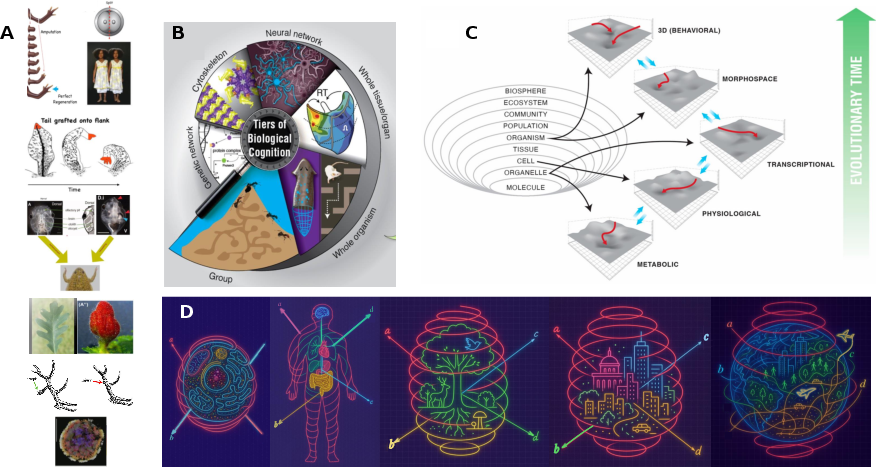
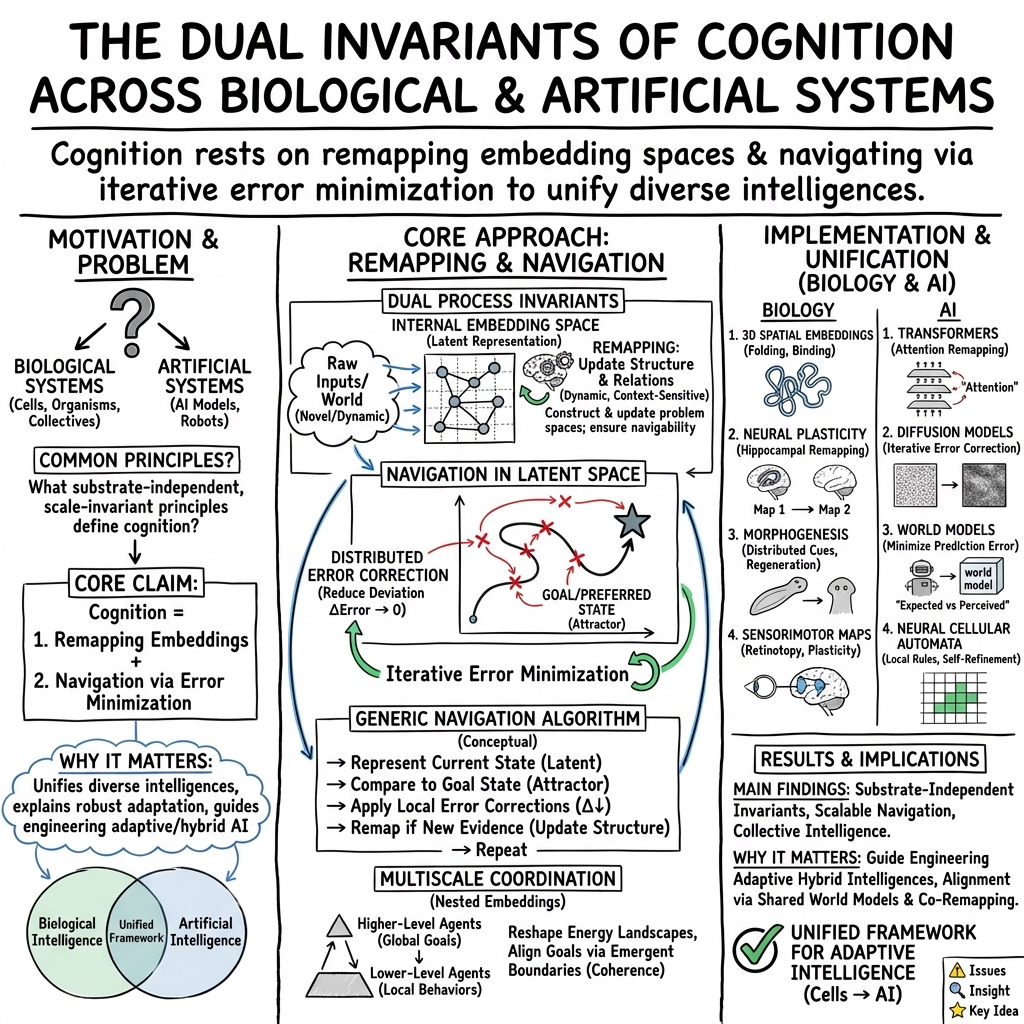
Remapping and navigation of an embedding space via error minimization: a fundamental organizational principle of cognition in natural and artificial systems
Abstract: The emerging field of diverse intelligence seeks an integrated view of problem-solving in agents of very different provenance, composition, and substrates. From subcellular chemical networks to swarms of organisms, and across evolved, engineered, and chimeric systems, it is hypothesized that scale-invariant principles of decision-making can be discovered. We propose that cognition in both natural and synthetic systems can be characterized and understood by the interplay between two equally important invariants: (1) the remapping of embedding spaces, and (2) the navigation within these spaces. Biological collectives, from single cells to entire organisms (and beyond), remap transcriptional, morphological, physiological, or 3D spaces to maintain homeostasis and regenerate structure, while navigating these spaces through distributed error correction. Modern AI systems, including transformers, diffusion models, and neural cellular automata enact analogous processes by remapping data into latent embeddings and refining them iteratively through contextualization. We argue that this dual principle - remapping and navigation of embedding spaces via iterative error minimization - constitutes a substrate-independent invariant of cognition. Recognizing this shared mechanism not only illuminates deep parallels between living systems and artificial models, but also provides a unifying framework for engineering adaptive intelligence across scales.
Paper Prompts
Sign up for free to create and run prompts on this paper using GPT-5.
Top Community Prompts
Explain it Like I'm 14
A simple guide to “Remapping and navigation of an embedding space via error minimization”
What is this paper about? (Overview)
This paper asks a big question: what do very different kinds of “smart” systems have in common? The authors look at living things (like cells, tissues, whole animals) and artificial systems (like LLMs and robots) and argue that they all follow the same basic rule to solve problems:
- They build and update an internal “map” of the world (remapping).
- They use that map to move step by step toward a goal by fixing mistakes (navigation by error minimization).
In short: intelligent systems redraw their maps as they learn, and they use those maps to find their way to what they want.
What questions did the authors ask? (Key objectives)
The paper focuses on four simple questions:
- Can we describe intelligence as two things working together: redrawing a map and finding a path on it?
- Do living systems and AI systems use the same kind of internal maps?
- Is “fixing errors” (reducing the gap between “where I am” and “where I want to be”) the engine that drives smart behavior?
- Does this idea work at many sizes and scales, from molecules to animals to AI models?
How did they study it? (Approach in everyday language)
This is a theory paper. The authors don’t run new lab experiments; instead, they collect many examples from biology and AI and show how the same idea explains them all.
First, some key ideas in plain language:
- Embedding space: Think of this as a map your system makes inside itself. It turns messy, complicated reality into a simpler picture with “directions” that matter (like a subway map that simplifies a whole city).
- Remapping: Redrawing or updating that internal map when something new happens, so it stays useful.
- Navigation: Using the map to move toward a goal, step by step.
- Error minimization: Like a “you’re getting warmer/colder” game. The system compares “where I am now” to “where I want to be” and takes steps that make the difference smaller.
How this looks in biology:
- Cells and tissues don’t just react; they aim for target states (like healthy shape or chemistry) and keep correcting until they get there.
- Brains make maps of space, sounds, smells, tools, and body movements; these maps can be redrawn when we learn or lose a sense (for example, in blindness, parts of the brain “remap” to process sound).
- Even without a brain, cell groups can “aim” for a body plan during development or regeneration by using chemical and electrical signals.
How this looks in AI:
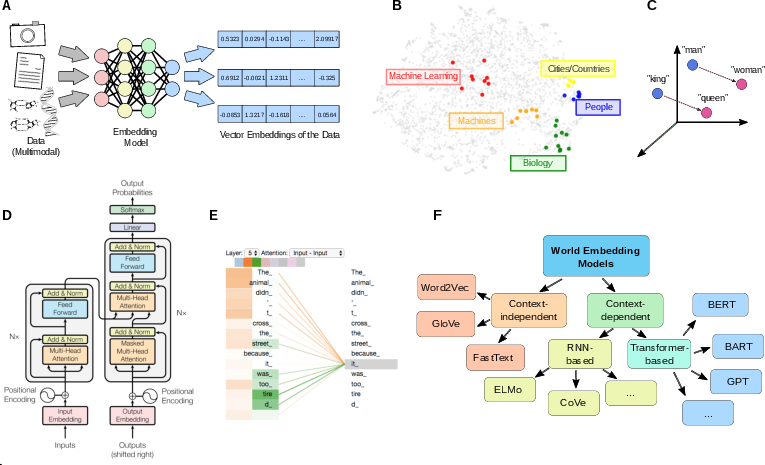
- LLMs turn words into vectors (numbers) in a space where similar meanings are close together. Attention layers keep refining these vectors to fit the current context—that’s remapping.
- These models then “walk” through that space token by token toward a likely next word—navigation driven by reducing prediction error.
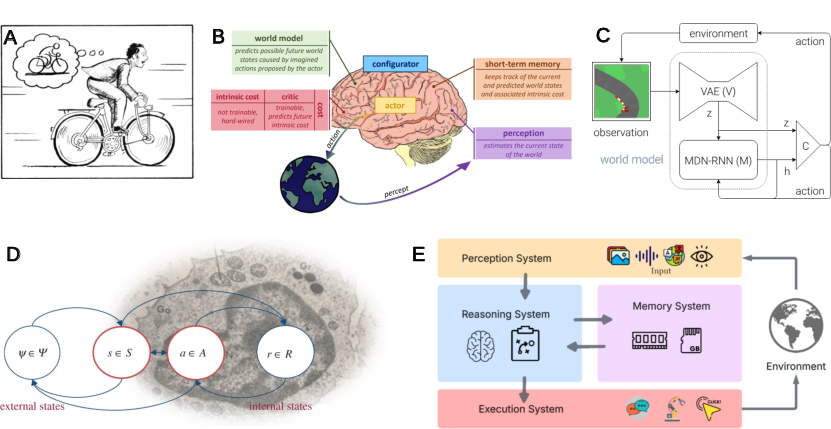
- Newer “world models” keep an internal state of the environment and try to make their predictions match reality, adjusting when they are wrong.
A helpful twist: the paper also shows that “embedding” into physical 3D space acts like a powerful constraint in biology. For example, molecules have to fit in 3D, and that limits which chemical states work. At larger scales, boundaries like cell membranes or skin help keep things coherent, so local rules add up to stable behavior.
What did they find? (Main results and why they matter)
The main claim is a simple, unifying principle:
- Intelligence—no matter the material (cells, brains, or computers)—comes from two linked processes: 1) Remapping: building and updating an internal map (embedding space). 2) Navigation: using that map to reach goals by reducing errors.
To make this concrete, the paper gives examples in both biology and AI.
Here are a few biology examples that match the principle:
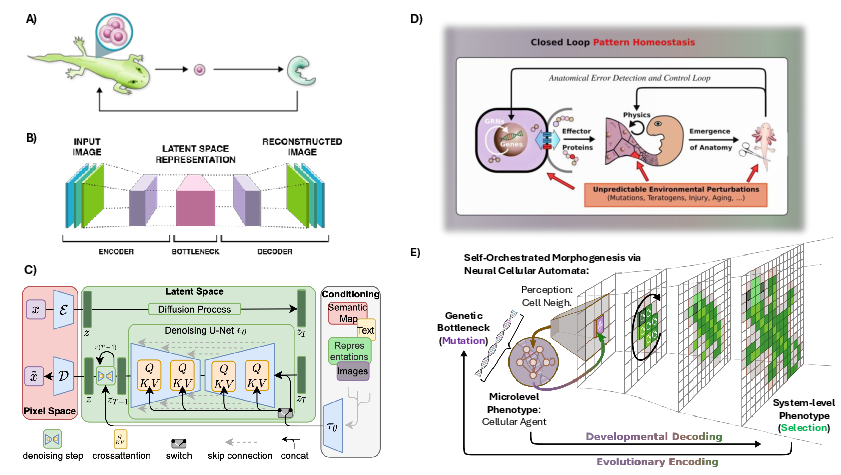
- Regeneration: Salamanders regrow limbs; frog face tissues can reorganize themselves toward a normal pattern. The cell collective “knows” the target shape and corrects errors to get there.
- Planarian flatworms: When exposed to a new chemical (barium), they change which genes are active until they reach a stable, healthy state—like “finding a safe zone” in gene-activity space.
- Immune system: It “remaps” its recognition space to match new germs by generating new antibodies.
- Brains: The hippocampus redraws “place maps” in new spaces; sensory and motor areas continually reorganize as we learn, use tools, or lose a sense.
Here are a few AI examples that match the principle:
- Transformers and embeddings: Words are mapped into a vector space; attention layers keep remapping these vectors as context changes; the model then navigates toward likely completions by reducing prediction error.
- Associative memory and high-dimensional computing: Models store patterns as “attractors” and retrieve the closest one when inputs are noisy—this is global error correction in a learned map.
- World models and agents: They keep an internal state of the world, act, check what happened, and revise their internal map to make error smaller next time.
Why this matters:
- It gives one common language for minds of many kinds.
- It explains how small, local corrections (like a cell’s local rules) can add up to big, smart outcomes (like a whole limb regenerating).
- It points to design rules for building more adaptable AI and for steering biological systems (like guiding tissue growth) by shaping their internal maps and their error signals.
What could this change? (Implications and impact)
- Better AI: Engineers can build systems that learn faster and adapt better by focusing on two things—how to create good embedding spaces (good internal maps) and how to navigate them through continual error correction. This includes smarter world models, memory, and tool use.
- Regenerative medicine and bioengineering: Doctors might learn to “nudge” tissues by adjusting their internal goals and feedback, helping bodies repair themselves more reliably.
- A unifying science of intelligence: Researchers can compare cells, animals, and algorithms using the same terms, speeding up discoveries across fields.
- Safer, more controllable systems: Understanding a system’s internal maps and its goals makes it easier to align, predict, and guide its behavior.
In one sentence: the paper argues that intelligence—everywhere—works by constantly redrawing an inner map of the world and using that map to move toward goals by fixing errors, and that this simple idea can guide how we understand, heal, and build complex living and artificial systems.
Knowledge Gaps
Below is a single, consolidated list of concrete knowledge gaps, limitations, and open questions the paper leaves unresolved; each item is phrased to be directly actionable for future research.
- Precise operationalization of “remapping,” “embedding space,” and “navigation” across substrates: what are the minimal mathematical definitions and measurable criteria that make a system count as remapping vs navigating in biology and AI?
- Falsifiability of the dual invariant claim: what specific predictions would distinguish the proposed framework from alternative theories (e.g., pure control-theoretic or purely information-theoretic accounts), and under what conditions should the invariants fail?
- Quantitative metrics of navigational competency and error minimization: how can we measure and compare “navigation efficiency” and “error-correction performance” across transcriptomic, morphogenetic, physiological, and semantic spaces?
- Empirical identification of latent spaces in living systems: what methods can infer the effective low-dimensional embeddings that biological collectives use (e.g., via topological data analysis, manifold learning, or causal discovery) and validate them experimentally?
- Mechanisms and heuristics for biological remapping at cell/tissue scales: what local rules (bioelectric, biochemical, mechanical) implement selection of effective transcriptional or morphological trajectories under novel perturbations (e.g., planarian barium resistance)?
- Timescales and stability of remapping: how fast do cells/tissues/organisms update internal maps under new contexts, and what trade-offs exist between plasticity, robustness, and catastrophic remapping (e.g., dysregulation, cancer)?
- Energy landscape deformation by higher-level systems: how can we experimentally quantify and model how organism- or tissue-level constraints reshape subsystem attractor landscapes to induce desirable local gradients?
- Active inference alignment: can morphogenesis and regeneration be cast as explicit free-energy minimization with testable generative models at tissue or organ scales; what priors and likelihoods are needed and how are Markov blankets identified empirically?
- Sheaf-theoretic coherence in biology: how can one detect and quantify coherence conditions (and their violations) in biochemical or bioelectric processes; what laboratory assays map “non-local” interference to coherence violations?
- Empirical validation of 3D spatial embedding as a constraint: what concrete mappings from chemical-potential spaces to 3D structure can be inferred or learned, and under what structural features does 3D embedding guarantee viability?
- Multi-scale embedding coherence: how can coarse-graining maps between molecular, cellular, tissue, organism, and community scales be constructed and validated so that high-resolution constraints “smear out” appropriately?
- Boundary emergence and Markov blanket dynamics: what experimental signatures indicate the formation, maintenance, and collapse of functional boundaries at multiple scales, and how do these impact cognition-like processes?
- Distinguishing learning from selection in adaptive transcriptional remapping: in cases like planarian barium adaptation, can we disentangle on-the-fly learning-like generalization from selection among pre-existing attractors?
- Disease as remapping/navigation failure: what pathologies (e.g., neurodevelopmental disorders, fibrosis, tumorigenesis) can be reframed as breakdowns of coherent embeddings or misdirected navigation, and how would interventions restore proper embeddings?
- Unifying benchmarks across spaces: what standardized tasks and datasets can assess cross-substrate navigation (transcriptomic/morphospace vs language/vision embeddings) and quantify transfer of competencies across domains?
- Attention–Hopfield equivalence in practice: can attractor dynamics and energy minimization be directly observed in transformer activations during inference, and do these dynamics predict error-correction properties and failure modes?
- Recurrent vs feedforward “cognitive depth”: what experimental paradigms reveal advantages of recurrent embedding architectures (RWKV/Mamba/xLSTM) for sustained remapping over time, and which tasks require internal states beyond transformers?
- Universal embedding geometry claim: under what training data, model capacities, and objectives do embedding spaces converge to the same geometry; what invariants (e.g., curvature, spectral properties) certify “universality” and when does it break?
- Platonic representation hypothesis: what empirical tests can confirm or refute the existence of modality-agnostic latent geometries; how can we probe whether biological and artificial agents discover the same conceptual bases?
- Disentangled vs fractured embeddings: how can one systematically induce and measure disentanglement in learned representations (e.g., via VICReg/JEPA) and tie it to generalization gains and robustness in downstream tasks?
- Agentic LLM remapping audits: what instrumentation can trace layer-wise embedding updates, tool-use interactions, and external-memory retrieval as explicit “remapping events,” and correlate them with iterative error-minimization during problem solving?
- Over-generalization vs precision trade-offs: how can we quantify when foundational models’ broad embeddings harm performance on narrow tasks, and what regularization or curriculum strategies improve precision without losing generality?
- Cross-modal embedding alignment in biology: do cells/tissues maintain consistent embeddings across bioelectric, mechanical, and biochemical modalities; what experiments test multimodal integration and remapping under perturbation?
- Intervention protocols to steer biological embeddings: what concrete bioelectric, mechanical, or pharmacological strategies can reshape latent maps to guide regeneration (e.g., target morphologies), and how are success criteria defined?
- Causality and necessity/sufficiency: are remapping and navigation via error minimization necessary and sufficient for intelligence-like behavior; can we construct counter-examples (systems that navigate without remapping or vice versa) to delimit the framework?
- Resource costs and scaling limits: what are computational/biophysical costs of maintaining and remapping embeddings at scale; where do bandwidth, noise, or energy constraints limit the dual invariant in biology and AI?
- Formal mapping from biological to AI embeddings: can we design cross-domain translation tools (e.g., category-theoretic functors or learned encoders) that map biological latent spaces to artificial ones and enable simulation or control across substrates?
Glossary
- Active inference: A theoretical framework where agents minimize prediction errors by aligning internal models with sensory inputs to maintain homeostasis. "Active inference \cite{friston2010}, for example, can only be maintained in a reasonably familiar and hence reasonably predictable environment;"
- Associative memories: Systems that retrieve stored patterns by converging to attractor states closest to an input, typical of Hopfield networks. "Hopfield networks retrieve the closest stored pattern and thus act as associative memories \cite{hopfield1982} of their training data."
- Attractor landscapes: Energy or state-space structures with basins (attractors) toward which system dynamics converge. "The principles observed in biological systems -- navigation of attractor landscapes, local error correction that scales to system-level outcomes, and nested embeddings that support higher-level organization --"
- Attractor regions: Specific areas in a state space toward which the system tends to evolve and stabilize. "The Fields-Levin framework defines these phenomena as navigation in morphospace toward specific attractor regions \cite{fields2022competency}"
- Bioelectric networks: Cell-level electrical signaling systems coordinating collective behaviors and morphogenesis. "developmental biology reveals that familiar bioelectric networks play key roles in coordinating cellular activities toward specific anatomical endpoints, long before nerves appear during morphogenesis,"
- Brain-machine interfaces: Technologies mapping neural activity to external device control in real time. "brain-machine interfaces demonstrate remapping in real-time, as motor cortex represents external device control, mapping internal neural activity onto external robotic movements \cite{lebedev2006brain}."
- Cognitive light cones: Conceptual horizons defining the spatiotemporal scope of an agent’s goals and influence. "binding competent subunits (e.g., cells) into collective intelligences that operate in spaces and with spatio-temporal goal horizons (cognitive light cones) much larger than those of their parts"
- Coherence conditions: Constraints ensuring consistent data assignments across overlapping regions in sheaf-theoretic embeddings. "Violations of the coherence conditions correspond to violations of the Kolmogorov axioms, e.g. local additivity of probabilities."
- Continuous Hopfield network: An associative memory model with continuous states and attractor dynamics; self-attention can be equivalent to it. "the self-attention mechanism in transformers has been shown to be mathematically equivalent to a form of continuous Hopfield network."
- Coarse-graining: Mapping high-resolution descriptions to lower-resolution ones while preserving essential structure. "Embeddings of the same data at different resolutions are coherent if, but only if, the higher-resolution embeddings coarse-grain to the lower resolution embeddings via some function that does not depend on the embedding-space coordinates."
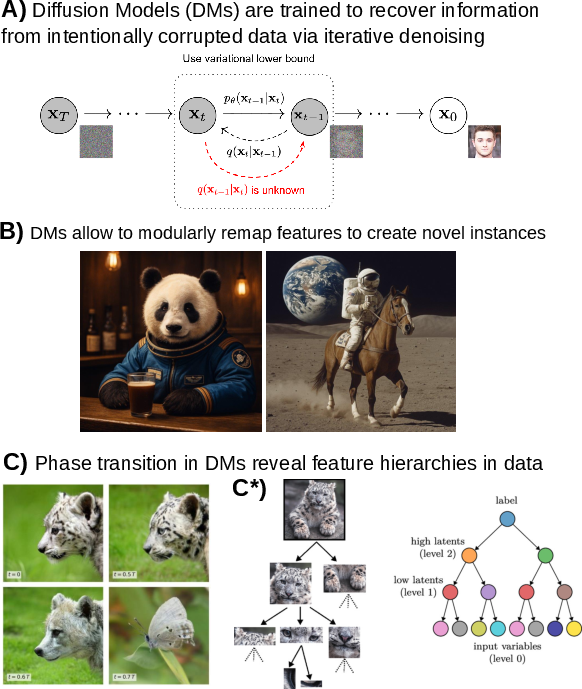
- Diffusion models: Generative models that iteratively denoise data to sample from complex distributions. "Modern AI systems, including transformers, diffusion models, and neural cellular automata enact analogous processes by remapping data into latent embeddings and refining them iteratively through contextualization."
- Disentangled representations: Latent features organized into independent, modular factors that generalize better. "neuroevolution techniques biased towards open-ended search tend to find modular \"disentangled\" representations that generalize well to further downstream tasks."
- Domain walls: Emergent boundaries in coarse-grained descriptions that constrain diffusion or dynamics. "High-resolution constraints that do not \"smear out\" at lower resolution may appear as domain walls or mesoscale structures at low resolution."
- Echolocation: Biological sonar used by bats; its evolution required auditory remapping into spatial maps. "The development of bat echolocation, for instance, necessitated remapping auditory processing in order to produce intricate internal spatial maps from external acoustic reflections"
- Embedding space: A structured vector space where complex data are mapped to preserve semantic or relational properties. "We argue that this dual principle -- remapping and navigation of embedding spaces via iterative error minimization -- constitutes a substrate-independent invariant of cognition."
- Energy landscapes: State-space surfaces guiding system dynamics via gradients and attractors. "Higher-level systems deform the energy landscapes for their subsystems, allowing lower-level components to follow local gradients that achieve goals beneficial to the higher-level system"
- Fractured entangled representation (FER): Disorganized, conceptually entangled latent structures from gradient-based training. "Without particular regularization techniques, gradient-based methods often suffer from disorganized \"entangled\" embedding structures, termed fractured entangled representation (FER)"
- Galilean metric: The classical spatial metric structure of 3D space used to constrain embeddings. "bounded volumes in 3D space with a simple Galilean metric."
- Gene regulatory networks (GRNs): Interacting gene control systems that can learn and store memory at the molecular level. "Even below cell level, gene regulatory networks are capable of various kinds of learning and memory"
- Generative models: Models that learn data distributions to synthesize or predict new samples. "Instead of relying on static representations, modern AI increasingly relies on generative models \cite{Foster2023generative} --"
- Glomeruli: Neural clusters in the olfactory bulb receiving inputs from specific receptor types. "olfactory receptors located throughout the nasal cavity connect to specific neural clusters called glomeruli in the olfactory bulb,"
- Hyperdimensional computing (HDC): Computing paradigm using very high-dimensional vectors for robust, holographic representations. "The related concept of hyperdimensional computing (HDC) is built on the principle that information can be stored and manipulated in extremely high-dimensional spaces -- often 10,000+ dimensions --"
- In-dream training: Training world models using internally generated simulations to improve performance. "potentially improving learning performance through \"in-dream\" training in several iterations of the model's own imagination"
- JEPA: Joint-Embedding Predictive Architecture for self-supervised learning imposing variance and decorrelation constraints. "Promising self-supervised techniques like VICReg \cite{Bardes2022} or JEPA \cite{Assran2023JEPA, assran2025vjepa2selfsupervisedvideo} impose constraints on variance, invariance, and conceptual decorrelation"
- Kolmogorov axioms: Foundational probability rules; their violation indicates non-classical context dependence. "Violations of the coherence conditions correspond to violations of the Kolmogorov axioms, e.g. local additivity of probabilities."
- Latent space: Lower-dimensional space capturing essential structure via an embedding from a high-dimensional parent space. "Embedding a high-dimensional parent space into a lower-dimensional latent space via a structure-preserving (i.e. non-trivial) map "
- LLM agents: LLM systems augmented with memory, tools, and control loops for autonomous task navigation. "build LLM agents capable of actively navigating custom problem domains~\cite{Wang2024SurveyLLMAgents, Park2023GenerativeAgents}."
- Mamba: A recurrent sequence model architecture maintaining internal embedding states over time. "RWKV \cite{peng2023rwkv}, Mamba \cite{gu2024mamba}, or xLSTM \cite{beck2025xlstm} architectures, to name but a few, maintain internal embedding states"
- Markov blanket: Boundary separating internal and external states, enabling localized inference and regulation. "these boundaries are Markov blankets \cite{pearl:88};"
- Markov blanket collapse: Breakdown of boundary-mediated inference under extreme environmental novelty. "completely unfamiliar environments -- e.g. environments involving radically different chemical potentials, EM fields, particle fluxes, or mechanical forces -- tend to induce Markov blanket collapse, generative-model dysregulation, and death."
- Mesoscale structures: Intermediate-scale emergent formations arising from fine-scale constraints. "may appear as domain walls or mesoscale structures at low resolution."
- Morphogenesis: Biological process shaping organisms during development via coordinated cellular activity. "familiar bioelectric networks play key roles in coordinating cellular activities toward specific anatomical endpoints, long before nerves appear during morphogenesis,"
- Morphospace: Abstract space of possible anatomical configurations navigated during development and regeneration. "During development and regeneration, cellular collectives navigate morphospace with high robustness,"
- Multimodal models: AI models integrating multiple modalities (e.g., text, vision) into a unified representational space. "multimodal models are navigating a merged space of different modalities \cite{baltruvsaitis2018multimodal}, etc."
- Nested embeddings: Hierarchically organized embeddings across scales that support higher-level structure and cognition. "nested embeddings that support higher-level organization -- exemplify scale-free cognitive dynamics"
- Neuroevolution: Evolutionary optimization of neural architectures emphasizing open-ended search and modular representations. "representations learned via open-ended neuroevolution paradigms show superior embedding representations as opposed to gradient-based methods"
- Neural cellular automata: Learnable, locally-updating grid-based models used for generative pattern formation and adaptation. "Modern AI systems, including transformers, diffusion models, and neural cellular automata enact analogous processes"
- Platonic representation hypothesis: The idea that learned embeddings converge to a universal geometry reflecting abstract forms. "raising fascinating questions about the Platonic representation hypothesis \cite{Huh2024PlatonicRepresentation, Levin2025IngressingMinds}."
- Retinotopic maps: Spatial neural mappings preserving retinal layout in visual cortices. "retinotopic maps in visual processing areas preserve spatial relationships from the retina"
- Retrieval-augmented generation: Technique augmenting LLMs with external memory to ground and improve outputs. "retrieval-augmented generation \cite{Lewis2020RAG, Borgeaud2022RAG},"
- RWKV: A recurrent architecture blending RNN-like state with attention-like mechanisms for efficient sequence modeling. "RWKV \cite{peng2023rwkv}, Mamba \cite{gu2024mamba}, or xLSTM \cite{beck2025xlstm} architectures, to name but a few, maintain internal embedding states"
- Sheaf theory: A category-theoretic framework for coherently attaching structured data to spaces across overlapping regions. "The idea of embeddings as imposing constraints can be made precise in the language of sheaf theory."
- Signal transduction: Cellular biochemical signaling pathways translating external cues into internal actions. "This is true not just of biochemical, e.g. metabolic or signal transduction, bioelectric, and mechanical processes,"
- Tonotopic maps: Auditory cortical mappings organized by sound frequency gradients. "neurons in tonotopic maps are organized following sound frequency gradients, with neighboring cells responding to similar frequencies"
- Transformer attention heads: Parallel attention mechanisms in transformers specialized for different data relationships. "Moreover, transformers maintain several attention heads."
- Transcriptional landscape: The global pattern of gene expression states navigated or remapped in response to context. "The already described planarian's transcriptional adaptation to barium demonstrates remapping of its internal transcriptional landscape"
- Variational free-energy principle: The proposition that self-organizing systems minimize an upper bound on surprise to remain viable. "this principle is formalized through the variational free-energy principle \cite{friston2010},"
- VICReg: Self-supervised method enforcing variance, invariance, and decorrelation to learn robust representations. "Promising self-supervised techniques like VICReg \cite{Bardes2022} or JEPA \cite{Assran2023JEPA, assran2025vjepa2selfsupervisedvideo} impose constraints on variance, invariance, and conceptual decorrelation"
- World models: Architectures that learn internal simulations of environments to plan, act, and minimize prediction errors. "World models \cite{ha2018worldmodels, dawid2023AutonomousMachineLearning, Friston2021worldmodel}, for instance, embody this principle probably most explicitly"
- xLSTM: An extended LSTM variant maintaining rich internal states to capture long- and short-term dependencies. "RWKV \cite{peng2023rwkv}, Mamba \cite{gu2024mamba}, or xLSTM \cite{beck2025xlstm} architectures, to name but a few, maintain internal embedding states"
Practical Applications
Immediate Applications
Below are actionable use cases that can be deployed now, derived from the paper’s dual invariant of cognition—remapping and navigation of embedding spaces via iterative error minimization.
Industry
- Embedding observability (“EmbOps”) for ML pipelines
- Sector(s): software/AI
- Tools/products/workflows: embedding-health dashboards with metrics inspired by VICReg/JEPA (variance, invariance, decorrelation), disentanglement indices, attention-as-Hopfield energy diagnostics, drift/coherence checks across training and inference
- Assumptions/dependencies: access to model internals and training data; standardized metrics; organizational MLOps maturity
- RAG with active-inference loops for enterprise AI
- Sector(s): software/enterprise IT
- Tools/products/workflows: agentic LLMs that iteratively minimize retrieval/answer error using planning, tool-use, self-reflection (Reflexion), and external memory; plan–act–evaluate cycles that remap context embeddings at each step
- Assumptions/dependencies: reliable knowledge bases, tool APIs, guardrails, audit logs for reasoning steps
- Hyperdimensional encoding for robust edge analytics
- Sector(s): robotics, IoT, embedded systems
- Tools/products/workflows: HDC libraries/hardware for sensor fusion and anomaly detection; superposition and binding operations to robustly represent multimodal data on-device
- Assumptions/dependencies: hardware support or optimized software kernels; task-specific benchmarking; energy/latency constraints
- Error-minimizing robotic controllers (fast adaptation)
- Sector(s): manufacturing, logistics, service robotics
- Tools/products/workflows: controllers that minimize deviation from task goals via online prediction–correction loops; model-predictive control augmented by learned latent states
- Assumptions/dependencies: high-quality sensing; safety verification; ability to update policies without downtime
- Multimodal embedding adapters (unified representation layer)
- Sector(s): AI platforms, developer tools
- Tools/products/workflows: CLIP-like adapters and universal embedding space alignment (per recent universal geometry findings) to interoperate CV/ASR/NLP models
- Assumptions/dependencies: access to foundation model embeddings; calibration datasets for cross-modal mapping
- Neuroevolution add-ons for representation learning
- Sector(s): AI tooling
- Tools/products/workflows: training services that evolve disentangled, modular representations (avoiding FER) before/alongside gradient fine-tuning to improve downstream generalization
- Assumptions/dependencies: compute budget; reproducibility and stability of evolutionary runs; comparison protocols with SGD baselines
Academia
- Morphospace navigation assays in regenerative biology
- Sector(s): developmental biology, regenerative medicine
- Tools/products/workflows: experiments that define target morphologies as attractors and measure error-minimization trajectories under bioelectric/chemical perturbations; ion-channel modulation screens (e.g., barium analogs, gap junction drugs)
- Assumptions/dependencies: suitable model organisms (planaria, amphibians); ethical approvals; high-resolution morphometrics and electrophysiology
- Sheaf-theoretic coherence analysis for complex datasets
- Sector(s): computational biology, complex systems, data science
- Tools/products/workflows: open-source library to test multi-scale coherence (compatibility across overlapping regions, emergent constraints) in biological and AI representations
- Assumptions/dependencies: expertise in category theory; translation of theory to interpretable diagnostics
- Cross-scale cognition curriculum and labs
- Sector(s): education
- Tools/products/workflows: course modules, interactive notebooks demonstrating remapping/navigation across cells, brains, and AI; hands-on attention/Hopfield labs
- Assumptions/dependencies: institutional buy-in; access to datasets and compute for teaching
Policy
- Representation transparency and robustness audits
- Sector(s): AI governance
- Tools/products/workflows: standards for embedding audit trails (disentanglement, drift, fairness via representation geometry), attention-head role reports, retrieval traces
- Assumptions/dependencies: regulator engagement; industry cooperation; privacy-preserving audit mechanisms
- Guardrails for agentic LLMs (reasoning logs, tool-use safety)
- Sector(s): AI safety/compliance
- Tools/products/workflows: requirements for logs of iterative self-correction, tool invocations, and boundary conditions; conformance testing for error-minimizing control policies
- Assumptions/dependencies: standardized testbeds; interoperability across vendors; incident reporting frameworks
Daily Life
- Personal planning assistants with iterative self-correction
- Sector(s): consumer software
- Tools/products/workflows: LLM-based agents that remap context embeddings as plans unfold; plan–execute–reflect loops to minimize deviation from user goals (calendar/tasks/travel)
- Assumptions/dependencies: data privacy controls; reliable integrations; transparent user feedback channels
- Adaptive homeostatic coaching in wearables
- Sector(s): consumer health
- Tools/products/workflows: trackers that learn personalized embeddings of physiology and iteratively minimize deviations from healthy baselines (sleep, glucose variability, HRV)
- Assumptions/dependencies: medical-grade sensing; personalized models; regulatory alignment for health claims
Long-Term Applications
Below are applications that will require further research, scaling, validation, or standardization before widespread deployment.
Industry
- Morphological compilers for regenerative medicine
- Sector(s): biotech/healthcare
- Tools/products/workflows: platforms that translate target anatomies into spatiotemporal bioelectric/chemical stimuli, steering cell collectives through morphospace via distributed error correction
- Assumptions/dependencies: validated maps from stimuli to anatomical attractors; long-term safety and efficacy; translational pipeline (animal → human trials)
- Universal embedding exchange standard
- Sector(s): AI platforms/standards
- Tools/products/workflows: cross-vendor protocols for the “universal geometry” of embeddings; adapters and conformance suites enabling seamless multimodal interoperability
- Assumptions/dependencies: consensus on metrics and schemas; governance bodies; IP/licensing resolution
- World-model powered generalist autonomy
- Sector(s): robotics, automotive, smart devices
- Tools/products/workflows: agents that maintain internal simulations and minimize prediction error for planning and imagination (“in-dream” training); robust nested boundaries for safe deployment
- Assumptions/dependencies: scalable, sample-efficient world models; formal safety guarantees; on-device compute and memory
Academia
- Biohybrid collective intelligences
- Sector(s): bioengineering/robotics
- Tools/products/workflows: xenobot-like systems with active-inference controllers that navigate nested embeddings (biophysical + algorithmic) to achieve complex tasks
- Assumptions/dependencies: ethical frameworks; reliable interfaces between living tissue and controllers; stability over time
- Multi-scale coherence verification (cells-to-systems)
- Sector(s): complex systems science
- Tools/products/workflows: sheaf-based toolkits to certify that local constraints coarse-grain to global behaviors (from GRNs → tissues → organisms → ecosystems)
- Assumptions/dependencies: high-resolution longitudinal data; computational scalability; interpretability to practitioners
- Platonic representation program
- Sector(s): AI theory/philosophy of representation
- Tools/products/workflows: comparative analyses across foundation models to test universality claims in embedding geometry and implications for multimodal integration
- Assumptions/dependencies: diverse large-scale model access; shared evaluation corpora; reproducible measurement protocols
Policy
- Multiscale competency governance
- Sector(s): bioethics/AI ethics
- Tools/products/workflows: frameworks for attributing limited agency/rights across cellular collectives, organisms, and AI agents; consent and welfare protocols for unconventional minds
- Assumptions/dependencies: societal consensus-building; cross-disciplinary ethics boards; empirical thresholds for competency
- Safety certification via error-minimization controllers
- Sector(s): autonomous systems regulation
- Tools/products/workflows: standardized proofs that agents maintain boundaries and minimize deviations from safe envelopes; scenario-based certifications
- Assumptions/dependencies: formal verification methods; regulatory infrastructure; incident forensics
- Privacy and provenance for universal embeddings
- Sector(s): data governance
- Tools/products/workflows: policies ensuring embeddings do not leak sensitive attributes; provenance tracking for remapped latent spaces used across applications
- Assumptions/dependencies: technical privacy guarantees (e.g., DP for embeddings); enforceable audit trails; global regulatory harmonization
Daily Life
- Neuroadaptive BCIs and prosthetics
- Sector(s): healthcare/assistive tech
- Tools/products/workflows: interfaces that remap motor/intention embeddings to device control, minimizing user–device error over time for seamless operation
- Assumptions/dependencies: clinical validation; individualized calibration; robust long-term biocompatibility
- Personalized immune engineering
- Sector(s): precision medicine
- Tools/products/workflows: therapies that guide VDJ remapping to navigate immune recognition space toward target antigens with minimal off-target risks
- Assumptions/dependencies: comprehensive antigen mapping; safety monitoring; regulatory approvals
- Active-inference smart homes and micro-grids
- Sector(s): energy/smart cities
- Tools/products/workflows: nested controllers that maintain stable local environments (thermal, acoustic, energy) via distributed error minimization and multi-modal sensing
- Assumptions/dependencies: interoperable IoT standards; cybersecurity; resilience under failure modes
Cross-Sector (Finance and Energy)
- Distributed error-minimization for grid stability
- Sector(s): energy
- Tools/products/workflows: controllers that treat demand/supply states as latent embeddings and iteratively reduce deviation from stable attractors at multiple scales (device → feeder → grid)
- Assumptions/dependencies: real-time telemetry; regulatory permission; formal stability analysis
- Hyperdimensional market agents for robust risk management
- Sector(s): finance
- Tools/products/workflows: high-dimensional embeddings of market microstructure and macro signals, enabling associative recall of stress patterns and rapid error-corrective hedging
- Assumptions/dependencies: high-quality data pipes; compliance with trading regulations; robust backtesting under regime shifts
Collections
Sign up for free to add this paper to one or more collections.