- The paper presents a search engine that extracts sentences stating scientific challenges and directions from extensive literature corpora.
- It employs fine-tuned domain transformer models, including PubMedBERT, achieving F1 scores up to 0.783 for challenge extraction.
- User studies and cross-domain evaluations demonstrate significant improvements over traditional systems for rapid research discovery.
Search-Centric Extraction of Scientific Challenges and Directions
Motivation and Problem Definition
Efficient identification of open scientific challenges and promising directions is crucial for the acceleration of research progress and mitigation of redundant efforts—particularly acute in hyper-prolific domains such as COVID-19. Traditional literature search systems fail to directly index or surface statements of limitations, open problems, or research hypotheses, causing missed opportunities for rapid knowledge discovery and evidence-based clinical decision-making.
The paper introduces and formalizes the task of extracting and indexing challenge/direction statements at the sentence level from scientific literature, with an initial focus on the interdisciplinary CORD-19 COVID-19 corpus. The authors define fine-grained semantic categories: “challenge” (problems, limitations, gaps) and “direction” (suggested future work, hypotheses), decoupled from explicit structural cues like section headers.
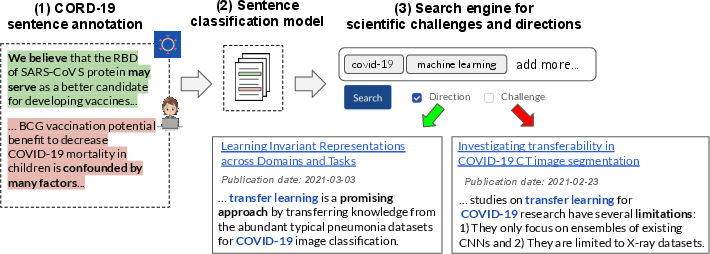
Figure 1: Overview of the extraction system, including expert annotation, model training, and deployment as a searchable index for the COVID-19 literature.
Corpus Creation and Expert Annotation
A dataset of 2,894 sentences with surrounding context was sampled from full-length CORD-19 papers across 1,786 sources. Emphasizing recall, sentences were upsampled by querying 280 weakly indicative keywords but also included a large stratum of keyword-negative samples. Annotation guidelines and training were employed for four domain experts, with a strong inter-annotator micro-F1 of 85% (challenges) and 88% (directions).
Multi-labeling captures the frequent entanglement of challenges and directions in scientific writing. The label distributions are imbalanced, with more frequent negatives and a nontrivial intersection category (challenge+direction).
Modeling Approach
Baselines and LLM Fine-Tuning
A diverse range of baselines was considered:
- Keyword and Sentiment: High recall/poor precision, revealing the inadequacy of lexicon matching and polarity heuristics for this task.
- Zero-shot NLI (BART-MNLI): Leveraged multiple class-label variants to improve detection but still underperformed compared to supervised approaches.
- Domain-pretrained Transformers: PubMedBERT, SciBERT, and (for comparison) RoBERTa-large were fine-tuned in a multi-label setup, with PubMedBERT yielding the best single-model results (F1: 0.770/0.766 for challenge/direction).
Context-aware modeling was studied: Hierarchical Attention (HAN) could trade precision for recall but did not yield superior overall F1. A "Slice-Combine" ensembling method, integrating independent predictions from various context input combinations at training and test time, produced the best observed F1 (0.783/0.773).
Error Analysis and Contextual Effects
False positives often reflected domain ambiguity (e.g., sentences describing general conditions or ambiguous problem status without domain knowledge). False negatives happened with implicit challenges or directions not overtly stated. Context can sometimes resolve these ambiguities but also insert noise, motivating the nuanced Slice-Combine strategy rather than naive context concatenation.
Model Generalization and Downstream Evaluation
High-confidence classifier predictions maintain >96% mean average precision (MAP) for both categories in CORD-19 and demonstrate robust zero-shot transfer to the S2ORC general biomedical corpus and the SciRex AI domain corpus (95–98% MAP), substantiating generalization beyond COVID-19 and biomedicine.
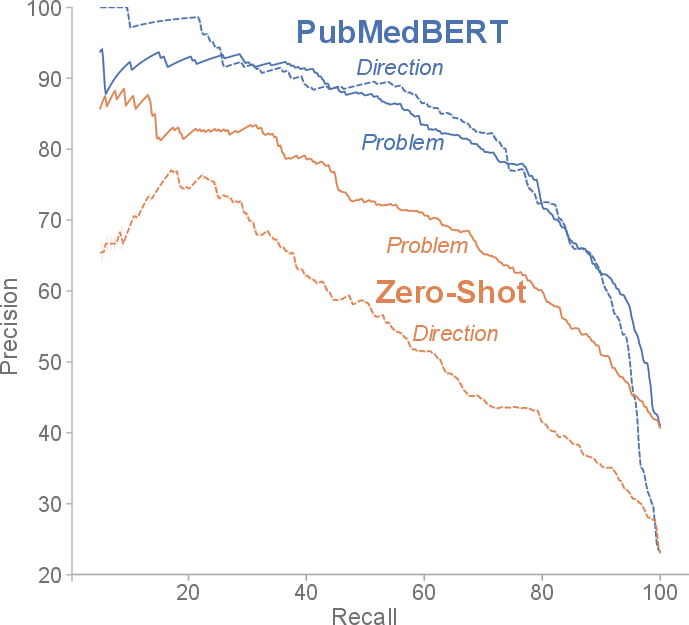
Figure 2: Precision/recall profile for PubMedBERT and the zero-shot baseline; PubMedBERT maintains high precision at practically relevant recall levels.
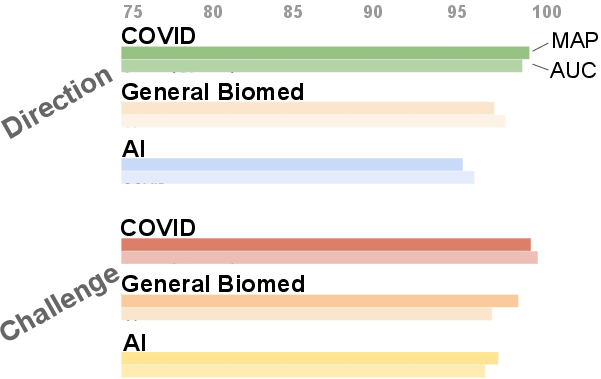
Figure 3: Domain transfer results: accuracy for challenge/direction extraction is high even in general biomedical and AI corpora, confirming cross-domain signal generality.
Search Engine Deployment and User Studies
The trained model indexed 2.2M sentences (high-confidence, with context) from 550K COVID-19 papers. Biomedical entity extraction (SciSpacy + MeSH linking) and faceted search interfaces were implemented for targeted retrieval of, e.g., “AI + diagnosis + pneumonia” challenge statements.
Utility was quantified via structured user studies:
- Study 1 (N=10 scientists): For over 70 distinct queries, participants found, on average, 4.46/6.43 challenge/direction instances per query using the new search engine versus 2.24/2.03 for PubMed (p < 0.002 for both).
- Study 2 (N=9 MDs): For controlled medical queries, satisfaction and perceived utility ratings (PSSUQ) were higher for the proposed system across all dimensions (overall, search, and utility—see below).
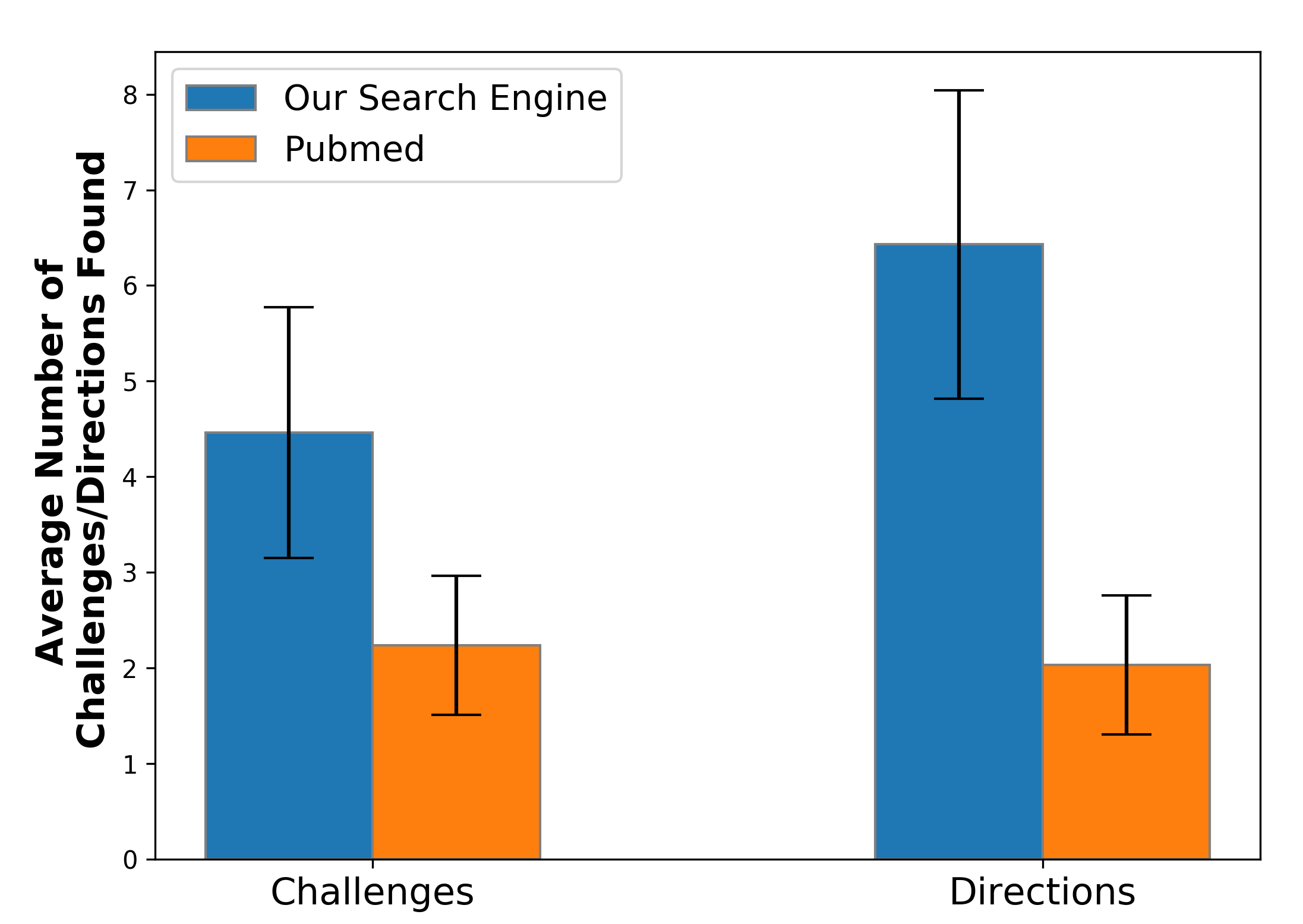
Figure 4: User study demonstrating that the specialized search engine uncovers more actionable challenge and direction statements per query than PubMed.
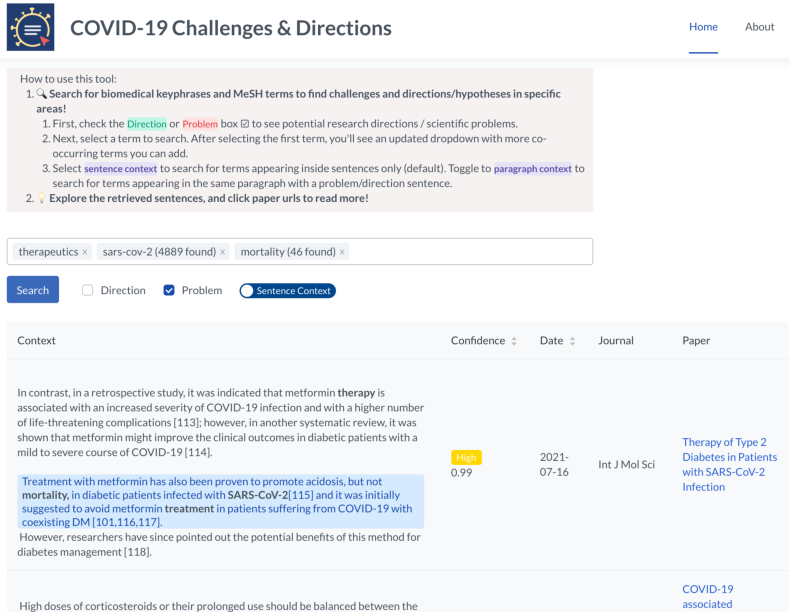
Figure 5: Screen capture of the deployed search interface central to the user experience.
Implications and Future Directions
This work establishes that scientific challenge/direction detection is feasible at scale via expert-annotated data and fine-tuned domain transformer models. The system demonstrates robust generalization, enabling extension to arbitrary domains affected by research volume overload. User studies with professional scientists and clinicians validate significant gains over generic search baselines for research ideation, literature review, and evidence synthesis.
On the practical axis, integrating such extraction models with knowledge synthesis and analytic tools (e.g., mechanistic relation graphs, hypothesis ranking, cross-discipline analogical retrieval) could accelerate targeted discovery and enable real-time literature triage during emergent crises (future pandemics, fast-evolving technological areas).
Theoretically, robust, multi-label challenge/direction detection transcends section-level rhetorical structure approaches, enabling automated surfacing of uncertainty and speculation, which are core to scientific progress but often overlooked in closed-loop, factoid-centric IE pipelines.
Conclusion
The paper offers a validated pipeline for automatic extraction and search of challenge and direction statements, demonstrating superior results to established scientific search systems in critical use cases. The approach is domain-agnostic, deployable, and directly bridges gaps between NLP, scientific information extraction, and researcher workflows, laying the groundwork for the next generation of search engines oriented toward unsolved scientific problems and opportunities (2108.13751).




