- The paper presents SAFE-T, a framework that unifies context-dependent molecule generation, scoring, and fragment-level interpretability via conditional chemical language modeling.
- It demonstrates state-of-the-art performance in zero-shot and goal-directed optimization, achieving >99% validity and high ROC-AUC in virtual screening tasks.
- The model's fragment-level attribution recovers known pharmacophores and SAR motifs, providing actionable insights for medicinal chemistry.
Versatile Conditional Chemical LLMs for Drug Discovery: A Critical Technical Assessment
Introduction
The paper "Conditional Chemical LLMs are Versatile Tools in Drug Discovery" (2507.10273) addresses key limitations of generative chemical LLMs (CLMs) in molecular design—particularly the lack of reward functions grounded in biological objectives and limited interpretability for medicinal chemistry applications. The proposed framework, SAFE-T (Sequential Attachment-based Fragment Embedding with Target-conditioning), unifies context-dependent molecule generation, scoring, and fragment-level interpretability within an end-to-end, biologically conditioned language modeling paradigm.
Unlike previous property-conditioned or structure-based generative models, SAFE-T leverages chemical fragment sequences (SAFE strings) and discrete biological prompts encoding target family, protein, and mechanism-of-action (MoA). This enables robust performance in zero-shot and few-shot drug discovery tasks without reliance on explicit structural or engineered objective functions.
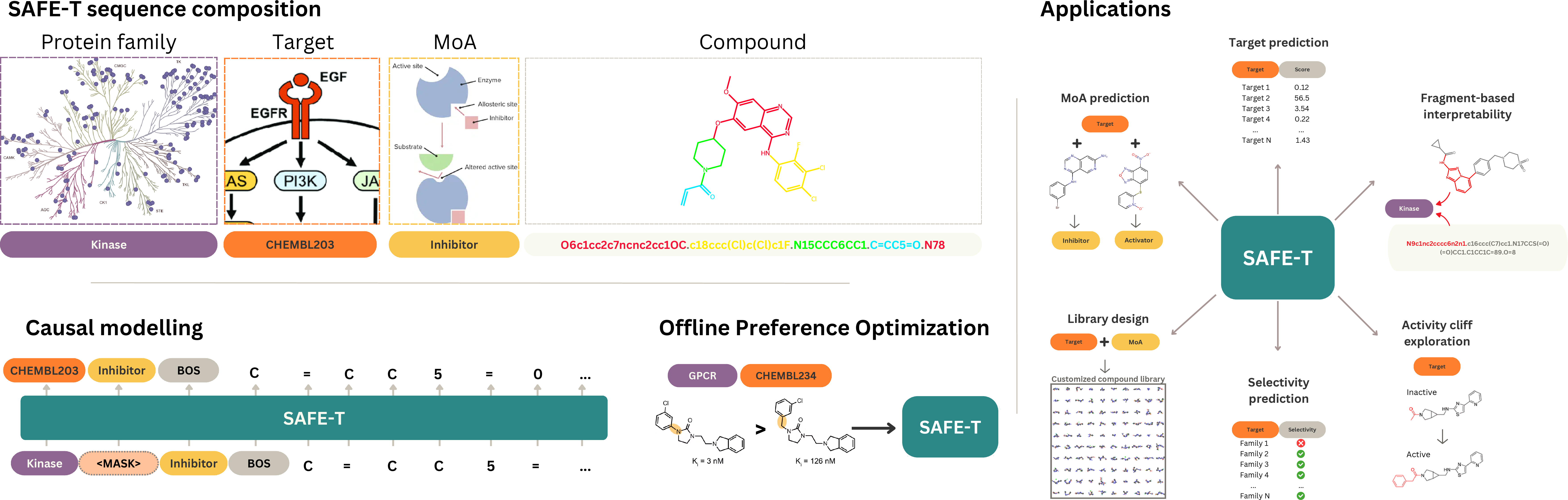
Figure 1: SAFE-T framework overview integrating fragment-based sequences and biological context tokens through staged pretraining, context fine-tuning, and preference optimization, supporting generalization in diverse drug discovery tasks.
SAFE-T: Framework and Learning Algorithm
SAFE-T adopts a Transformer-based autoregressive CLM architecture that operates over molecular fragments and appended biological context tokens. Training proceeds in three sequential stages:
- Chemical Pretraining: The model is first exposed to fragment-based molecular representation using a large corpus of unlabeled chemical space with masked biological context, ensuring baseline validity and reliable molecular syntax generation.
- Biological Context Fine-tuning: SAFE-T is then conditioned on triplets (target family, target ID, MoA) using the curated MoAT-DB dataset, enabling explicit learning of bio-structural associations and making the model robust to missing or partial biological specification via aggressive random context masking.
- Preference Calibration: Pairwise preference data (obtained from activity cliff datasets) is used to calibrate conditional likelihoods for ranking and prioritization, enhancing the reflection of subtle SAR nuances and improving detection of large activity discontinuities from minor structural modifications.
Crucially, all use-cases—goal-directed generation, prioritization, selectivity profiling, and local SAR interpretability—are derived from the same core conditional probability model.
Zero-shot and Goal-directed Molecular Generation
SAFE-T demonstrates high validity (>99%) and competitive diversity in both unconditional and context-conditioned generation tasks across multiple standard benchmarks. The integration of context conditioning does not degrade generative performance or constraint satisfaction, as shown by high synthetic accessibility and QED values. Notably, SAFE-T achieves state-of-the-art performance in zero-shot goal-directed optimization on PMO benchmarks (DRD2, JNK3, GSK-3β), with top-10 AUCs exceeding 0.99, representing a substantial delta over RL- or search-based approaches and highlighting strong generalization without target-specific retraining.
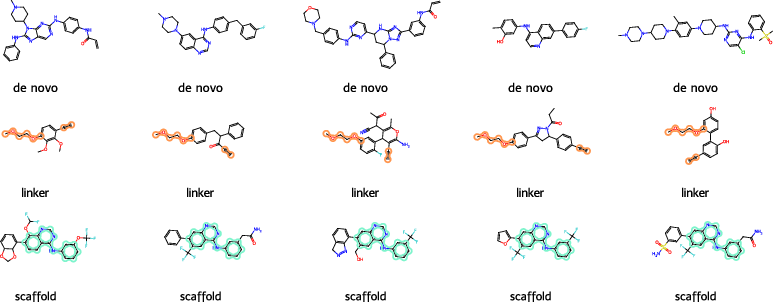
Figure 3: Representative molecules generated by SAFE-T under various target conditions; chemical validity and diversity are preserved.
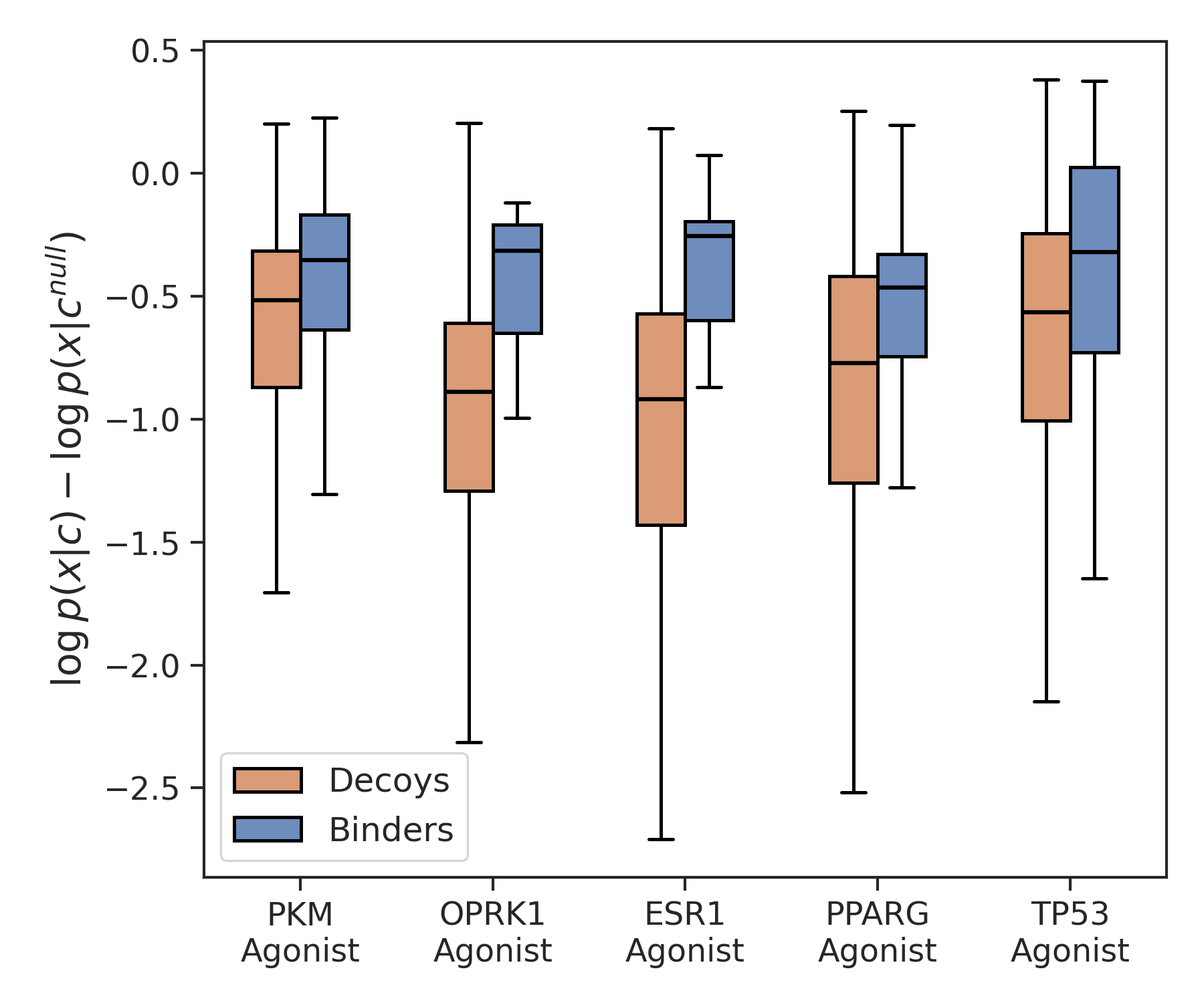
SAFE-T's conditional likelihoods serve directly for both context classification (target/MoA inference for a molecule) and virtual screening (ranking by likelihood within a context). It achieves strong ROC-AUC and enrichment (EF@1%) across realistic and challenging datasets.
Activity Cliff Detection and SAR Interpretability
Preference tuning in SAFE-T significantly enhances the detection of activity cliffs, yielding ROC-AUC up to 0.95—on par with GNN and fingerprint-based ensembles. The model's fragment-level attribution, performed via counterfactual fragment replacement and comparison of likelihood shifts, recovers known pharmacophores, selectivity motifs, and non-trivial SARs as validated on JAK inhibitors (e.g., Tofacitinib, Ruxolitinib). SAFE-T achieves superior attribution accuracy relative to competitive fragment-aware models, associating largest scores with essential functional groups in line with medicinal chemistry literature.
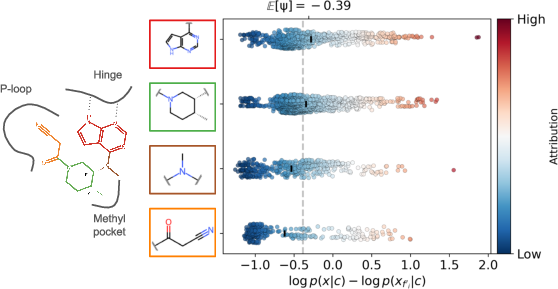
Figure 5: Fragment-level attribution on Tofacitinib reveals correspondence between likelihood shifts and established hinge-binder, P-loop, and selectivity motifs.
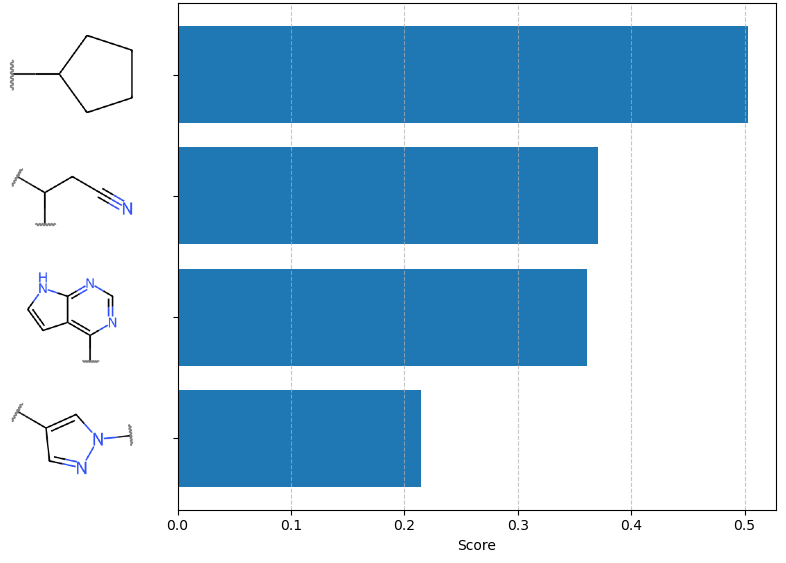
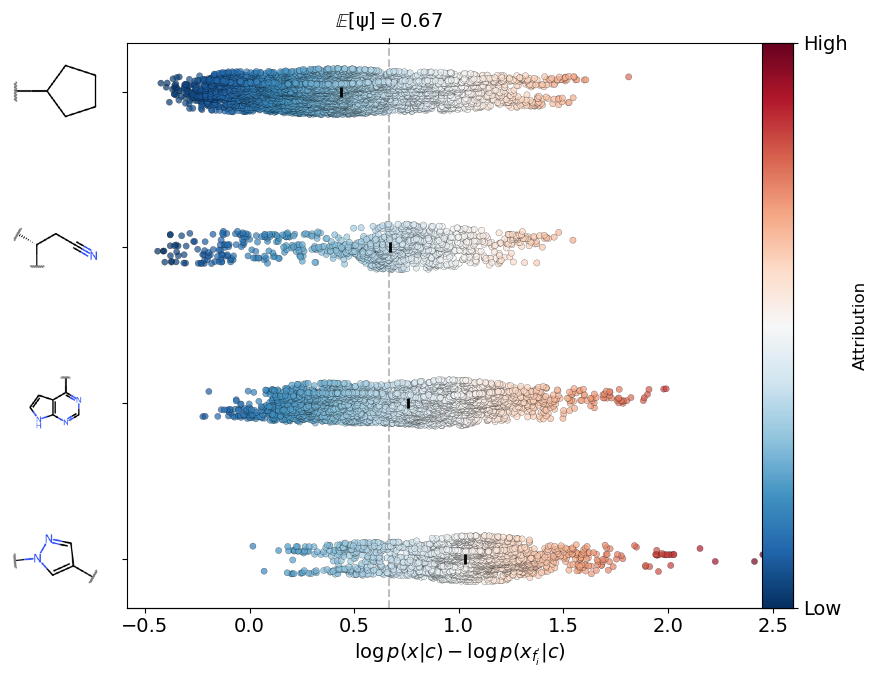
Figure 7: Comparative fragment attribution on Ruxolitinib; SAFE-T highlights the pharmacophore core over non-essential fragments, outperforming alternative models (e.g., GEAM) in interpretability.
Chemical and Biological Contextual Generalization
SAFE-T generalizes robustly across both unseen chemotypes and target–MoA pairs, as well as to new protein and MoA encodings absent from the original training set. Ablation studies show that model scaling (>25M parameters) enhances both predictive and generative performance. However, preference tuning, though essential for fine-grained SAR and cliff detection, can adversely affect generation validity in smaller models—a trade-off mitigated by increased capacity.
Detailed analysis of MoAT-DB reveals that SAFE-T's training data represents a broad swath of protein families and compound diversity, which is essential for the scalable nature of the model's generalization.
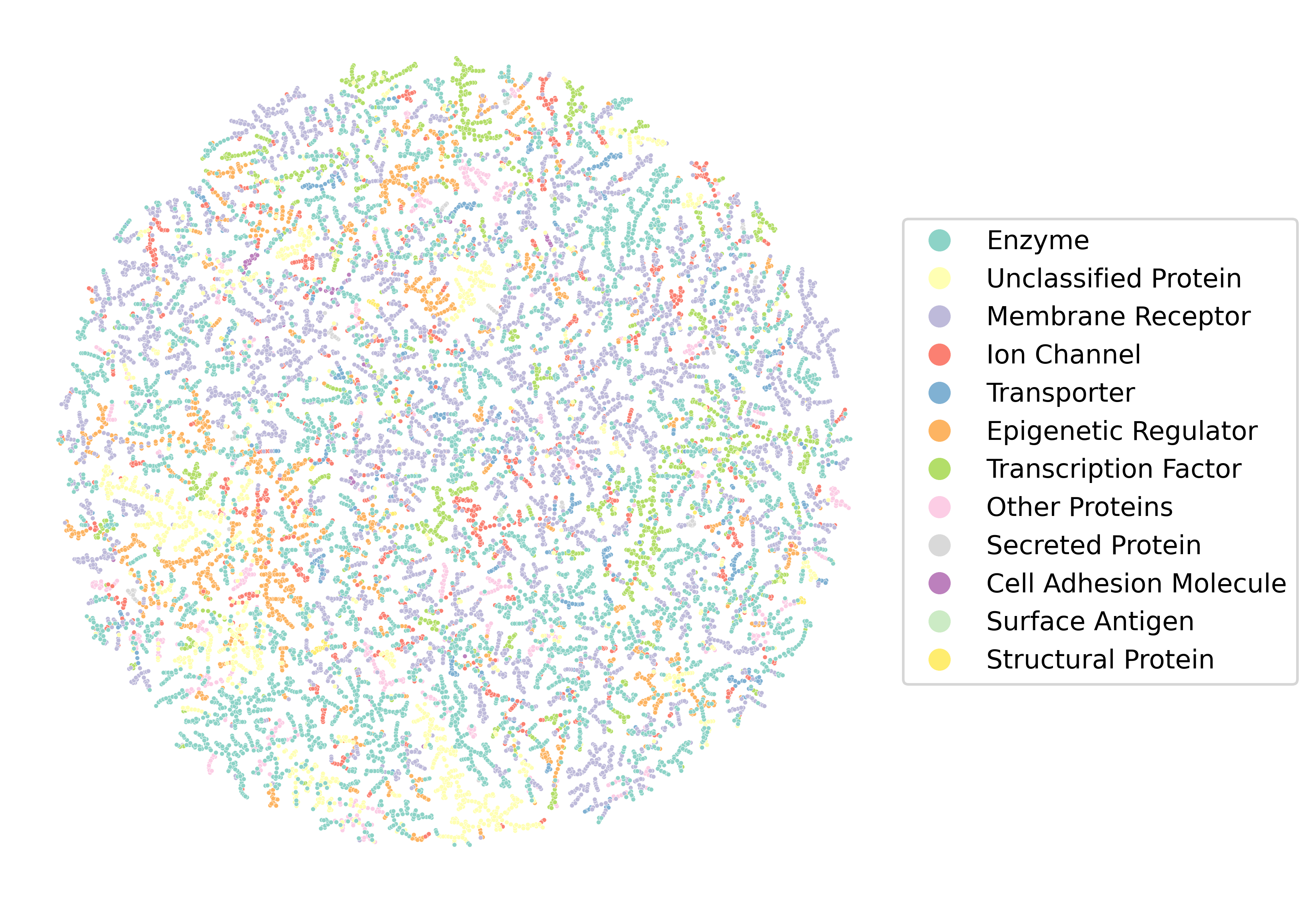

Figure 2: MoAT-DB's chemical space visualized by TMAP; family-level clusters support the model's context-aware generalization.
Implications and Future Directions
SAFE-T exemplifies a trend toward unifying scoring, generation, and interpretability in fragment-based, conditional CLMs for therapeutic discovery. This framework eliminates the need for reward heuristics, structure-based objective functions, or costly target-specific retraining, substantially reducing early-stage discovery cycle times. Its capacity to provide interpretable attribution and actionable SAR insights, without sacrificing generative flexibility or throughput, is highly pertinent for both computational and experimental medicinal chemistry pipelines.
Several limitations persist: data coverage and annotation quality (particularly across the inactive chemical space) remain the main bottleneck for further performance gains, especially in SAR-facing tasks such as activity cliff prediction. Masking strategies for incomplete context and adaptation to rare targets indicate robustness, but practical application in polypharmacology and lead optimization may benefit from further extension to continuous, structural, or pathway conditioning. Integration with protein structural embeddings or LLM-driven reasoning over broader biological signals is a logical progression, as are direct coupling with in silico or high-content screening platforms for iterative, closed-loop design.
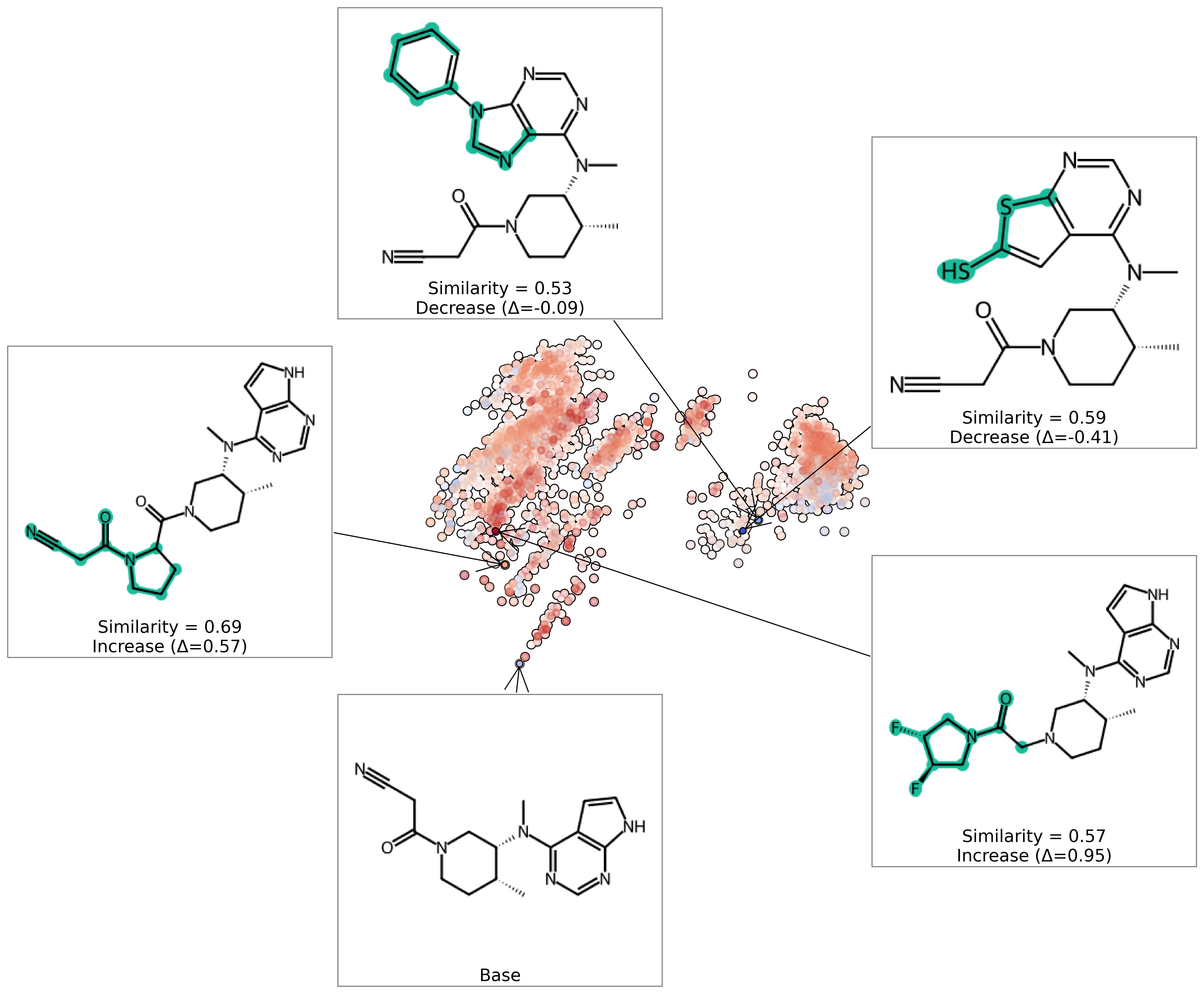
Figure 9: Counterfactual analysis around Tofacitinib demonstrates that targeted fragment substitutions systematically shift predicted bioactivity probability, supporting lead optimization via minimal edits.
Conclusion
SAFE-T demonstrates that conditional generative CLMs over fragment-based molecular representations, with explicit biological context conditioning, can produce unified, high-performing, and interpretable models for early-stage drug discovery. The framework addresses critical limitations of prior generative paradigms and opens new directions for scalable, generalist, and mechanistically-grounded AI in medicinal chemistry. While further integration with richer biological modalities and larger annotated datasets will push performance ceilings, the reported results set a strong benchmark for biologically informed generative modeling in therapeutics.
Reference:
"Conditional Chemical LLMs are Versatile Tools in Drug Discovery" (2507.10273)