- The paper introduces SF2Former, a dual-branch transformer model that fuses spatial and frequency features from MRI for accurate ALS identification.
- It leverages Vision Transformers for spatial encoding and the Global Filter Network for frequency analysis, outperforming traditional CNNs.
- The framework employs a majority voting scheme and integrates multi-center MRI modalities to enhance robustness in ALS diagnosis.
Introduction
The study introduces SF2Former, a novel framework designed to differentiate patients with Amyotrophic Lateral Sclerosis (ALS) from healthy controls using MRI data from multiple centers. The research leverages the capabilities of Vision Transformers (ViTs) to distinguish subtle neurodegenerative changes in ALS, often poorly captured by traditional convolutional networks due to insignificant structural variations in neuroimaging data. By integrating features from both spatial and frequency domains—where MRI captures raw data before conversion—the framework capitalizes on the advantages of frequency domain data representation. This section summarizes the notable contributions of SF2Former, including its effective capturing of discriminative features and the implementation of a majority voting scheme for enhanced classification decisions.
Methodology
The SF2Former architecture combines spatial and frequency domain features using a dual-branch approach (Figure 1). The left branch encodes spatial features employing ViT, known for its capability to model long-range dependencies through self-attention mechanisms. Simultaneously, the right branch captures frequency domain interactions using the Global Filter Network (GFNet), applying Fourier transforms to better exploit inherent MRI data properties.
After preprocessing using tools like FreeSurfer and FSL, coronal slices are selected based on empirical analysis. The methodology also incorporates a linear fusion module to integrate spatial and frequency representations, thus refining the aggregated signal for final classification. Majority voting across coronal slices ensures robustness against false positives, effectively accommodating slice variation from single subject scans.
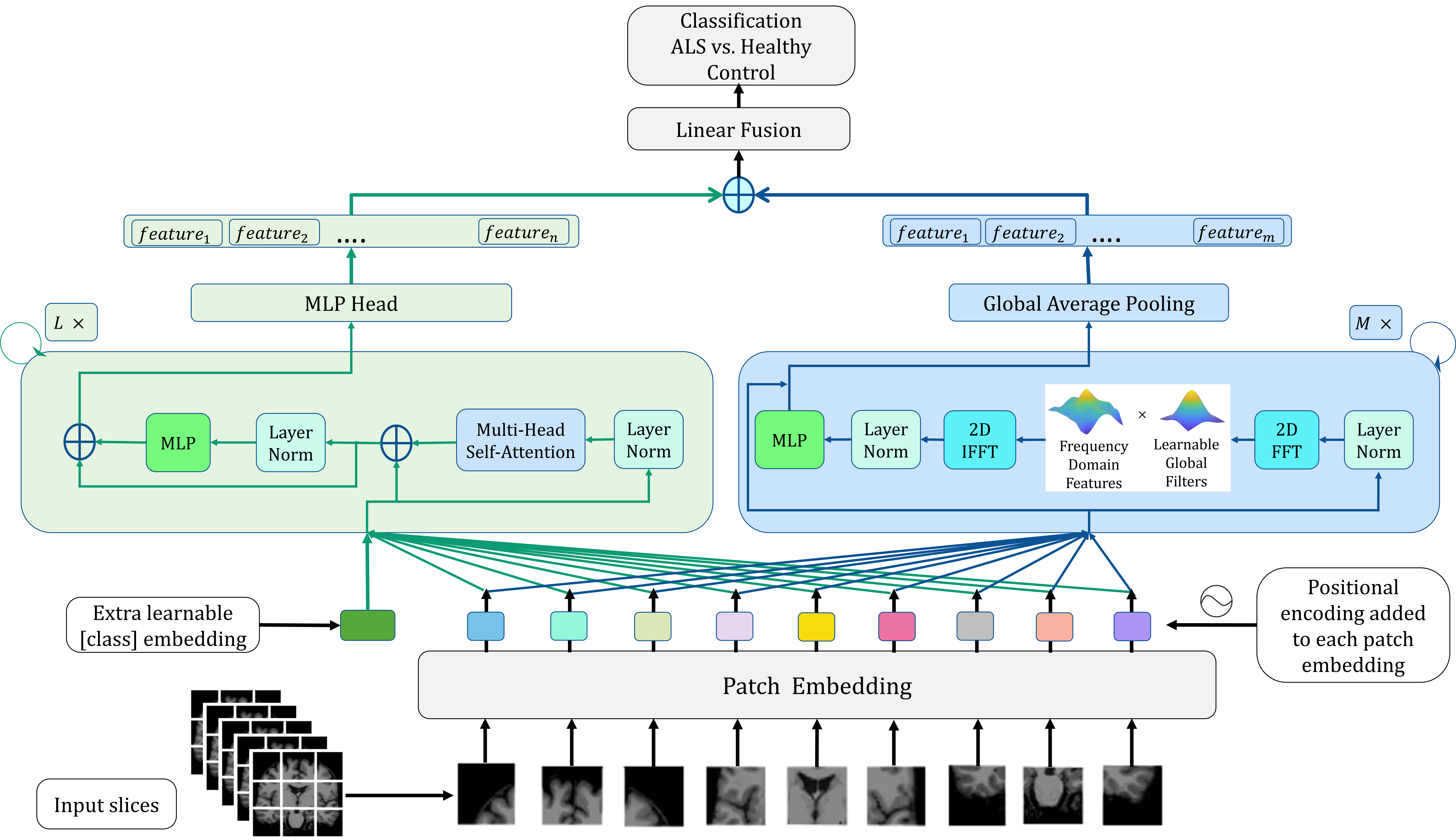
Figure 1: Proposed SF2Former architecture. The left branch encodes features from the spatial domain, whereas the right branch encodes features from the frequency domain. The linear fusion module assembles the classification decision for each 2D slice.
Experimental Results
Experimental validation employed multi-modal datasets from CALSNIC, focusing on three MRI modalities: T1-weighted, R2*, and FLAIR. The system demonstrated superior performance, exhibiting improved classification accuracy over existing CNN-based approaches (Table 1). Key metrics included Accuracy (ACC), Sensitivity (SEN), and Specificity (SPE), all indicating SF2Former's robustness in multi-modal, multi-center contexts.
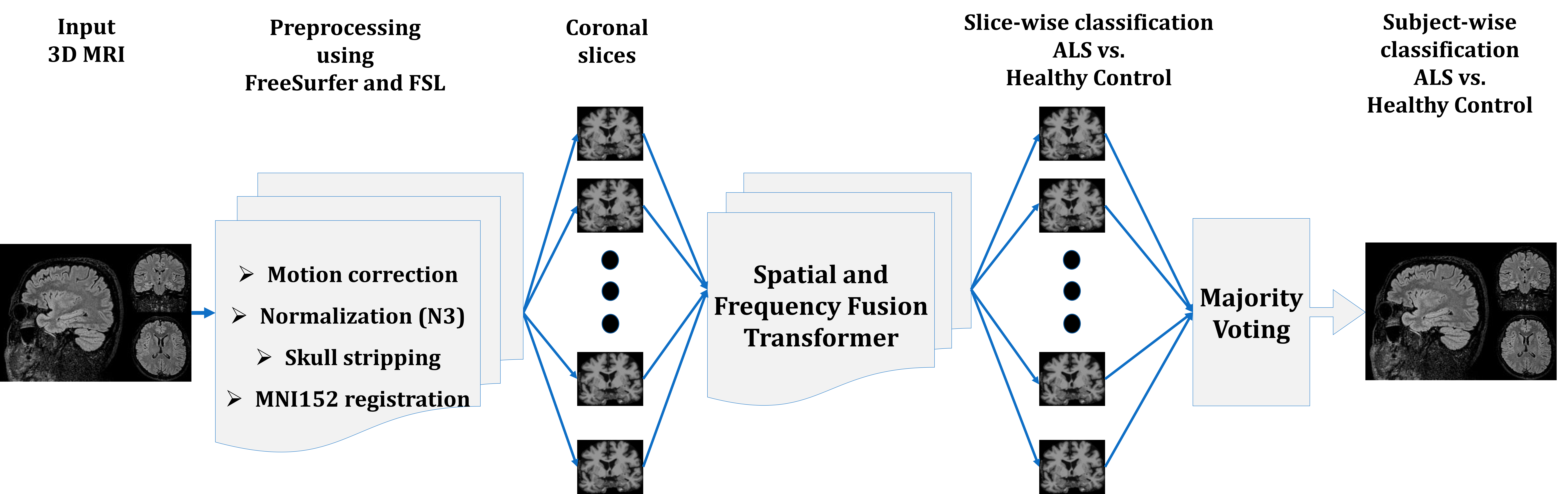
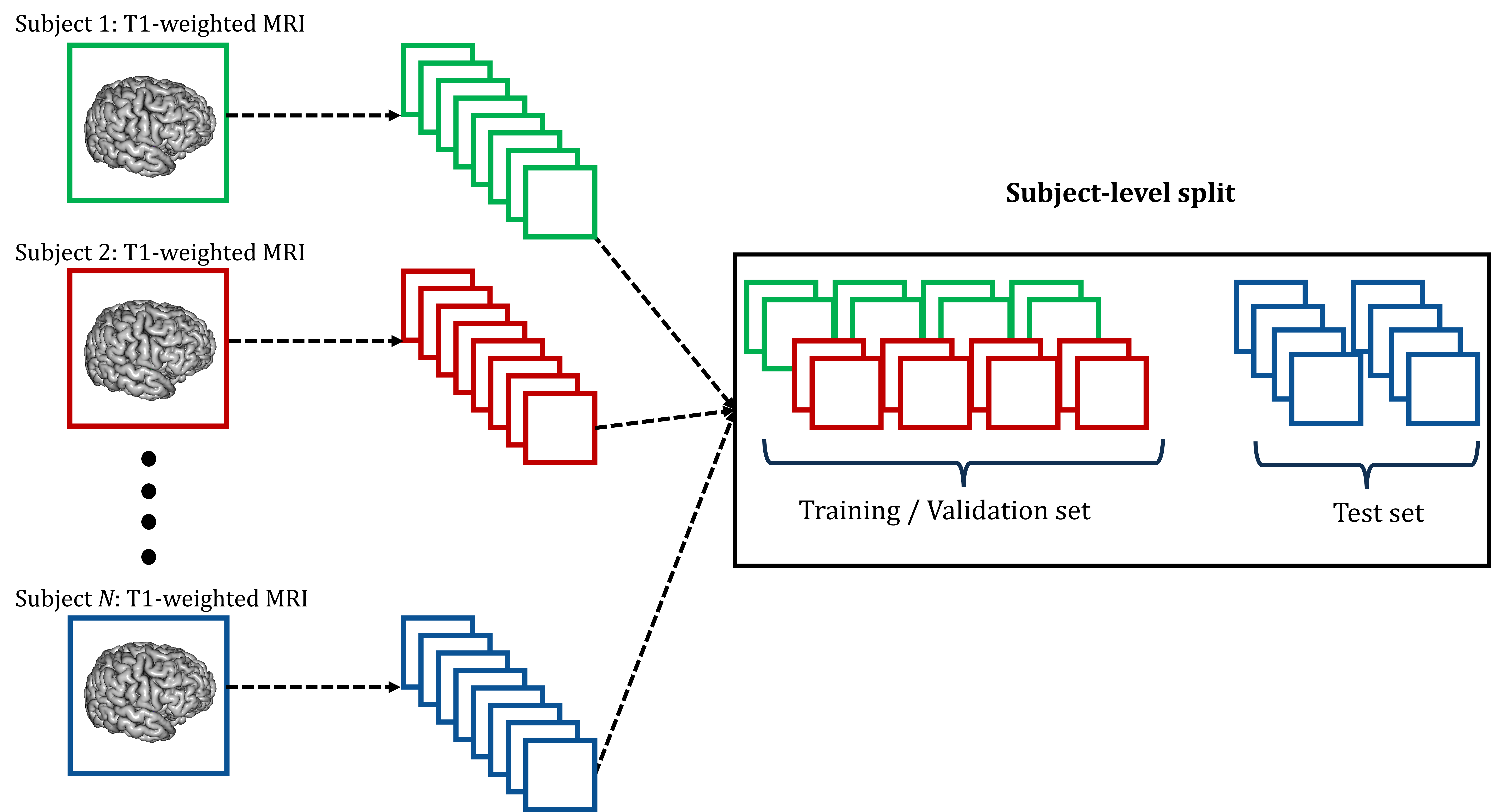
Figure 2: The overall workflow of the proposed stages.
Figure 3: Subject-level split process for the data to train our proposed model.
Ablation and Comparative Analysis
The ablation study underscores the contribution of each architectural and methodological component to the overall performance. Removal of data normalization, augmentation, or transfer learning significantly diminished accuracy. Furthermore, comparative analysis revealed SF2Former’s distinct advantage over 2D and 3D CNN architectures, as well as previous texture-based methods, particularly with T1-weighted images.
Discussion and Implications
SF2Former's deployment of multi-domain feature extraction positions it as a notable advancement in ALS imaging biomarker research. The dual-branch transformer approach is especially appropriate for handling the multi-slice nature of MRI data while remaining robust across various imaging modalities and scanner types. The integrated majority voting scheme reliably accounts for slice variability inherent in scanned subjects.
The implications of this research are significant, fostering the future utility of MRI as a diagnostic biomarker for neurodegenerative disorders. Prospective work could further incorporate clinical and functional imaging data, enhancing diagnostic accuracy and generalization across neurodegenerative conditions.
Conclusion
SF2Former exemplifies a sophisticated approach to ALS classification using MRI, effectively overcoming limitations of previous CNN-centric models. This framework’s adaptability to different imaging modalities and its robustness in multi-center datasets make it a promising tool for ALS diagnosis and progression monitoring. Future studies could extend its applicability to integrate alternative neuroimaging and clinical data, offering expansive avenues for diagnostic exploration.


