- The paper presents DINOCell, a framework that integrates DINOv2 self-supervision, domain adaptation, and supervised fine-tuning to achieve robust cell segmentation.
- DINOCell leverages a three-stage training protocol with 130,000 unlabeled microscopy images to align visual representations with biological structures.
- Quantitative evaluations demonstrate significant performance improvements over prior models, with notable gains in segmentation accuracy and zero-shot generalization.
Self-Supervised Pretraining of Cell Segmentation Models: DINOCell
Introduction
Cell segmentation in microscopy images underpins quantitative cell biology by enabling automated analysis of cellular morphologies. Progress in deep models for cell instance segmentation is hampered by the scarcity of diverse, high-quality labeled datasets and by a considerable distribution shift between pretraining datasets (typically, natural images) and target microscopy images. "Self-supervised Pretraining of Cell Segmentation Models" (2604.10609) introduces DINOCell: a self-supervised vision transformer-based framework that leverages DINOv2 pretraining and domain adaptation on unlabeled microscopy data to achieve robust instance segmentation. The work delivers both strong quantitative performance and improved representational alignment for microscopy tasks relative to prior methods, directly addressing foundational limitations of segmentation-pretrained models such as SAMCell and Cellpose-SAM.
Methodology
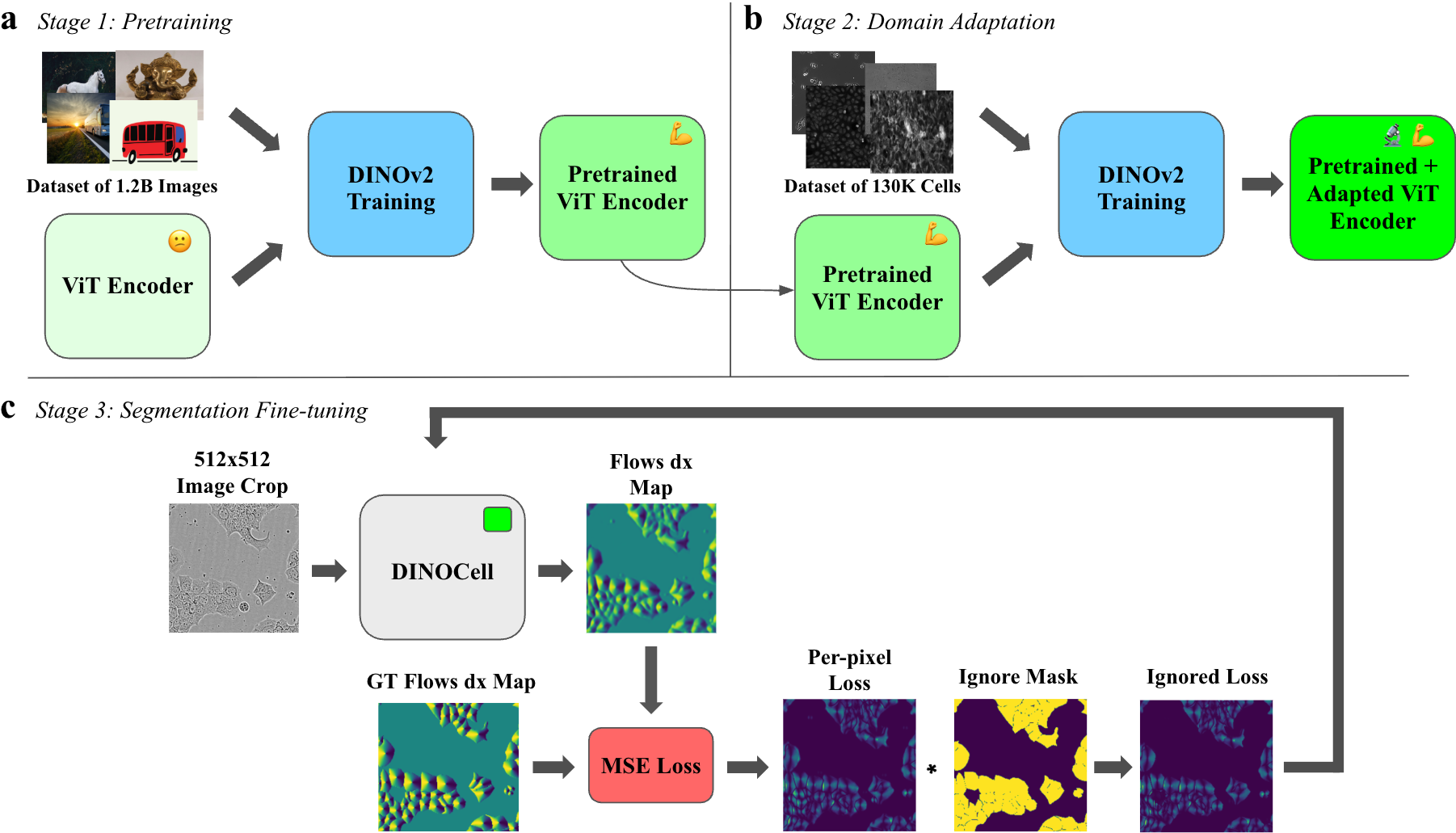
DINOCell employs a three-stage training protocol, integrating large-scale self-supervised visual pretraining with task-agnostic domain adaptation and supervised segmentation fine-tuning:
The model architecture pairs a DINOv2-pretrained ViT-B encoder (re-patched to 8×8, embedding dim 768) with a lightweight upsampling decoder and compact convolutional heads, predicting dense flows and cell probabilities. The segmentation mask is derived via gradient-tracking over predicted flows.
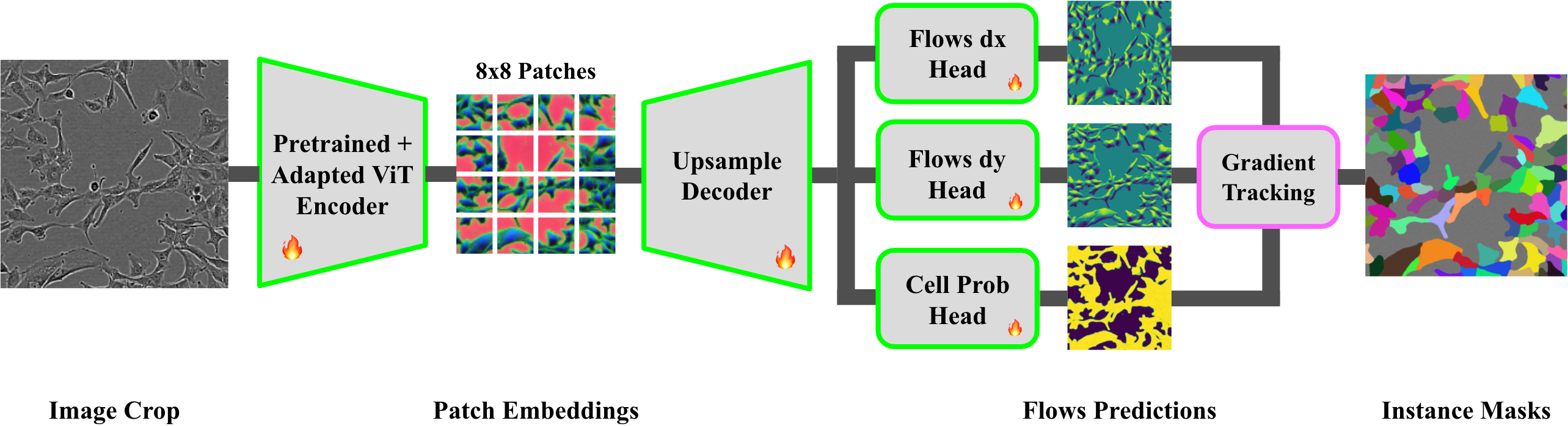
Figure 2: DINOCell architecture, combining a DINOv2-adapted ViT encoder, an upsampling decoder, and convolutional heads for segmentation via flow field decoding.
Preprocessing includes CLAHE and normalization, ensuring robust learning under microscopy-specific image artifacts.
Effects of Pretraining and Domain Adaptation
Ablation studies dissect the impact of different pretraining and adaptation schemes. Models initialized from DINOv2 with or without further domain adaptation, and models pretrained solely using microscopy images, are contrasted with naïvely initialized counterparts. The effect on learned visual representations is visualized using PCA projections of encoder patch embeddings.
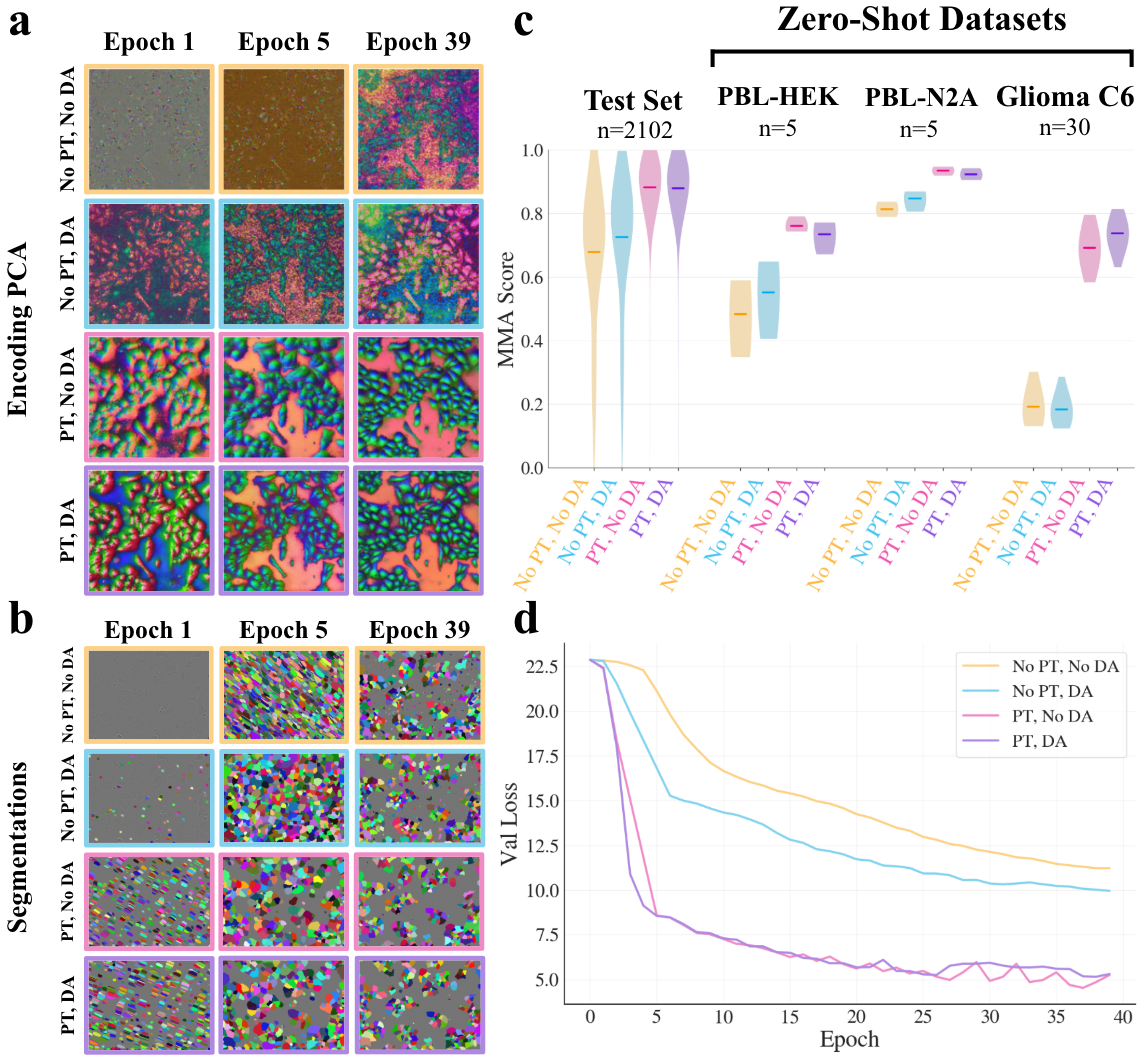
Figure 3: DINOv2 pretraining yields highly structured, object-aligned representations in early fine-tuning; domain adaptation offers modest additional benefit for OOD robustness, especially for highly variable datasets.
The results indicate:
- DINOv2 pretraining produces substantial improvements in both in-domain (fine-tuning test set) and out-of-distribution (OOD) datasets, accelerating convergence and drastically improving representational structure and segmentation accuracy.
- Domain adaptation introduces marginal benefit for challenging OOD data (e.g. Glioma C6), but has negligible impact on in-distribution generalization or convergence for most cases.
- Random initialization or microscopy-only pretraining yields significantly worse performance, slower convergence, and less discriminative encoder feature organization.
Encoder Evolution and Representation Quality
The progressive alignment of encoder representations with cellular substructure is elucidated by tracking the PCA projections through the three training stages.
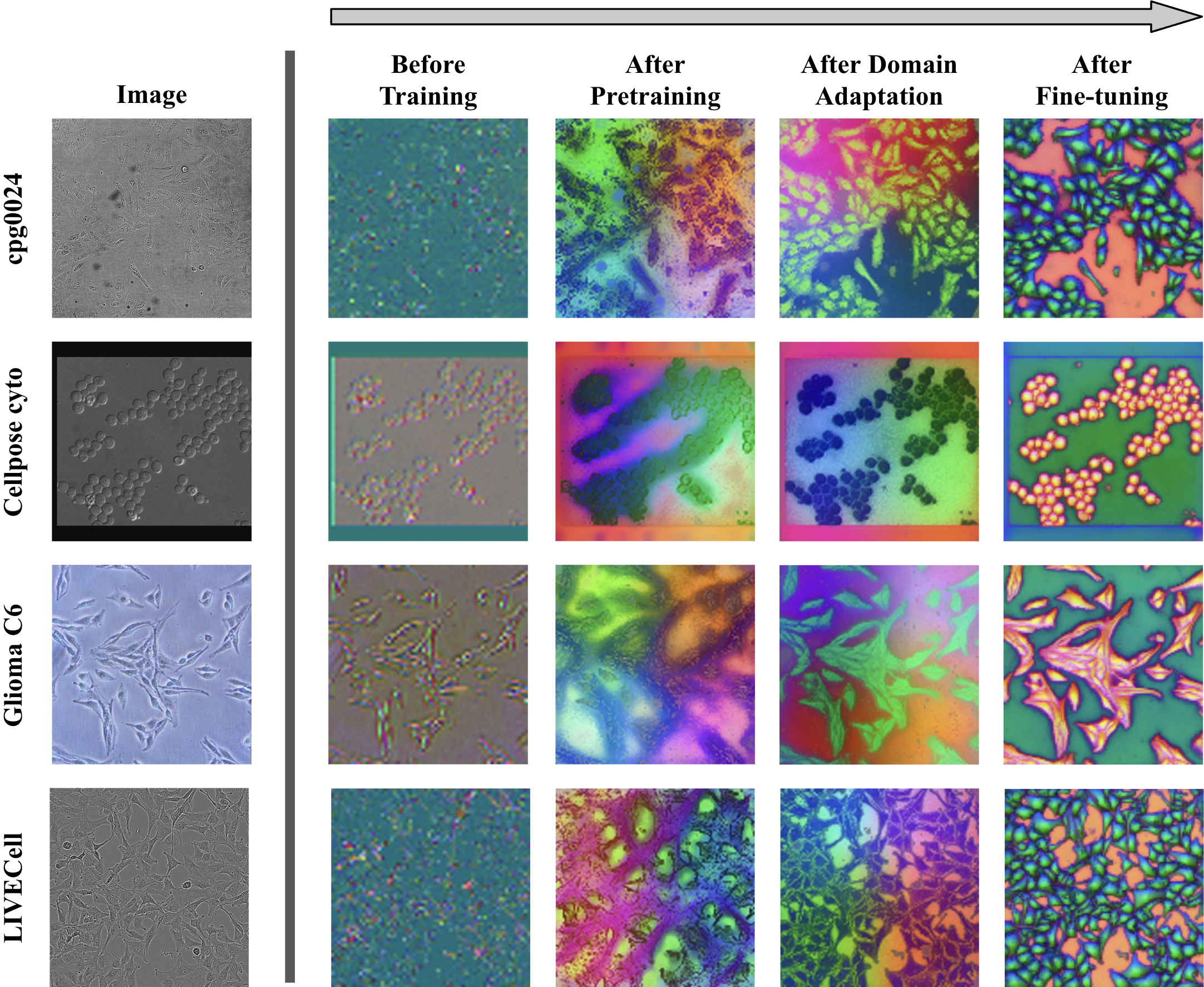
Figure 4: Initial DINOv2 representations are already foreground-background and edge-aware. Domain adaptation clusters features more closely with cell structure, but fine-tuning is essential for sharp object-level organization.
DINOCell’s encoder output transitions from noisy, weakly organized embeddings to highly object-aware, spatially coherent groupings demarcating cell interiors and borders.
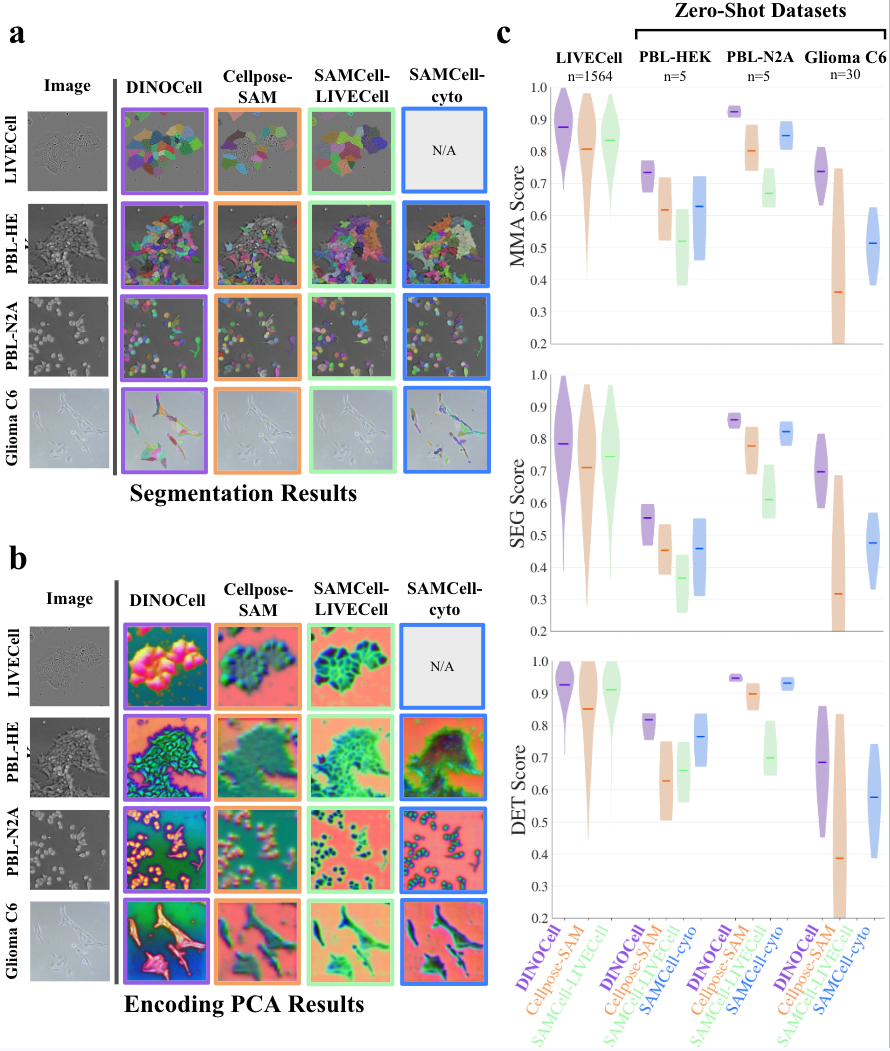
DINOCell’s performance is benchmarked on both aggregate and per-dataset splits using Maximum Matching Accuracy (MMA), SEG, and DET metrics. Results show high accuracy for morphologically simple datasets and strong robustness across diverse imaging conditions and cell types.
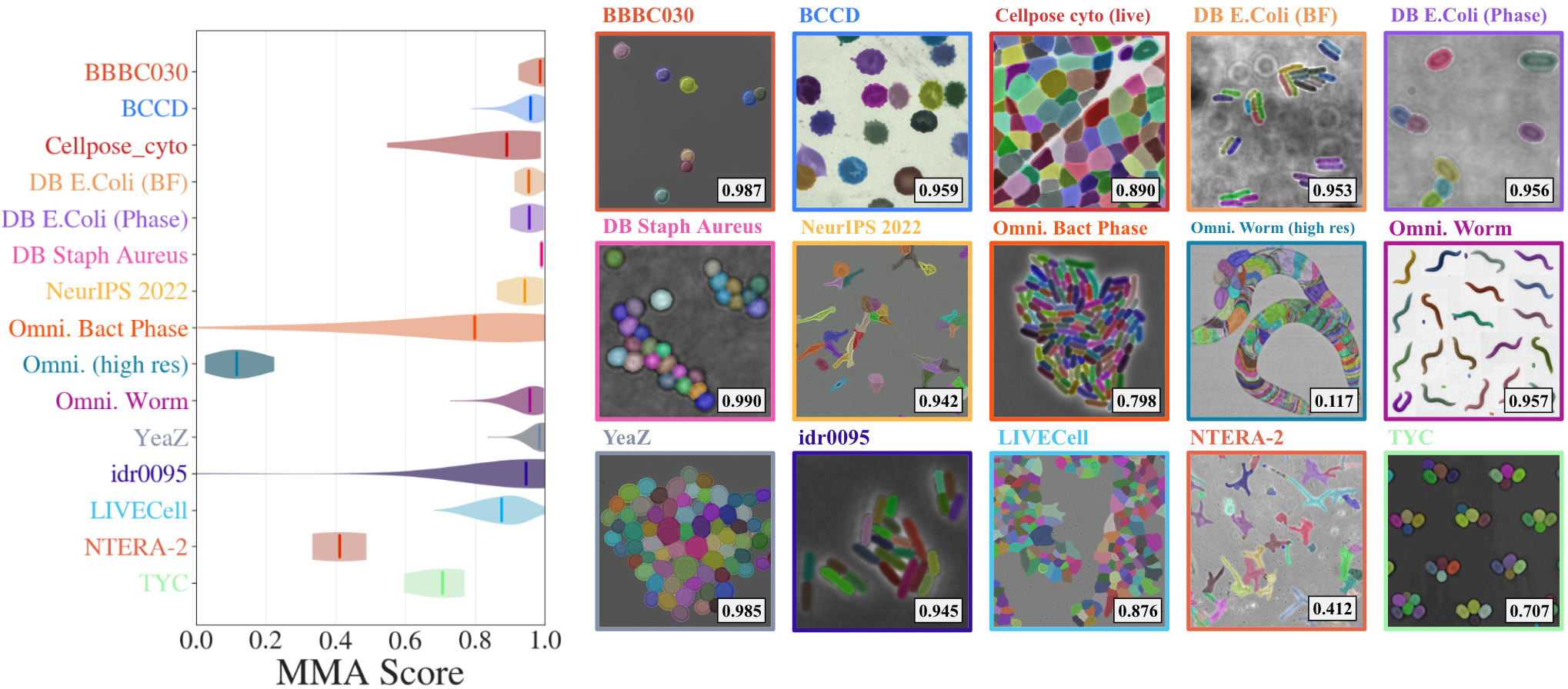
Figure 5: Per-dataset MMA scores reveal near-saturating performance for typical cell morphologies; the model handles variable density, shape, and modality with consistently high segmentation fidelity.
Certain classes—elongated or highly connected instances (e.g., C. elegans—Omnipose Worm) or complex embryonal cells with thin processes (NTERA-2)—pose challenges, resulting in over-segmentation or fragmented masks.
Comparison with Prior Segmentation Models
Head-to-head evaluation with state-of-the-art segmentation-pretrained models (SAMCell, Cellpose-SAM) highlights DINOCell’s quantitative and qualitative superiority:
Principal component analysis of encoder output further confirms that DINOCell’s features are more object-aligned and spatially coherent than those derived from task-pretrained ViTs not domain-adapted for microscopy.
Discussion and Implications
DINOCell establishes that large-scale self-supervised pretraining with subsequent domain adaptation provides a robust foundation for cell segmentation, outperforming segmentation-pretrained models, especially under significant domain shift. This separates the concerns of "what to segment" (domain-level visual structure) from "how to segment" (segmentation objective and decoding mechanism), suggesting that domain-aligned representations are more transferable than task-aligned but domain-mismatched ones.
The negligible effect of extended domain adaptation post-DINOv2 points to an inherent robustness and generalizability in features learned from vast, curated datasets via self-supervision; essential for deploying segmentation models to new biological imaging modalities where annotated data is limited or non-existent.
The introduced ignore-masking technique to address annotation incompleteness in biological datasets also advances practical segmentation performance, reducing overfitting and instability caused by label noise.
The results imply that, for downstream biomedical imaging tasks, leveraging foundation models with self-supervised, domain-aligned pretraining yields models more robust to domain shift than those relying on cross-domain segmentation annotation transfer (e.g., SAM). Further, this approach is likely extensible beyond cell segmentation—other imaging tasks in histology, pathology, and organ segmentation may benefit from similar self-supervised pipelines employing large-scale domain adaptation before target-specific fine-tuning.
Conclusion
DINOCell provides a compelling demonstration that self-supervised vision transformer pretraining, with minimal domain adaptation, offers substantial improvements over task-pretrained models for cell instance segmentation under diverse imaging conditions. The model achieves strong in-domain accuracy, exceptional zero-shot transfer, and qualitatively superior object representations, underlined by consistent outperformance of prior methods across metrics and datasets. This advances the state of the art in label-efficient, domain-robust cell segmentation and motivates further investigation into domain-adaptive self-supervised learning pipelines for broader applications in biological image analysis.