Biological computations: limitations of attractor-based formalisms and the need for transients
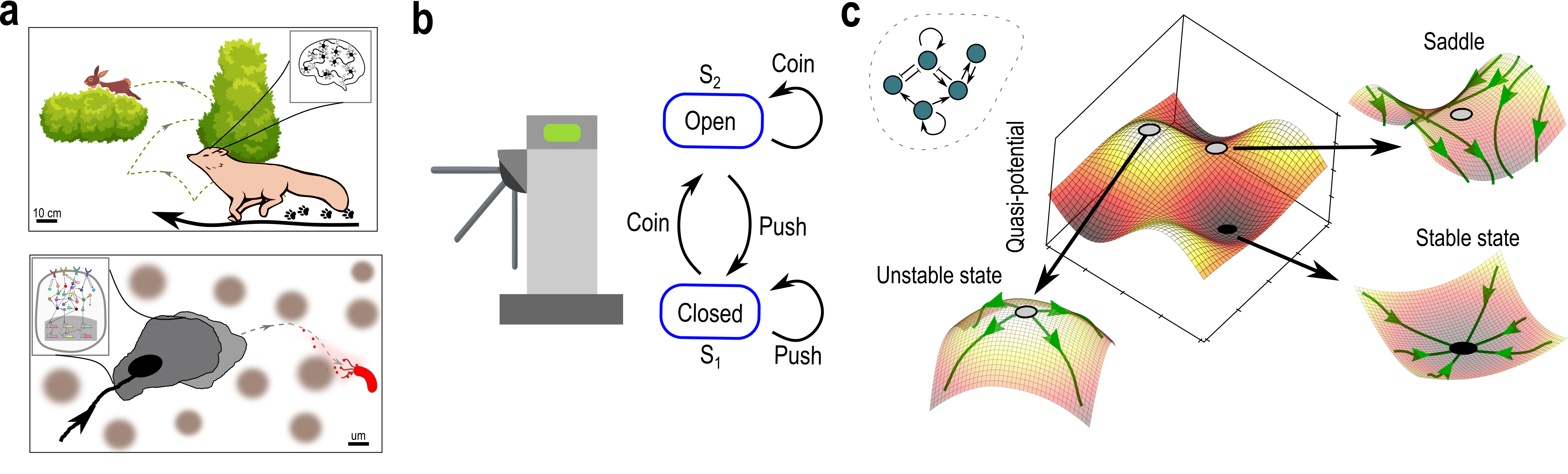
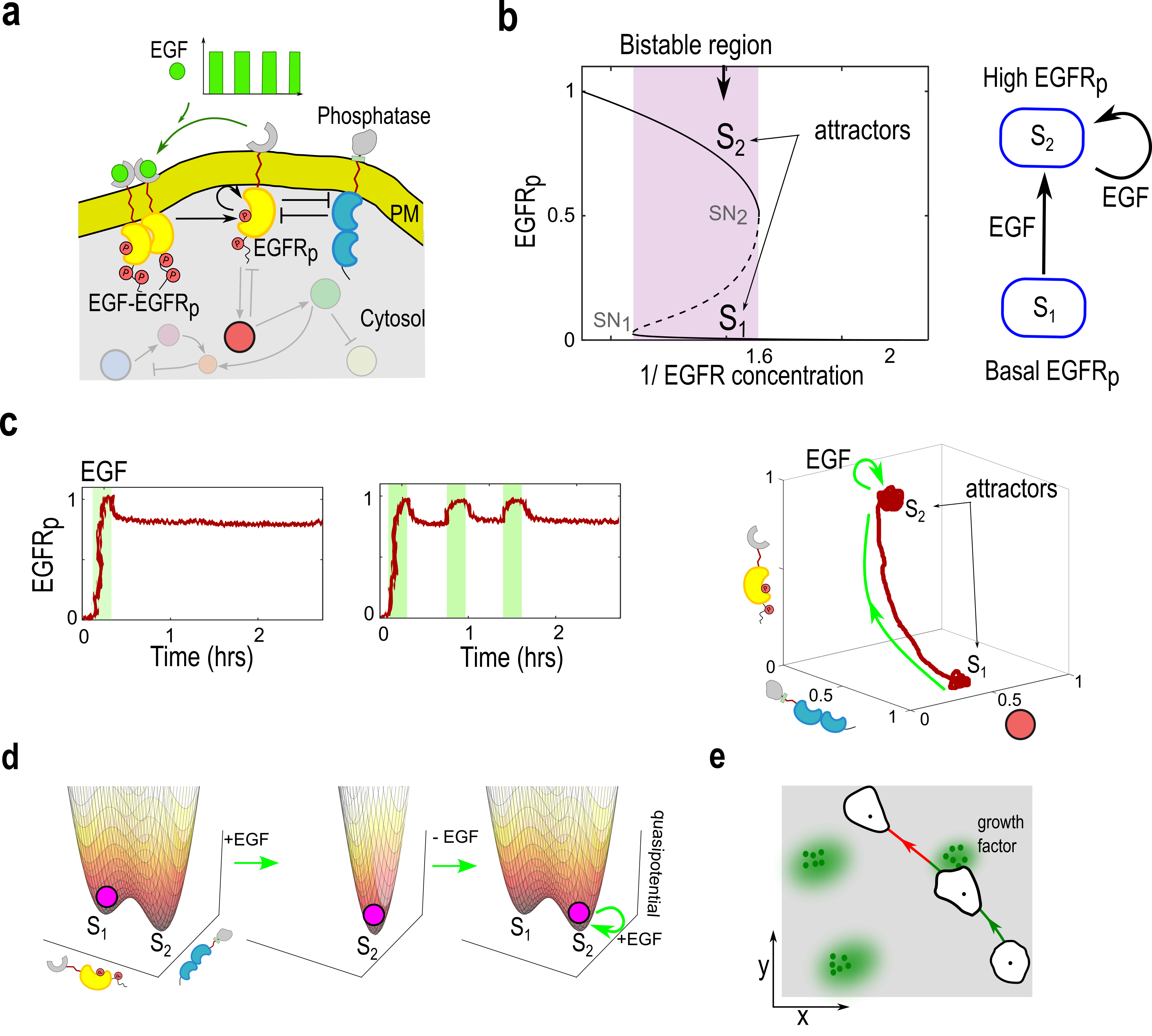
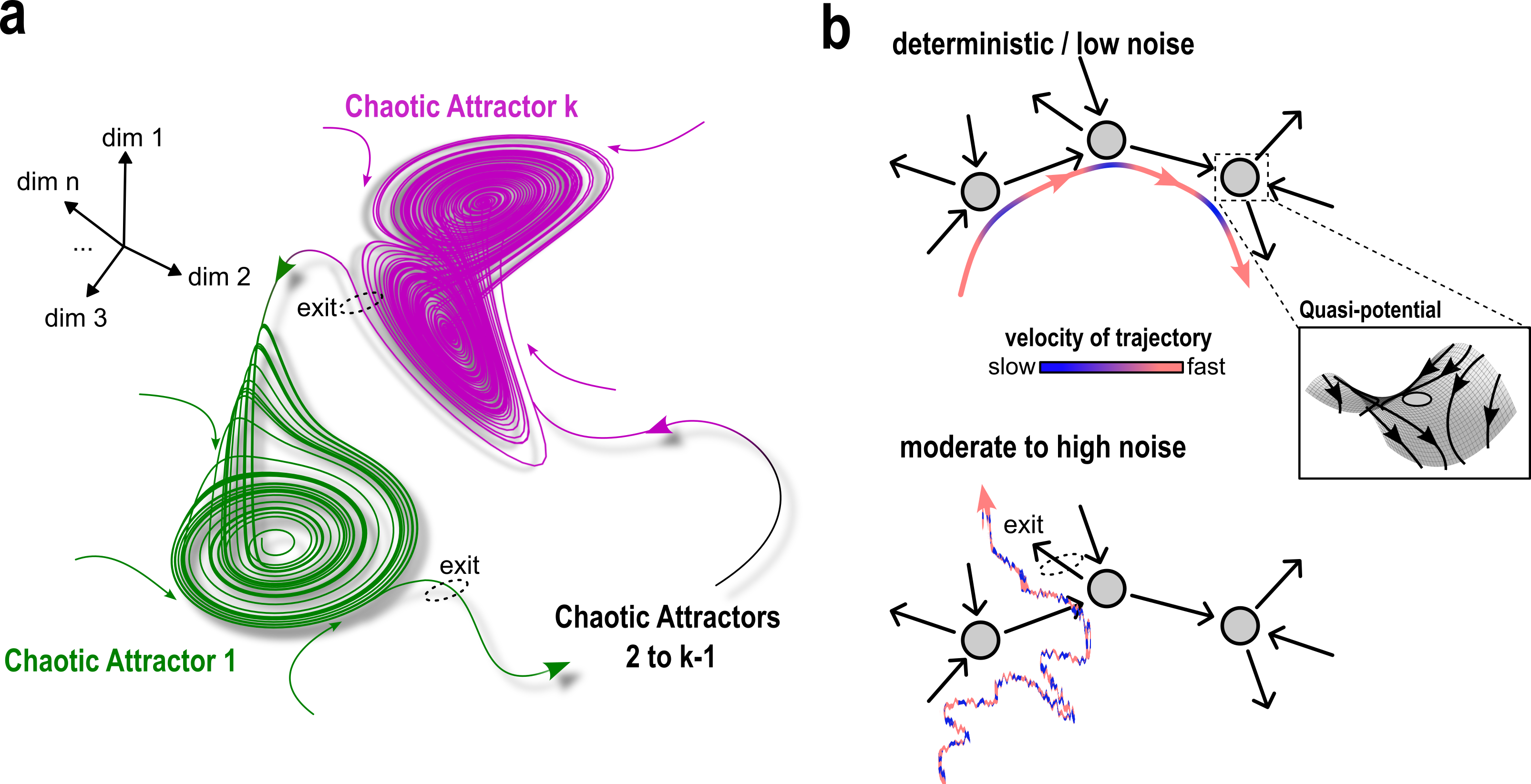
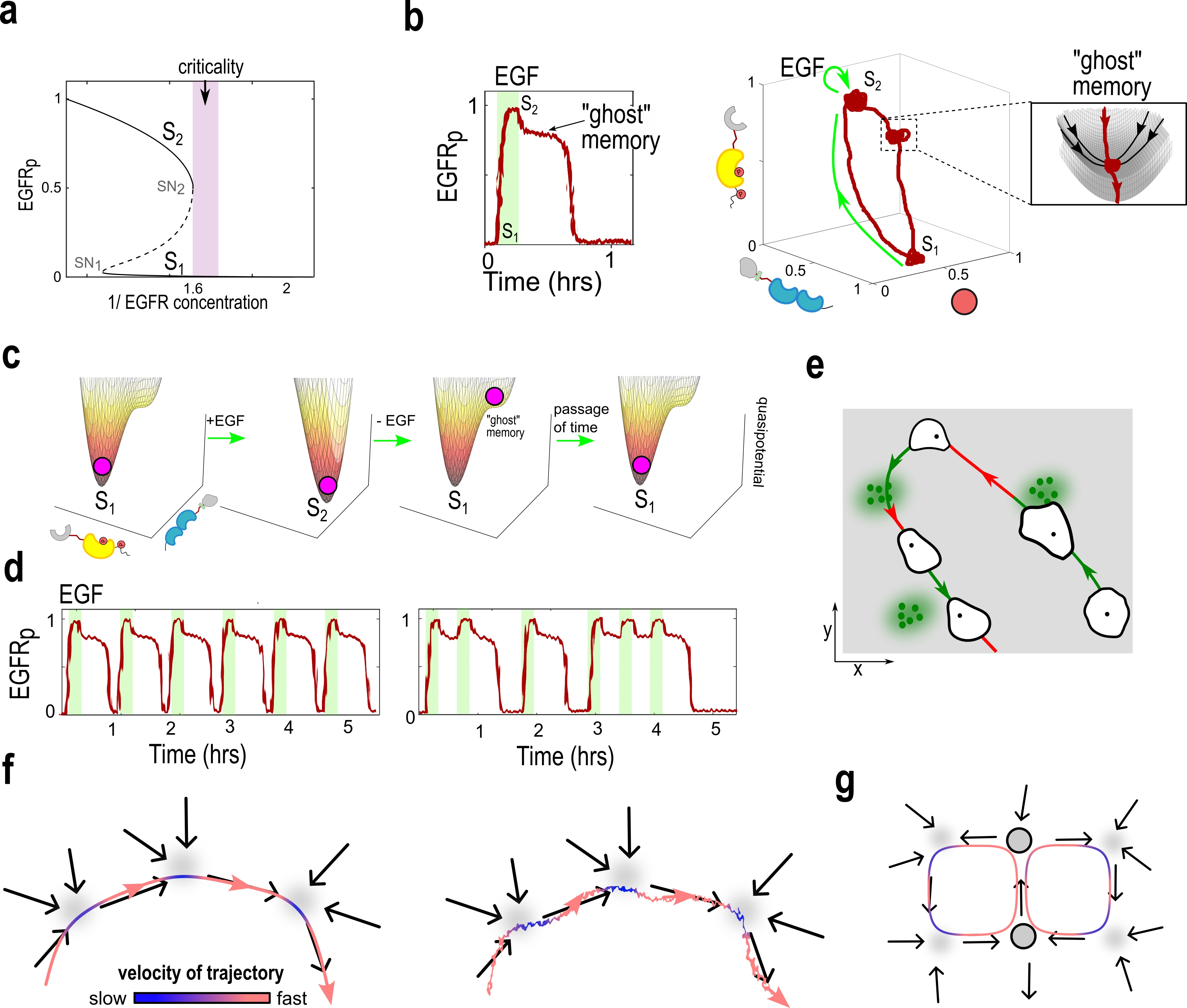
Abstract: Living systems, from single cells to higher vertebrates, receive a continuous stream of non-stationary inputs that they sense, e.g., via cell surface receptors or sensory organs. Integrating these time-varying, multi-sensory, and often noisy information with memory using complex molecular or neuronal networks, they generate a variety of responses beyond simple stimulus-response association, including avoidance behavior, life-long-learning or social interactions. In a broad sense, these processes can be understood as a type of biological computation. Taking as a basis generic features of biological computations, such as real-time responsiveness or robustness and flexibility of the computation, we highlight the limitations of the current attractor-based framework for understanding computations in biological systems. We argue that frameworks based on transient dynamics away from attractors are better suited for the description of computations performed by neuronal and signaling networks. In particular, we discuss how quasi-stable transient dynamics from ghost states that emerge at criticality have a promising potential for developing an integrated framework of computations, that can help us understand how living system actively process information and learn from their continuously changing environment.
- Six distinct NFκ𝜅\kappaitalic_κb signaling codons convey discrete information to distinguish stimuli and enable appropriate macrophage responses. Immunity, 54(5):916–930.e7, 2021.
- Associative mechanisms allow for social learning and cultural transmission of string pulling in an insect. PLOS Biology, 14(10):e1002564, 2016.
- U. Alon. An introduction to systems biology: design principles of biological circuits. CRC press, 2019.
- P. Ashwin and J. Borresen. Discrete computation using a perturbed heteroclinic network. Physics Letters A, 347(4-6):208–214, 2005.
- P. Ashwin and M. Timme. When instability makes sense. Nature, 436(7047):36–37, 2005.
- Slower prefrontal metastable dynamics during deliberation predicts error trials in a distance discrimination task. Cell Reports, 35(1):108934, 2021.
- G. Bondanelli and S. Ostojic. Coding with transient trajectories in recurrent neural networks. PLOS Computational Biology, 16(2):e1007655, 2020.
- A. Buttenschön and L. Edelstein-Keshet. Cell repolarization: A bifurcation study of spatio-temporal perturbations of polar cells. Bulletin of Mathematical Biology, 84(10), 2022.
- Soft-wired long-term memory in a natural recurrent neuronal network. Chaos: An Interdisciplinary Journal of Nonlinear Science, 30(6), 2020.
- D. Chialvo. Emergent complex neural dynamics. Nat. Phys., 6:744, 2010.
- C. Choudhary and M. Mann. Decoding signalling networks by mass spectrometry-based proteomics. Nature Reviews Molecular Cell Biology, 11(6):427–439, 2010.
- A. Dancin. Bacteria as computers making computers. FEMS Microbiol. Rev., 33:3–26, 2009.
- M. Davis. What is a computation? In: Mathematics Today Twelve Informal Essays, In: Lynn Arthur Steen (ed.), Mathematics Today Twelve Informal Essays, 1978.
- A complex hierarchy of avoidance behaviors in a single-cell eukaryote. Current Biology, 29(24):4323–4329.e2, 2019.
- A. Dussutour. Learning in single cell organisms. Biochemical and Biophysical Research Communications, 564:92–102, 2021.
- Gene regulatory networks and their applications: understanding biological and medical problems in terms of networks. Frontiers in Cell and Developmental Biology, 2:38, 2014.
- Novel generic models for differentiating stem cells reveal oscillatory mechanisms. Journal of The Royal Society Interface, 18(183), 2021.
- W. T. Fitch. Information and the single cell. Current Opinion in Neurobiology, 71:150–157, 2021.
- Integrating conflicting chemotactic signals: The role of memory in leukocyte navigation. J of Cell Biol., 147(3):577–588, 1999.
- K. J. Friston. Transients, metastability, and neuronal dynamics. NeuroImage, 5(2):164–171, 1997.
- Reconsidering the evidence for learning in single cells. eLife, 10:e61907, 2021.
- A. N. Gorban. Singularities of transition processes in dynamical systems: Qualitative theory of critical delays. Electronic Journal of Differential Equations, 1(Mon. 05), 2004.
- J. Gunawardena. Learning outside the brain: integrating cognitive science and systems biology. Proceedings of the IEEE, 110(5):590–612, 2022.
- R. M. Harris-Warrick and E. Marder. Modulation of neural networks for behavior. Annual review of neuroscience, 14(1):39–57, 1991.
- D. O. Hebb. The organization of behavior: A neuropsychological theory. Psychology press, 2005.
- M. W. Hirsch and B. Baird. Computing with dynamic attractors in neural networks. Biosystems, 34(1-3):173–195, 1995.
- J. J. Hopfield. Neural networks and physical systems with emergent collective computational abilities. Proceedings of the National Academy of Sciences, 79(8):2554–2558, 1982.
- Maxwell-bloch turbulence. Prog. Theor. Phys. Suppl., 99:295, 1989.
- Designing spontaneous behavioral switching via chaotic itinerancy. Science Advances, 6(46), 2020.
- Distinguishing mechanisms underlying EMT tristability. Cancer Convergence, 1:2, 2017.
- The EGFR phosphatase RPTPγ𝛾\gammaitalic_γ is a redox-regulated suppressor of promigratory signaling. The EMBO Journal, 42(10), 2023.
- Geometric models for robust encoding of dynamical information into embryonic patterns. eLife, 9:e55778, 2020.
- K. Kaneko. Clustering, coding, switching, hierarchical ordering, and control in network of chaotic elements. Physica D, 41:137–172, 1990.
- K. Kaneko. Globally coupled circle maps. Physica D, 54:5–19, 1991.
- K. Kaneko and I. Tsuda. Chaotic itinerancy. Chaos: An Interdisciplinary Journal of Nonlinear Science, 13(3):926–936, 2003.
- Epigenetic inheritance of gene silencing is maintained by a self-tuning mechanism based on resource competition. Cell Systems, 14(1):24–40.e11, 2023.
- Global brain dynamics embed the motor command sequence of caenorhabditis elegans. Cell, 163(3):656–669, 2015.
- Cyclical fate restriction: a new view of neural crest cell fate specification. Development, 148(22), 2021.
- R. Kim and T. J. Sejnowski. Strong inhibitory signaling underlies stable temporal dynamics and working memory in spiking neural networks. Nature Neuroscience, 24(1):129–139, 2020.
- Beyond fixed points: transient quasi-stable dynamics emerging from ghost channels and ghost cycles. arXiv, 2309.17201, 2023.
- Dynamics of a mutual inhibition circuit between pyramidal neurons compared to human perceptual competition. The Journal of Neuroscience, 41(6):1251–1264, 2020.
- Dissipative structures, machines, and organisms: A perspective. CHAOS, 27:104607, 2017.
- D. Krishnamurthy and M. Prakash. Emergent programmable behavior and chaos in dynamically driven active filaments. PNAS, 120(28):e2304981120, 2023.
- R. Legenstein and W. Maass. What makes a dynamical system computationally powerful. New directions in statistical signal processing: from systems to brain (ed. SS Haykin). Cambridge, MA: MIT Press, 2007.
- Dynamical synapses causing self-organized criticality in neural networks. Nature Physics, 3(12):857–860, 2007.
- Neutrophil swarms require LTB4 and integrins at sites of cell death in vivo. Nature, 498(7454):371–375, 2013.
- Real-time computing without stable states: A new framework for neural computation based on perturbations. Neural Computation, 14(11):2531–2560, 2002.
- B. J. MacLennan. Natural computation and non-turing models of computation. Theoretical Computer Science, 317(1-3):115–145, 2004.
- Canalization of gene expression and domain shifts in the drosophila blastoderm by dynamical attractors. PLoS Computational Biology, 5(3):e1000303, 2009.
- O. Mazor and G. Laurent. Transient dynamics versus fixed points in odor representations by locust antennal lobe projection neurons. Neuron, 48(4):661–673, 2005.
- J. Milnor. On the concept of attractor. Comm. Math. Phys., 99(2):177–195, 1985.
- T. Mora and W. Bialek. Are biological systems poised at criticality? J. Stat. Phys., 144:268–302, 2011.
- Cells use molecular working memory to navigate in changing chemoattractant fields. eLife, 11:e76825, 2022.
- A. Nandan and A. Koseska. Non-asymptotic transients away from steady states determine cellular responsiveness to dynamic spatial-temporal signals. PLOS Computational Biology, 19(8):e1011388, 2023.
- F. Neves and M. Timme. Computation by switching in complex networks of states. Phys. Rev. Lett., 109:018701, 2012.
- F. Neves and M. Timme. Bio-inspired computing by nonlinear network dynamics—a brief introduction. Journal of Physics: Complexity, 2(4):045019, 2021.
- D. J. Nicholson. Is the cell really a machine? J Theor Biol., 21(477):108–126, 2019.
- D. Papatsenko and M. Levine. The drosophila gap gene network is composed of two parallel toggle switches. PLoS ONE, 6(7):e21145, 2011.
- Quantitative landscapes reveal trajectories of cell-state transitions associated with drug resistance in melanoma. iScience, 25(12):105499, 2022.
- Dynamical encoding by networks of competing neuron groups: Winnerless competition. Physical Review Letters, 87(6), 2001.
- Metastable attractors explain the variable timing of stable behavioral action sequences. Neuron, 110(1):139–153.e9, 2022.
- How organisms come to know the world: Fundamental limits on artificial general intelligence. Frontiers in Ecology and Evolution, 9:806283, 2022.
- Control of cell state transitions. Nature, 609(7929):975–985, 2022.
- Frequency modulation of erk activation dynamics rewires cell fate. Molecular Systems Biology, 11(11):838, 2015.
- Control of lateral migration and germ cell elimination by the emphDrosophila melanogaster lipid phosphate phosphatases wunen and wunen 2. The Journal of Cell Biology, 171(4):675–683, 2005.
- Growth factor-induced MAPK network topology shapes erk response determining PC-12 cell fate. Nature Cell Biology, 9(3):324–330, 2007.
- J. Sardanyés and R. V. Solé. Delayed transitions in non-linear replicator networks: About ghosts and hypercycles. Chaos, Solitons and Fractals, 31(2):305–315, 2007.
- L. F. Seoane. Evolutionary aspects of reservoir computing. Philosophical Transactions of the Royal Society B: Biological Sciences, 374(1774):20180377, 2019.
- M. Sheetz. The cell as a machine. Cambridge University Press, 2018.
- S. Smale. On gradient dynamical systems. Annals of Mathematics, 74(1):199–206, 1961.
- R. Solé and L. F. Seoane. Evolution of brains and computers: The roads not taken. Entropy, 24(5):665, 2022.
- Interdependence between EGFR and phosphatases spatially established by vesicular dynamics generates a growth factor sensing and responding network. Cell Systems, 7(3):295–309.e11, 2018.
- Organization at criticality enables processing of time-varying signals by receptor networks. Molecular Systems Biology, 16(2):e8870, 2020.
- S. H. Strogatz. Nonlinear Dynamics and Chaos: With Applications to Physics, Biology, Chemistry and Engineering. Westview Press, 2000.
- D. Sussillo and O. Barak. Opening the black box: Low-dimensional dynamics in high-dimensional recurrent neural networks. Neural Computation, 25(3):626–649, 2013.
- Prevalence of unstable attractors in networks of pulse-coupled oscillators. Phys. Rev. Lett., 89:154105, 2002.
- I. Tsuda. Chaotic itinerancy as a dynamical basis of hermeneutics in brain and mind. Microcomputers and Attention, World Futures, 32:167, 1991.
- I. Tsuda. Dynamic link of memory-chaotic memory map in non equilibrium neural networks. Neural Networks, 5:313, 1992.
- I. Tsuda. Chaotic itinerancy and its roles in cognitive neurodynamics. Current Opinion in Neurobiology, 31:67–71, 2015.
- A. M. Turing. Systems of logic based on ordinals. Proceedings of the London Mathematical Society, Series 2, 45:161–228, 1939.
- A. M. Turing. The chemical basis of morphogenesis. Bulletin of mathematical biology, 52:153–197, 1990.
- A. M. Turing. Intelligent Machinery (1948), in: The Essential Turing. Oxford University Press, 2004.
- Classification of transient behaviours in a time-dependent toggle switch model. BMC Systems Biology, 8:1–19, 2013.
- Cell commitment motif composed of progenitor-specific transcription factors and mutual-inhibition regulation. IET Systems Biology, 8(4):129–137, 2014.
- The dynamics of pattern matching in camouflaging cuttlefish. Nature, 619(7968):122–128, 2023.
Paper Prompts
Sign up for free to create and run prompts on this paper using GPT-5.
Top Community Prompts
Collections
Sign up for free to add this paper to one or more collections.