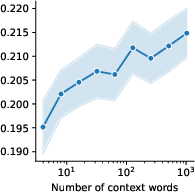
Figure 6: Model ablations.
Conclusion
The TRIBE architecture represents a pioneering effort to encapsulate multimodal and multisubject brain encoding within a singular framework. Its ability to successfully predict whole-brain responses to complex stimuli marks a substantial advancement in achieving integrative models for understanding human cognition, extending broader implications in neuroscience research.
Limitations remain prevalent, including spatial resolution limitations inherent in the use of parcel-level mappings and dependency on fMRI data. Future advances should aim to encompass voxel-level predictions and additional cognitive dimensions like memory and decision-making processes.
The broader impact of TRIBE underscores its applicability for probing deeply into cognitive complexities via naturalistic stimuli, promoting interdisciplinary alignment across neuroscience domains by leveraging AI advancements. This work sets the stage for future endeavors in computational cognition and in silico neuroexperimental frameworks.












