- The paper presents HEG-TKG, a two-tier temporal knowledge graph method that reduces the Provenance Gap by over 50%, ensuring 100% citation verifiability in rare disease reasoning.
- It employs a structured approach with GOLD, SILVER, and BRONZE tiers, combining manual curation with automated extraction to maintain temporal precision and evidence quality.
- Empirical analysis shows conventional LLMs often fabricate citations, whereas HEG-TKG achieves robust clinical feature coverage while significantly enhancing auditability and clinician trust.
Evidence-Traceable Temporal Knowledge Graphs Addressing the Provenance Gap in Clinical AI
Introduction and Context
Clinical reasoning and decision support are increasingly augmented by LLMs. Despite their fluency and high performance on medical benchmarks, these models remain fundamentally limited by citation verifiability deficits, especially for rare diseases where trust and auditability are critical. This systematic inability to ground clinical assertions in checkable primary literature is termed the Provenance Gap: the measurable distance between a model's claimed coverage of clinical features and the capacity to trace those features to specific, verifiable sources. The work introduces both a metric and a method—HEG-TKG (Hierarchical Evidence-Grounded Temporal Knowledge Graphs)—to bridge this gap, structuring temporal and quality-stratified evidence from primary literature and curated sources, and evaluating its impact on both numerical metrics and clinician trust.
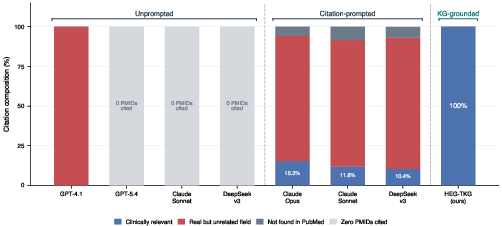
Figure 1: Citation verifiability across five frontier LLMs and HEG-TKG, showing only HEG-TKG achieves fully relevant (clinically verifiable) PMIDs while LLMs frequently fabricate or misattribute citations.
Empirical Analysis of the Provenance Gap
The benchmarking protocol involves five state-of-the-art LLMs (OpenAI GPT-4.1/5.4, Anthropic Claude Sonnet/Opus, DeepSeek-v3) on 36 clinically validated scenarios spanning three phenotypically confusable rare disease pairs. In vanilla inference, all LLMs produced zero correct PMIDs; in citation-prompted mode, the best performing (Claude Opus) achieved only 15.3% relevant PMIDs, with the majority resolving to unrelated but real PubMed records. The result is robust: real but contextually irrelevant citations create false plausibility, undermining clinician safety audits.
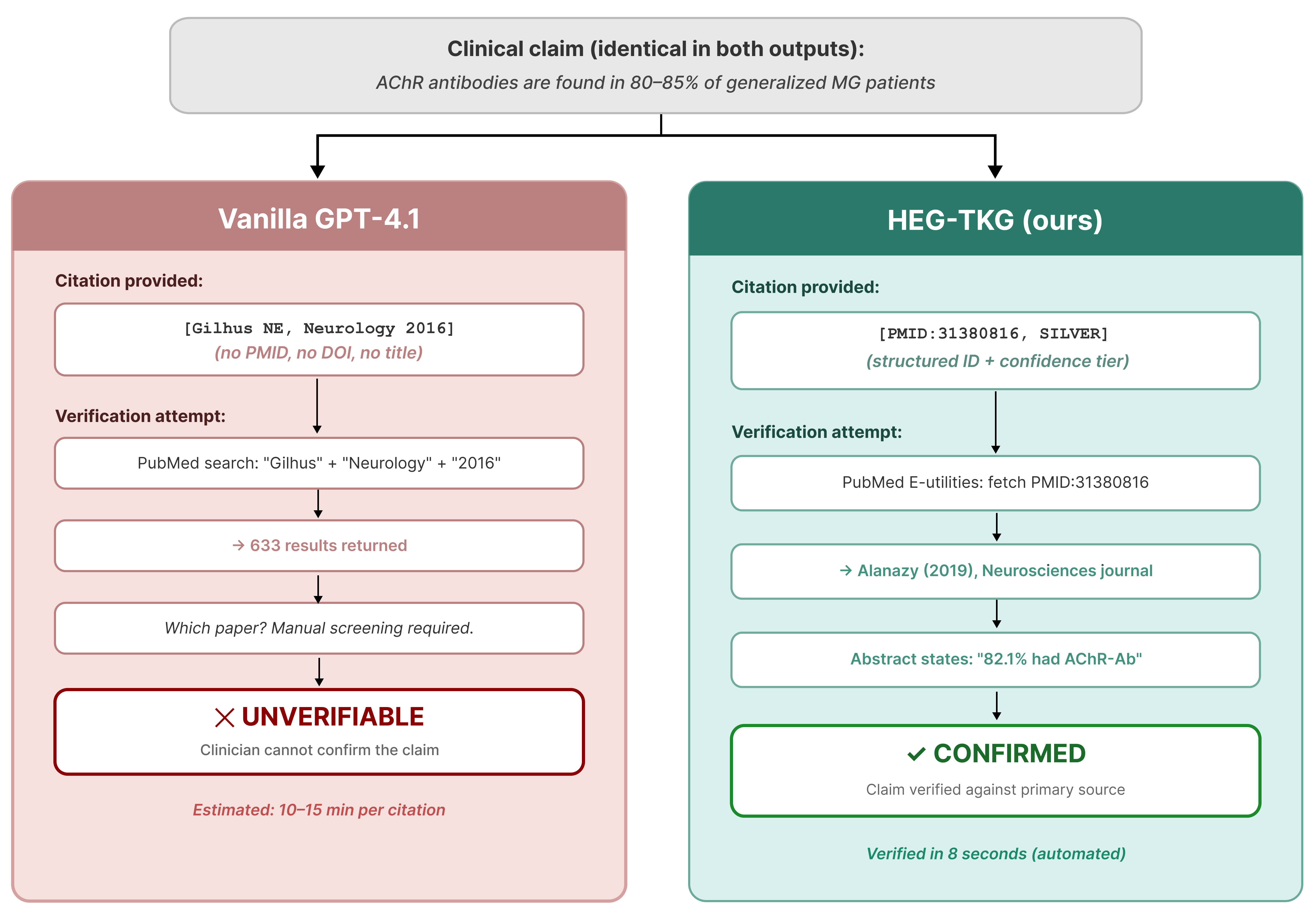
Figure 2: Vanilla LLMs (left) produce vague or unverifiable author-year citations; HEG-TKG (right) emits a precise, rapidly verifiable PMID.
Prompt engineering is insufficient; explicit citation requirements guide models to produce more plausible identifiers, but not more reliable provenance. Manual PubMed lookup demonstrates that models produce superficially correct but contextually decoupled citations, revealing a structural limitation in LLM parametric memory.
HEG-TKG: Methodology and System Architecture
The HEG-TKG pipeline operationalizes clinical verifiability using a two-plane, two-tier architecture:
- Tier 1 ("GOLD"): programmatically curated backbone drawn from GeneReviews, OMIM, Orphanet, and CDC guidelines, guaranteeing a floor of clinically authoritative edges.
- Tier 2 ("BRONZE"/"SILVER"): automated extraction from 4,512 PubMed records using a schema-guided, multi-LLM, consensus-driven pipeline with entity normalization, semantic correction (18 rule sets), and explicit temporal anchoring (ISO 8601 precision levels).
Edges carry provenance metadata: PMID(s), evidence tiers reflecting extraction/model consensus, and precise temporal anchorings absent from PrimeKG, UMLS, and prior biomedical KGs [chandrasekaran2023primekg]. The pipeline is disease pair-configurable, supporting direct expansion.
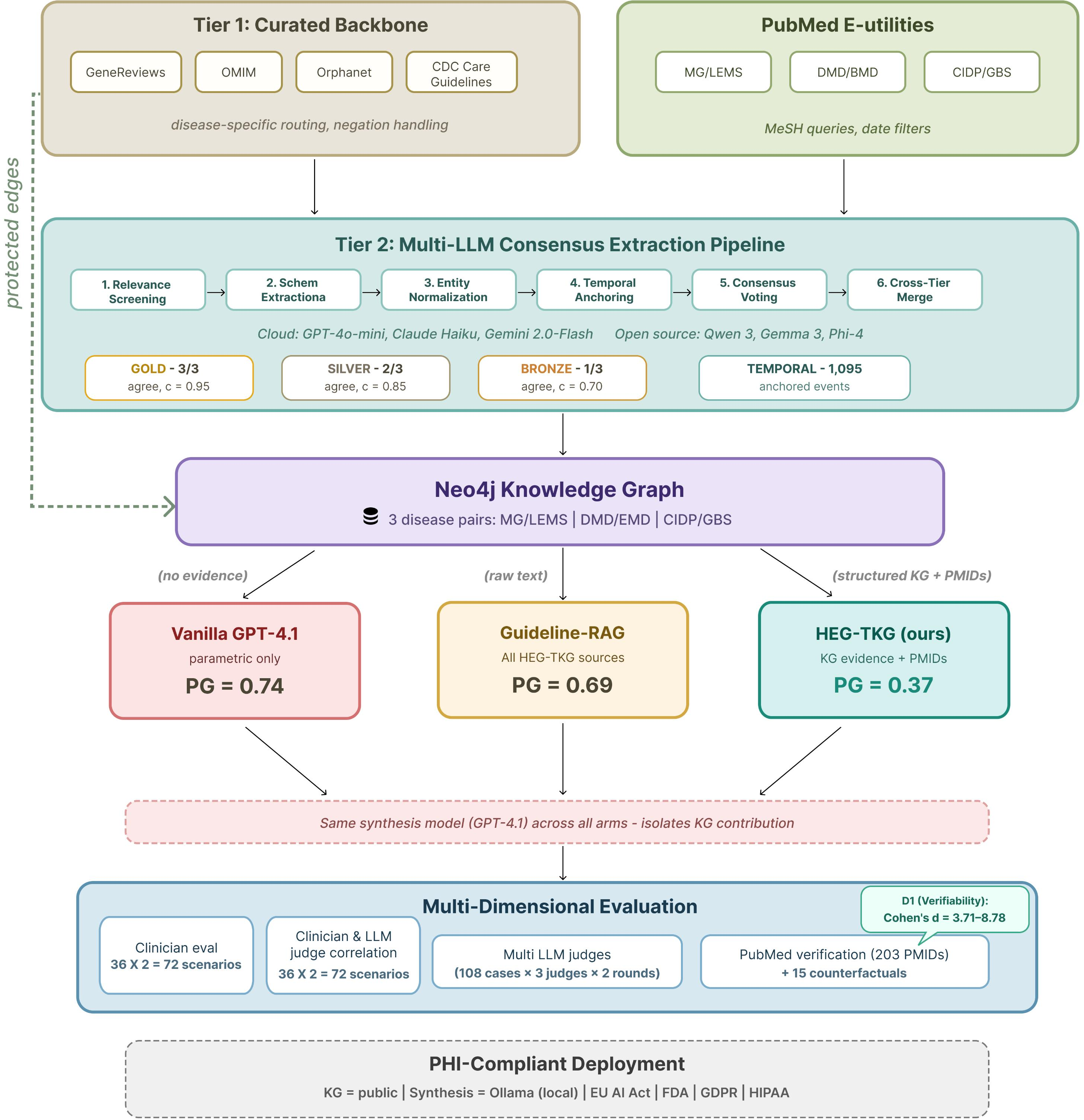
Figure 4: HEG-TKG system architecture, with separation between curated Tier 1, extraction-based Tier 2, and downstream clinical inference.
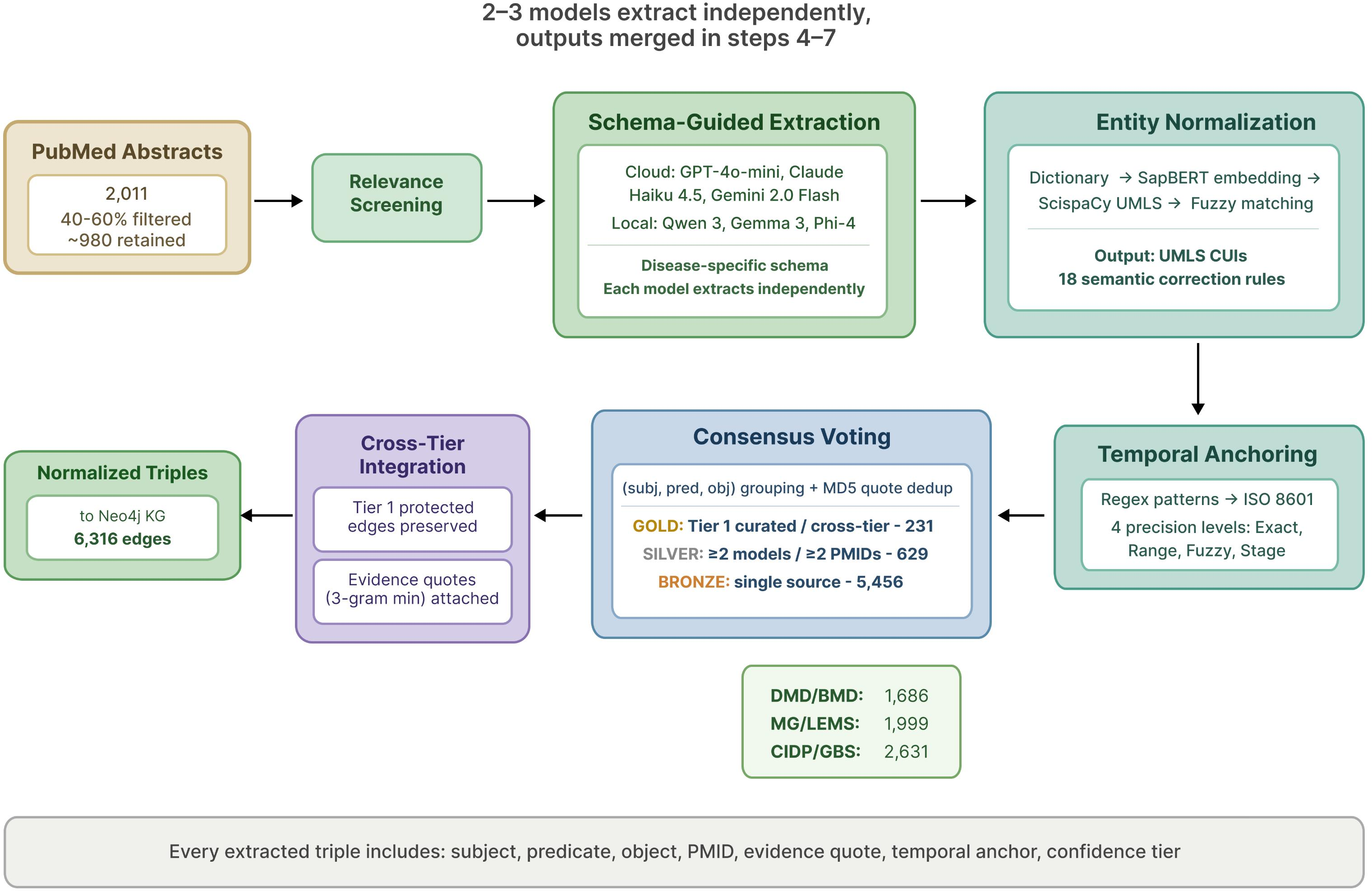
Figure 6: Tier 2 literature extraction pipeline, highlighting relevance screening, normalization, temporal anchoring, consensus voting, and integration.
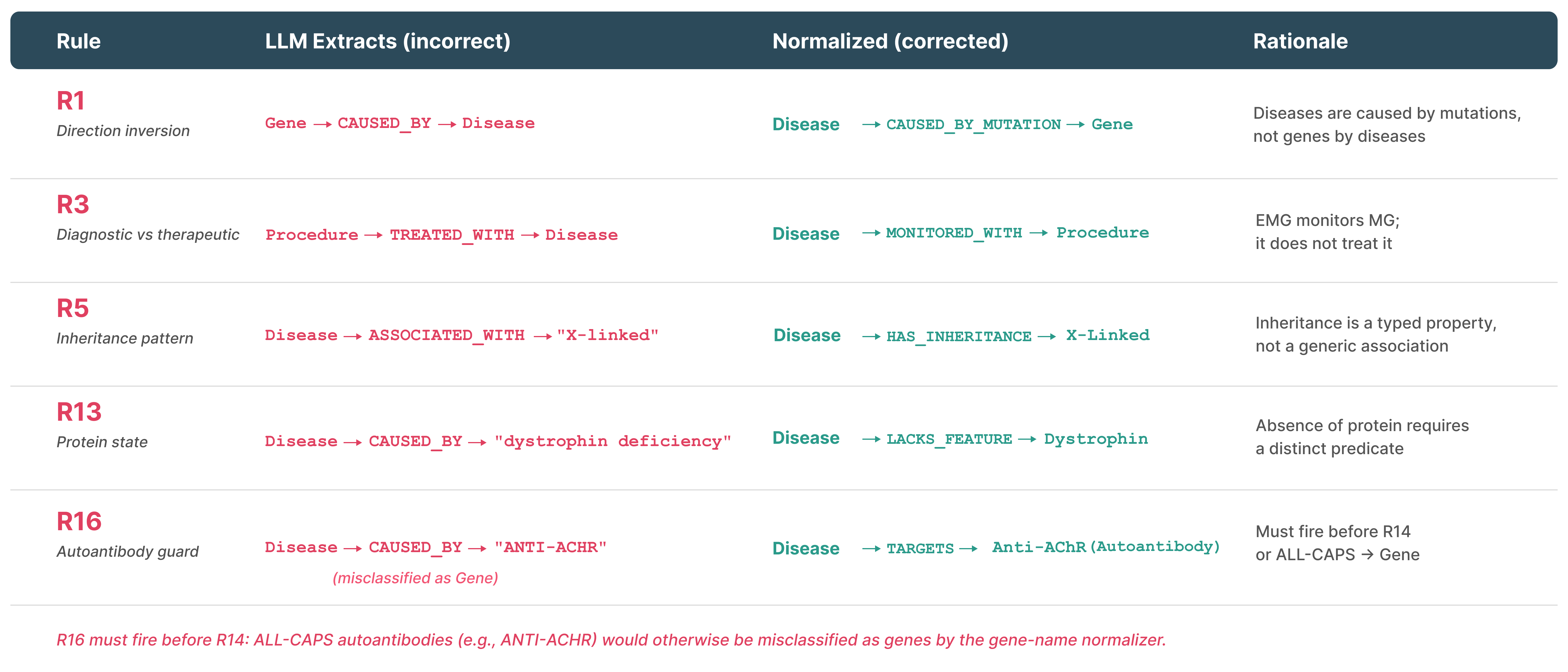
Figure 8: Representative semantic normalization and correction rules mitigating extraction errors.
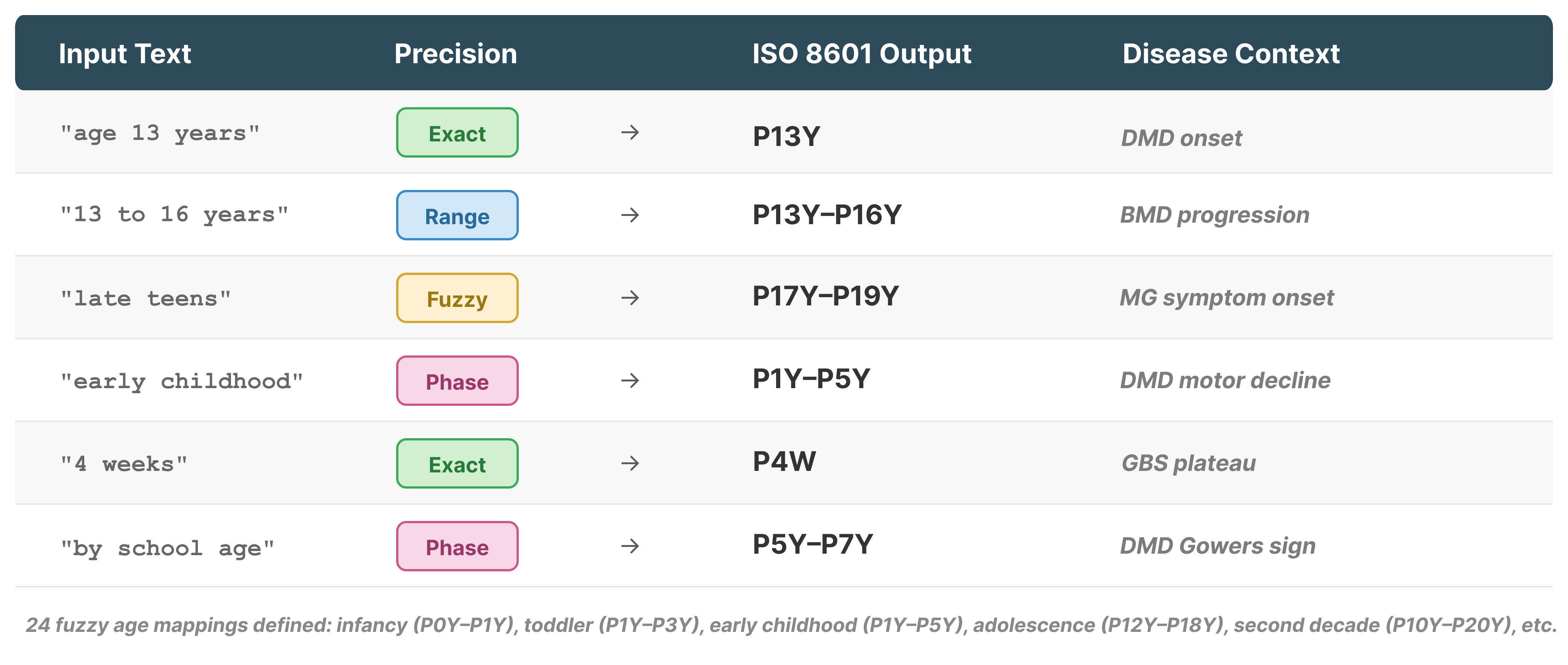
Figure 3: Examples of temporal anchor resolution, systematically mapping narrative time expressions to formal ISO 8601 intervals.
Three-Arm Comparative Evaluation: Structured Grounding versus Flat Retrieval
The controlled experiment contrasts three modes: vanilla (parametric only), Guideline-RAG (retrieved, overlapping raw guideline text), and HEG-TKG (structured graph evidence). All arms use GPT-4.1 for synthesis, isolating the effect of evidence structure.
HEG-TKG matches vanilla and RAG arms on automated clinical feature coverage (71.7% vs 74.3/68.8%), but reduces the Provenance Gap by >50% (0.37 vs 0.69–0.74)—every claim is traceable to specific PMID-backed evidence, with 100% of citations audited as correct and relevant.
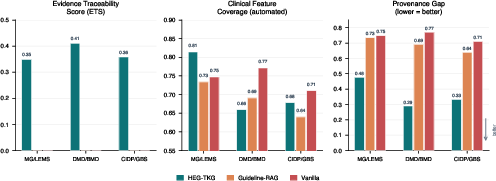
Figure 5: Three-arm comparison of Evidence Traceability Score, Feature Coverage, and Provenance Gap—HEG-TKG achieves comparable coverage with far higher verifiability.
Guideline-RAG, despite being given the same clinical source material as HEG-TKG (as free text), produces zero verifiable PMIDs; only structured citation metadata produces programmatically checkable evidence trails.
Quality Stratification and Temporal Reasoning
The knowledge graphs span 5,481 nodes, 6,316 edges, 987 unique PMIDs, and 1,280 temporal anchors. Each edge is dynamically assigned a confidence weight: GOLD (0.95, cross-tier confirmed/curated), SILVER (0.85, multi-model or multi-PMID consensus), BRONZE (0.70, single source/model). Temporal anchors encode progression, onset, and intervention windows at multiple precision levels, enabling sequence-aware phenotype evolution and management.
Temporal claims (e.g., ages of diagnostic milestones, onset of specific complications) are a central innovation; LLMs and baseline KGs cannot support such queries beyond static associations.
Clinician and LLM-Judge Evaluation
Blinded clinician assessment confirms a statistically significant advantage for evidence-grounded reasoning on verifiability (D1: Δ=+1.64; Cohen’s d=1.81; p<0.001). Actionability is also increased, with no sacrifice in safety or completeness. Notably, experience-dependent variation is observed: some clinicians regard citation of consensus knowledge as redundant, flattening inter-rater differences.
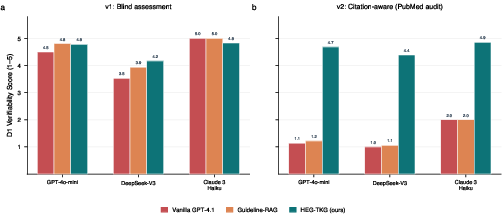
Figure 7: LLM judges cannot reliably evaluate verifiability without external ground-truth; only when presented with audit data can they distinguish fabrications from valid citation structures.
A two-stage LLM-Judge protocol further reveals that automatic (LLM) evaluation of citation quality tracks only surface plausibility absent external audits—exposing a critical flaw in routine automated evaluation designs. With explicit PubMed verification data, LLM-judge reliability (Krippendorff's α) increases from -0.09 to 0.89.
Counterfactual Robustness and Error Detection
A targeted safety analysis involves injecting clinically incorrect evidence into the knowledge graph. Remarkably, 80% of spurious clinical claims are resisted by the synthesis model’s parametric prior, and 100% of injected errors are traceable through the citation graph—even when accepted by the synthesis model, a human auditor can instantly localize and verify the provenance of any anomalous recommendation.
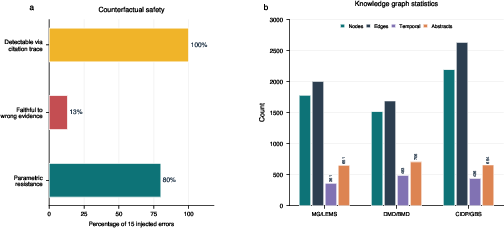
Figure 9: Of 15 injected counterfactual errors, 80% are ignored, and all remain fully auditable via citation path.
Practical and Regulatory Implications
HEG-TKG supports full on-premise deployment with open-source LLMs (Ollama/Qwen/Gemma), sidestepping PHI exposure to cloud APIs and aligning with GDPR and HIPAA mandates. Verification time per claim drops from minutes to seconds. The system design enables real clinical workflows (e.g., attaching PMID-anchored outputs to referral pathways), addresses auditability requirements under EU AI Act and FDA CDS guidance, and establishes a baseline for reproducibility—something not systematically achievable with either parametric or agentic retrieval models.
Methodological Contributions
The rigorous separation of evidence planes, programmatic citation auditing, and multi-judge, multi-arm, multi-rater statistics establish a framework for benchmarking and for future model accountability. The work demonstrates that neither model scale, provider, nor prompt engineering closes the Provenance Gap; only structured provenance-grounded synthesis does. Automated judges are shown to be insufficient for evaluation of clinical citation quality absent ground-truth data, with direct implications for future clinical AI validation standards.
Limitations and Future Directions
The evaluation scope is limited to three disease pairs in neuromuscular medicine, with coverage primarily at the abstract level. Expansion to other specialties, full-text literature, further clinician panels, and integration with agentic retrieval are planned. The direct measurement and minimization of the Provenance Gap should become standard practice for clinical AI systems.
Conclusion
Addressing clinical verifiability in AI is not a matter of scaling LLMs or iterating prompts. The consequences of fabricated, irrelevant, or unverifiable citations are of utmost importance where trust is non-negotiable. Through HEG-TKG, the evidence-grounded, temporally structured, and quality-stratified approach enables LLM outputs that are as auditable as they are accurate. This work sets a new standard for evidence traceability in clinical AI, shifting the focus from surface knowledge to structured, reviewable, and temporally contextualized reasoning.
Reference:
"The Provenance Gap in Clinical AI: Evidence-Traceable Temporal Knowledge Graphs for Rare Disease Reasoning" (2604.17114)








