- The paper demonstrates that domain fine-tuning yields a 6.8 percentage point accuracy boost over general models for USMLE-style MCQA on 4B-parameter LLMs.
- It details a controlled 2x2 experimental framework comparing DAFT and RAG, revealing that external retrieval does not statistically enhance accuracy.
- The findings suggest that embedding domain knowledge directly into model weights is more effective than incorporating external context for small, deployable medical QA models.
Domain Fine-Tuning Versus Retrieval-Augmented Generation for Medical MCQA at the 4B Scale
This work investigates, under strict experimental control, the relative efficacy of domain-adaptive fine-tuning (DAFT) versus retrieval-augmented generation (RAG) for medical multiple-choice QA in small, open-weight, 4B-parameter LLMs. The core question is which intervention—adapting model weights to the medical domain or providing external domain information at inference—provides greater improvements in accuracy for USMLE-style questions at a fixed parameter budget and deployment profile.
The authors select two backbones: Gemma3-4B (general-purpose, instruction-tuned) and MedGemma-4B (domain-adapted via continued pretraining on medical data). Both are 4-bit quantized and served via Ollama for deployment relevance. RAG is implemented with a ChromaDB-based pipeline retrieving from MedMCQA explanation fields, with careful filtering to avoid answer leakage and tuned hybrid reranking. A 2×2 design with three repetitions per item evaluates all combinations of backbone (general, domain-tuned) and context paradigm (question-only, question + retrieved passages) on the MedQA-USMLE 4-option test split.
Accuracy and Statistical Analysis
Majority-vote accuracy reveals a substantial effect for domain fine-tuning and no significant benefit from RAG, irrespective of the backbone. The gap in accuracy between the generalist and domain-adapted models is +6.8 percentage points (46.4% for Gemma3-4B vs. 53.3% for MedGemma-4B), with a McNemar p-value < 10−4, indicating a highly significant improvement derived solely from encoding domain knowledge into model weights.
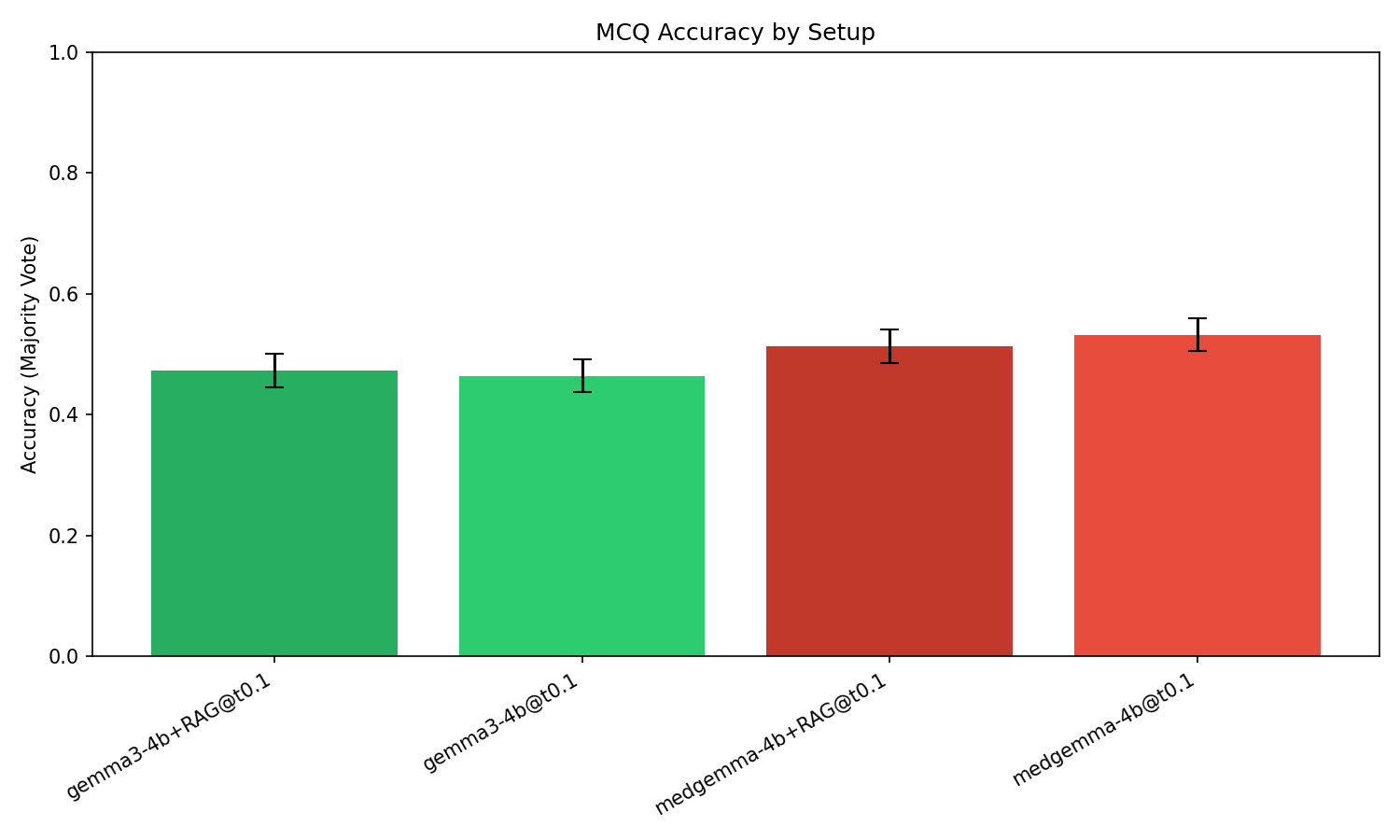
Figure 1: Majority-vote accuracy with 95% confidence intervals highlights the significant improvement from domain fine-tuning (+6.8 pp), while the introduction of RAG does not yield a statistically meaningful change.
For RAG, adding retrieved textbook-style explanations (MedMCQA) does not produce statistically significant accuracy gains for either Gemma3-4B (p=0.56) or MedGemma-4B (−1.9 pp, p=0.16). All pairwise McNemar significance tests confirm only backbone switch comparisons are significant; toggling RAG does not change accuracy reliably.
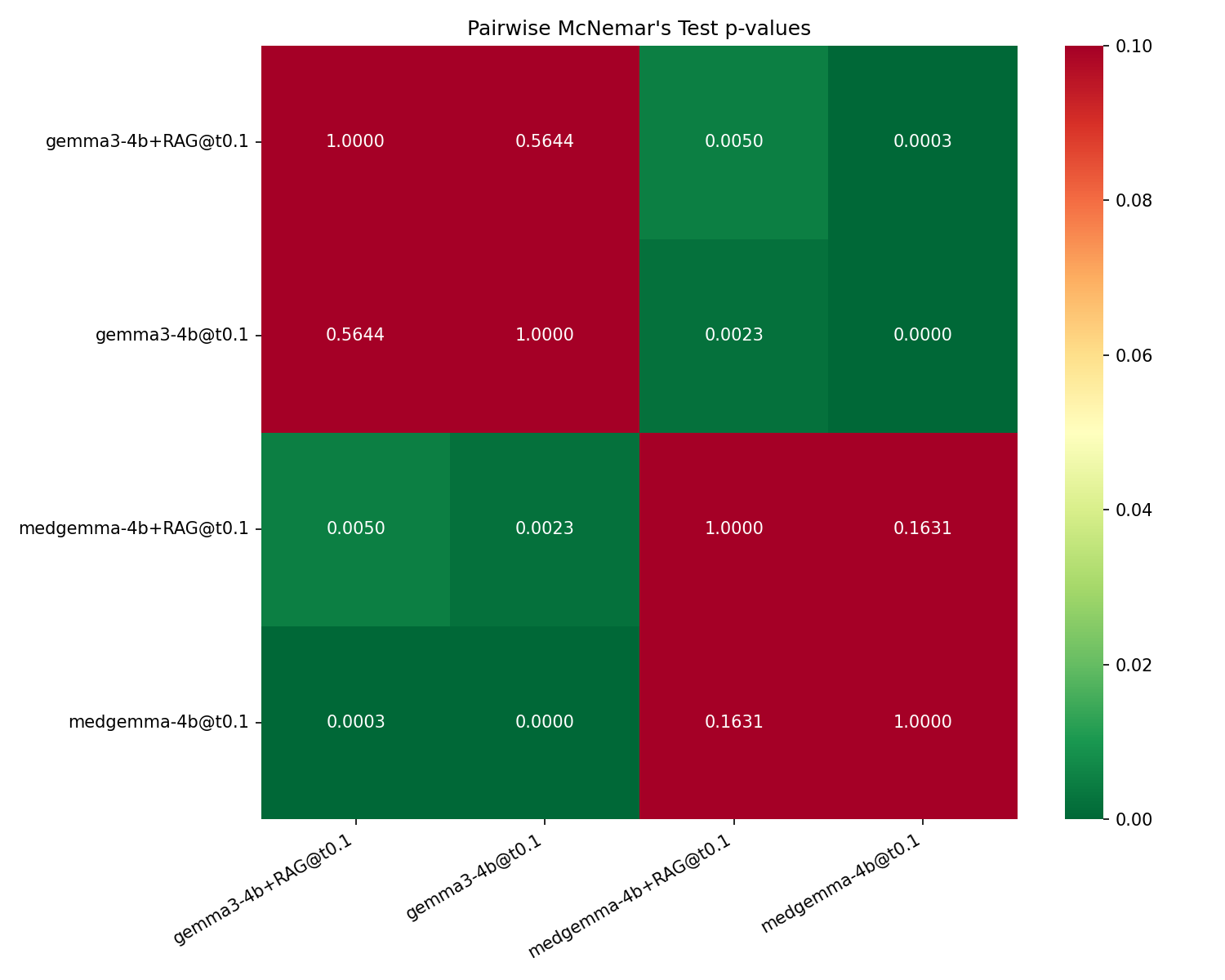
Figure 2: Pairwise McNemar p-values for the four experimental setups, where non-significant cells are confined to within-backbone RAG toggles.
Consistency and Output Variability
All four configurations exhibit extremely high within-setup answer consistency (≥0.99), and the parse-fail rate is negligible. Variability caused by stochastic decoding at T=0.1 is therefore minimal, and the effect sizes/descriptive statistics are robust to aggregation choice.
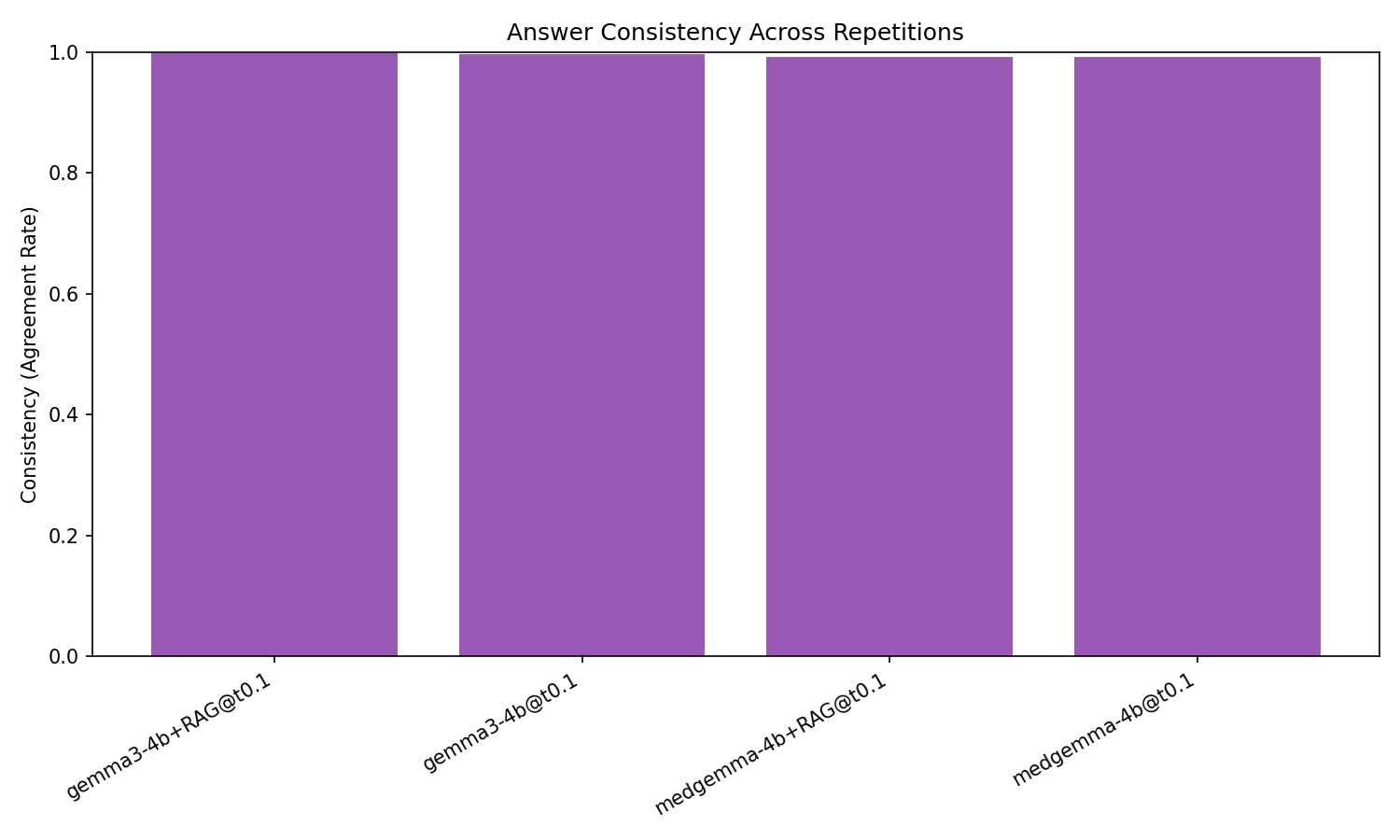
Figure 3: All evaluated configurations demonstrate nearly perfect output consistency across decoding repetitions, validating aggregation reliability.
Interpretation and Potential Mechanisms
The experimental evidence demonstrates that, at the 4B scale, DAFT on medical corpora is the substantially more effective engineering intervention for boosting USMLE-style MCQA accuracy. Several plausible explanations exist for the lack of RAG effect:
- Task intrinsic reasoning demands: USMLE items often require complex pattern recognition, integration of multiple cues, and chaining of clinical reasoning, rather than recall of explicit facts likely to be surfaced by retrieval.
- Corpus mismatch and non-authoritativeness: The MedMCQA explanations, while broad, may not offer highly aligned or authoritative context for the MedQA benchmark, limiting their value when injected.
- Model capacity constraints: 4B-parameter models may lack the representational capacity for effective late fusion of multiple retrieved passages, especially where deep reasoning and grounding are required.
- Domain redundancy/interference: For MedGemma-4B, in-weights knowledge absorbed during DAFT likely overlaps with the content retrieved, possibly leading to interference or confusion rather than synergy.
Summarily, domain knowledge encoded directly into model parameters is more impactful than context-window augmentation from an external corpus for small LLMs on this MCQA task.
Practical and Theoretical Implications
Practitioners developing deployable medical QA systems at this scale should prioritize high-quality DAFT/backbone selection over RAG pipeline engineering when resources are constrained. While RAG does not appear actively harmful, it lacks efficacy parity with DAFT given fixed context length and model capacity. This shifts immediate model-stack design priorities for local clinical and educational deployments.
From a methodological perspective, the results argue for scale-awareness: RAG-boosted architectures that improve performance for large (≥70B) backbones—where context integration and retrieval grounding are more tractable—may not transfer the same benefits to smaller, cost-sensitive models. These findings recommend target-task, scale, and context-relevancy-alignment-aware evaluation for the design of medical and domain-specific LLM systems.
Limitations and Future Directions
The investigation is limited by its focus on a single benchmark (MedQA-USMLE), one retrieval corpus, and the 4B parameter/4-bit quantization regime. Results cannot be uncritically extrapolated to settings with verbatim-answerable open-book tasks, higher-capacity models, or more authoritative retrieval corpora. Further exploration could address:
- The incremental effect of RAG for larger open-weight or API-scale backbones
- The impact of highly curated or dataset-aligned retrieval corpora
- Alternative fusion methods to increase grounding efficiency in small models
The released code and experiment artifacts provide a strong base for such studies.
Conclusion
This controlled, head-to-head comparison at deployment-relevant scale demonstrates that for medical MCQA with 4B-parameter models, domain fine-tuning yields significant and robust accuracy improvements while RAG does not provide statistically meaningful gains. For tasks requiring substantive domain reasoning, in-weights adaptation should be prioritized over retrieval-based augmentation under limited parameter budgets. These results serve to re-calibrate engineering strategy in the rapidly evolving landscape of small, local LLMs for medical applications.


