- The paper introduces a self-supervised deep learning framework that leverages fine-grained segmentation feature maps to enhance registration accuracy.
- The method employs a UNet-based segmentation network and k-means clustering to generate anatomical pseudo-labels, eliminating the need for manual annotations.
- Empirical evaluations demonstrate improved performance on the ANHIR dataset and brain MRI, achieving a median rTRE of 0.00099 and competitive segmentation metrics.
Deep Learning Registration of Histopathology Images Using Fine-Grained Structural Feature Maps
Introduction
The registration of histopathology images underpins the morphological comparison of tissue sections required for accurate pathological interpretations. Variations in tissue preparation, staining protocols, and sample handling induce nonlinear, elastic deformations that complicate inter-image alignment. Traditional registration approaches, often reliant on iterative optimization and intensity-based metrics, encounter scalability, robustness, and efficacy issues when faced with diverse tissue morphologies and repetitive textures. Recent deep learning (DL) models have accelerated registration workflows, but their dependence on manually annotated segmentation maps limits practicality, particularly in clinical scenarios where expert-provided structural delineations are unavailable.
This paper introduces a self-supervised, annotation-independent method that integrates structural information gleaned from unsupervised segmentation feature maps into a DL-based registration pipeline. Unlike prior methods, it leverages segmentation features from a pre-trained segmentation network coupled with k-means clustering to generate fine-grained anatomical maps, enhancing registration accuracy without requiring manual labels.
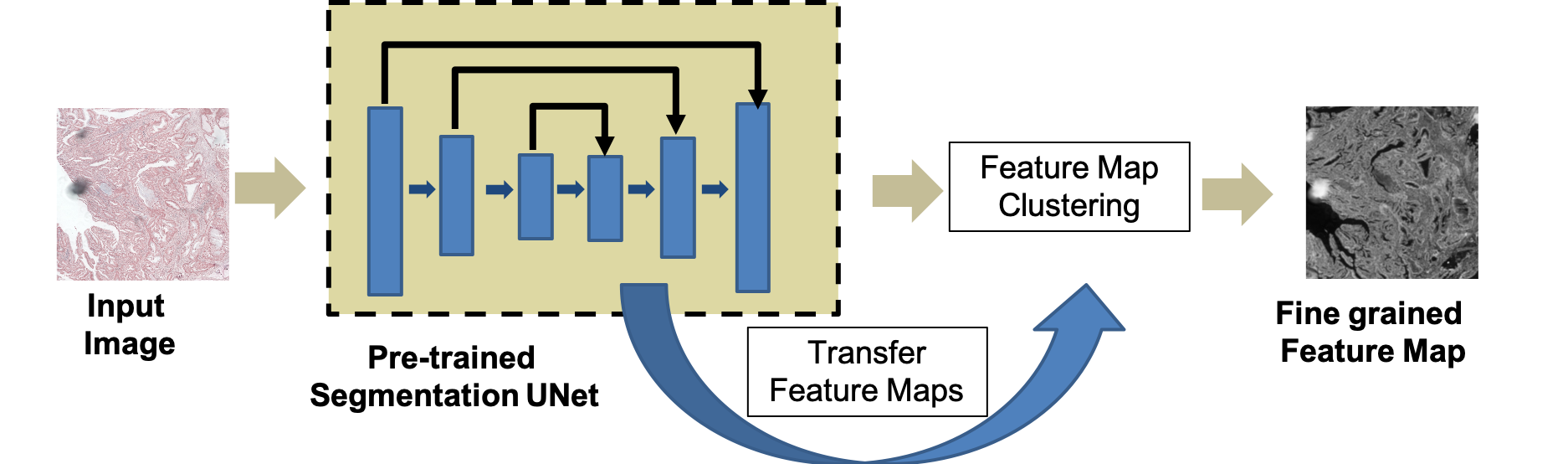
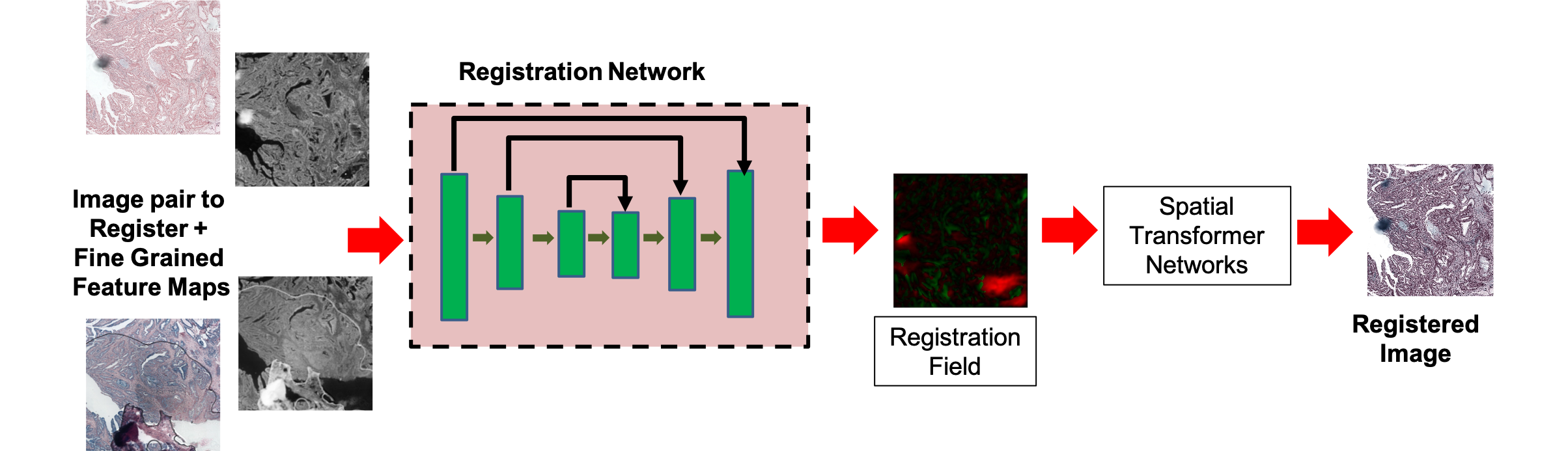
Figure 1: Workflow for generation of fine-grained self-supervised feature maps and integration within the registration pipeline.
Methodological Framework
The proposed workflow commences with the reference and floating images processed by a Fine-Grained Segmentation Network (FGSN) based on a UNet backbone pre-trained on the Glas dataset for histopathological segmentation. Encoder and decoder feature maps are concatenated and fused at multiple scales, capturing both low-level and high-level structures. Subsequent k-means clustering assigns granular pseudo-labels, with cluster counts (k) dynamically refined via the Gap Statistic method to optimize structural segmentation, yielding dense, fine-grained anatomical maps.
Segmented feature maps from both images are input to a UNet-like registration network. The deformation field φ is estimated, transforming the floating image via a Spatial Transformer Network (STN) to yield the registered output. The training loss amalgamates image similarity (mean squared error and local cross-correlation), smoothness regularization, and a segmentation loss defined as the pixel-wise mean squared error between self-supervised segmentation maps of the reference and registered images:
Lus=Lsim+λ1Lsmooth+λ2Lseg
This approach is architecture-agnostic but is demonstrated with VoxelMorph for benchmarking, replacing its manual segmentation maps with self-supervised analogs.
Empirical Evaluation and Results
Evaluation on the ANHIR challenge dataset, encompassing 481 histopathology image pairs of diverse tissue types, employed the normalized target registration error (rTRE) metric for quantitative assessment. The proposed SR-Net achieved a median rTRE of 0.00099 across all tissue types, outperforming top-ranked challenge methods and demonstrating significant gains over ablated variants lacking segmentation loss (SR-NetwLSeg). Statistical tests (paired Wilcoxon, p=0.003) confirmed the substantive impact of self-supervised segmentation.
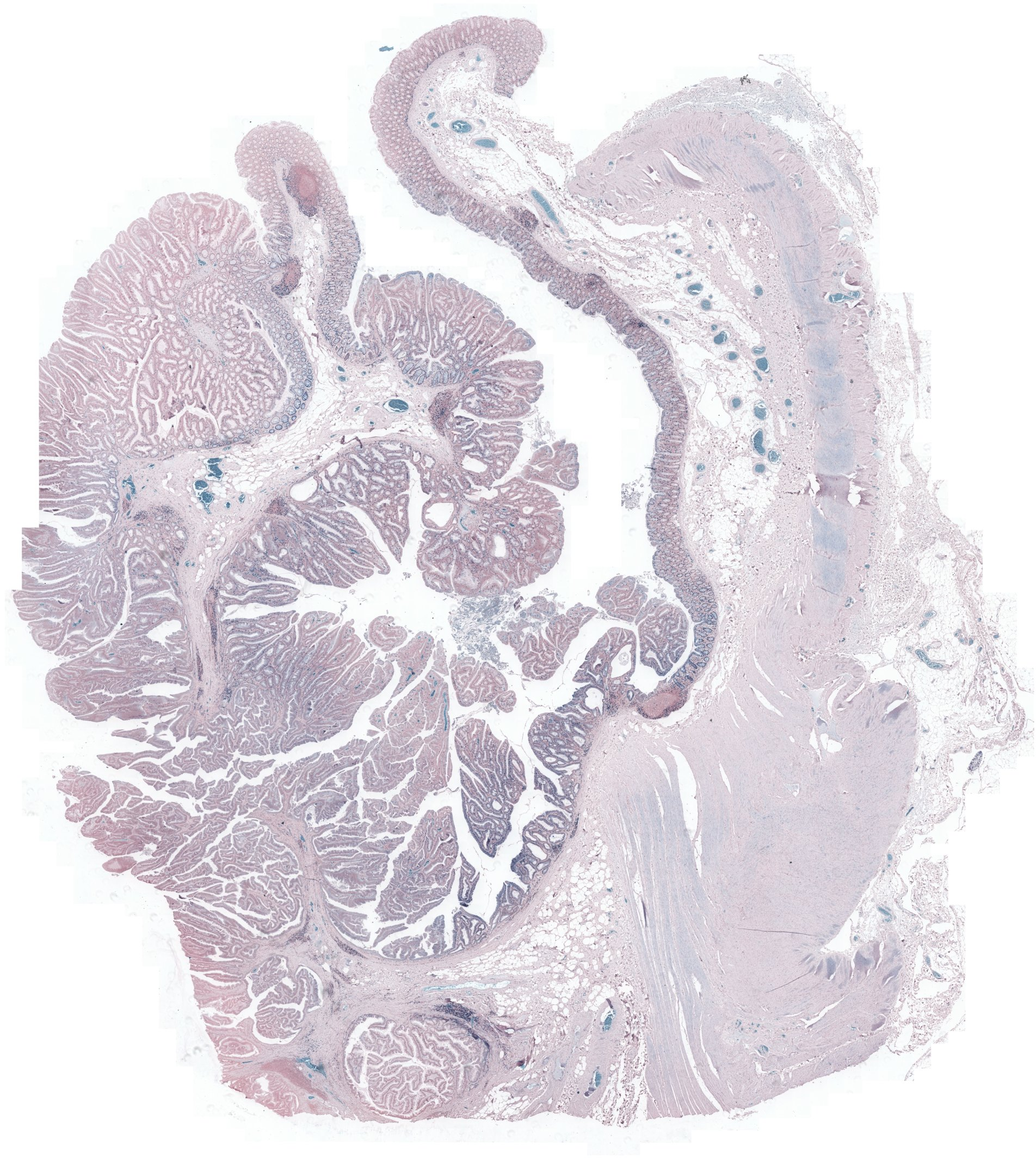
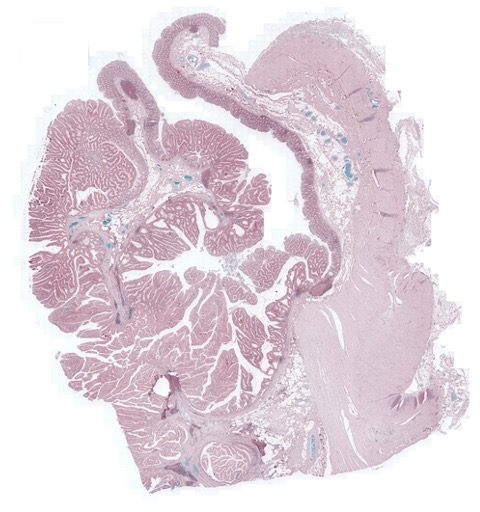
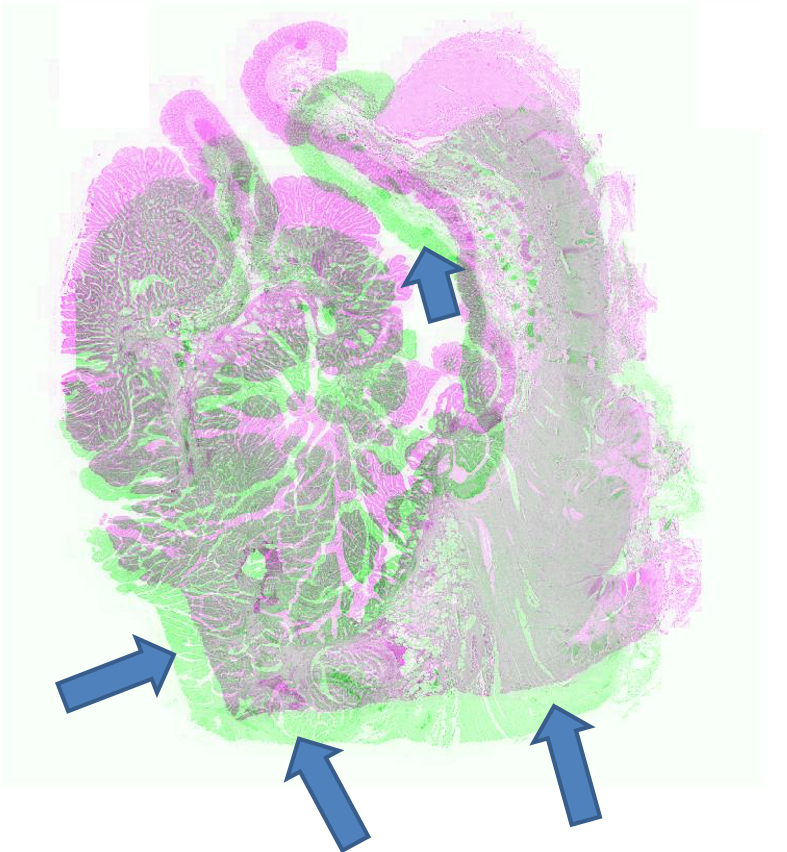
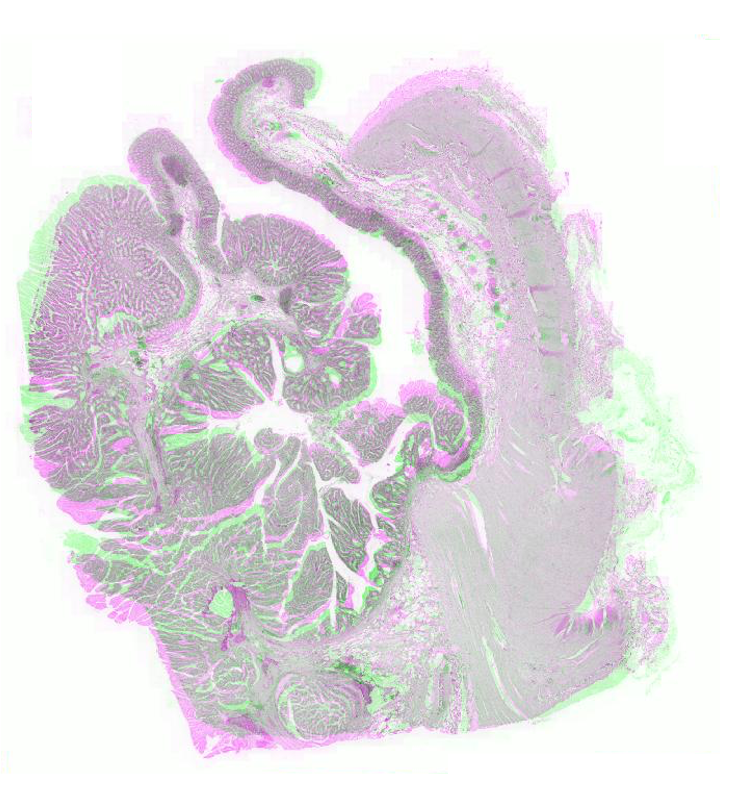
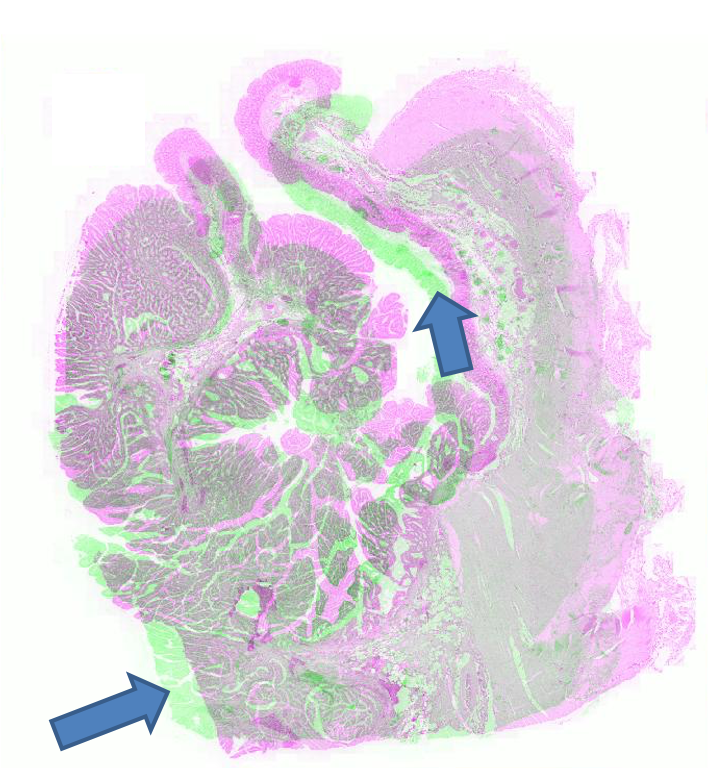
Figure 2: Example pathology image registration — misalignment pre-registration, results post-registration with SR-Net and SR-NetwLSeg, highlighting improved structural congruence.
Brain MRI Atlas Registration
SR-Net was applied to a subset of ADNI-1 brain MRI images with simulated elastic deformations. Dice Metric (DM) and Hausdorff distance (HD95) evaluated segmentation overlap and spatial accuracy. SR-Net achieved DM 79.2%, closely matching VoxelMorph (79.5%) and surpassing ablated models, with no statistically significant difference (p=0.031), thus validating the efficacy of self-supervised maps in replacing manual counterparts.
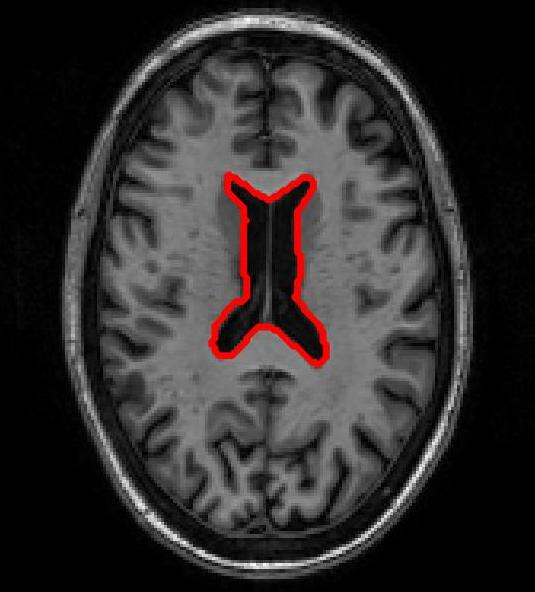
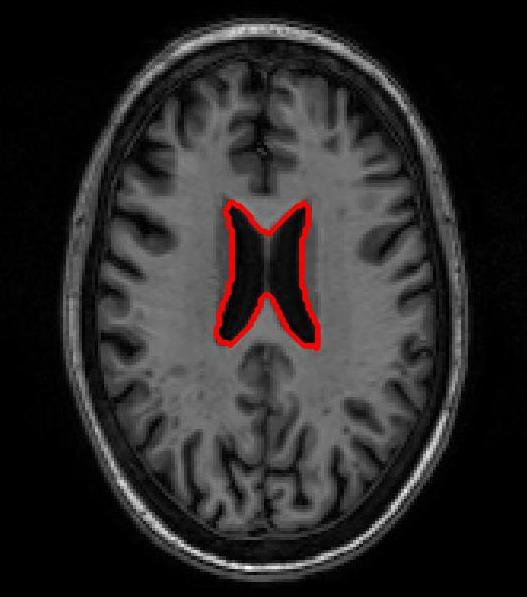
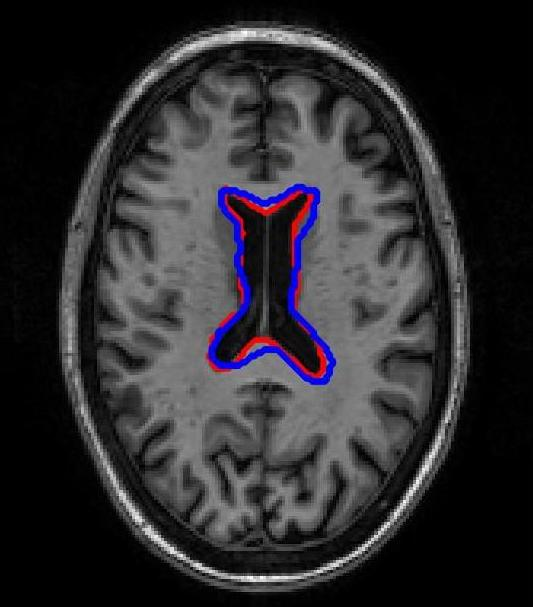
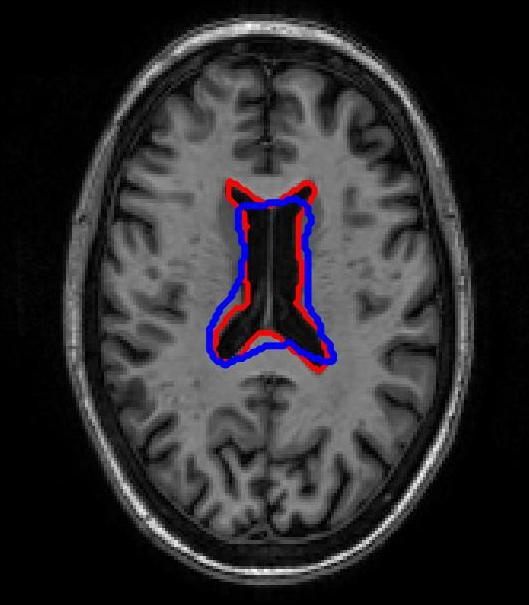
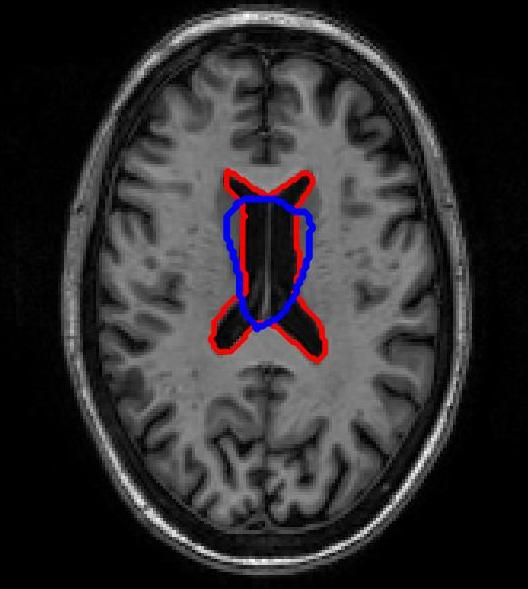
Figure 3: Atlas-based brain MRI registration — comparison of manual segmentation (red) and registered mask (blue) using SR-Net, VoxelMorph, and SR-NetwLSeg.
Ablation Analysis
Ablation studies isolating loss components (MSE, CC, segmentation) confirmed the dominant role of segmentation-derived structural features in reducing registration error and aligning critical anatomical structures, with adverse effects observed when these were omitted.
Implications and Future Directions
The presented registration paradigm, leveraging self-supervised fine-grained feature maps, offers substantial practical advantages for histopathology and neuroimaging: it circumvents the need for manual segmentations, generalizes across tissue types, and supports deployment in real-world clinical workflows where annotated datasets are scarce. Theoretical implications include further unification of segmentation and registration tasks within DL frameworks and extension toward fully unsupervised structure-aware learning.
Potential future advancements may involve:
- Adapting the segmentation network backbone for cross-modality and multi-scale registration.
- Integrating other clustering or representation learning techniques for improved anatomical delineation.
- Evaluating framework robustness on whole-slide digital pathology images and volumetric scans.
- Extending structural loss functions to exploit higher-order anatomical relationships.
Conclusion
This work establishes that self-supervised segmentation maps, generated via feature clustering from a pre-trained segmentation network, supply sufficient structural guidance to achieve registration performance comparable with manual segmentation-based DL methods. The proposed approach enhances registration accuracy, promotes scalability, and lays groundwork for annotation-independent medical image analysis pipelines, with broad applicability across histopathological and neuroimaging domains.
(2007.02078)











