- The paper pioneers a domain-specific CLIP-like architecture that integrates semantic-aware contrastive pre-training with a diagnostic taxonomy.
- It presents the US-365K dataset and a structured Ultrasonographic Diagnostic Taxonomy to standardize and improve ultrasound image-text alignment.
- Empirical results demonstrate superior diagnostic classification, retrieval performance, and robust zero-shot transfer across ultrasound tasks.
Semantic-Aware Contrastive Pre-training for Ultrasound Image-Text Understanding: An Expert Analysis of Ultrasound-CLIP
Introduction
"Ultrasound-CLIP: Semantic-Aware Contrastive Pre-training for Ultrasound Image-Text Understanding" (2604.01749) addresses core limitations in multi-modal ultrasound analysis by introducing a unified diagnostic taxonomy, a large-scale dedicated ultrasound dataset (US-365K), and a domain-specific CLIP-like architecture. The framework strategically integrates semantic-aware supervision and structured graph-based reasoning to achieve enhanced discrimination and generalization in image-text understanding—outperforming general medical VLP models, especially in zero-shot and transfer settings.
Motivation and Context
Although ultrasound is the most widely deployed radiology imaging modality outside plain X-ray, it remains grossly underrepresented in the construction and benchmarking of large-scale vision-language pre-trained (VLP) models. Existing datasets such as PMC-15M, ROCO, and MedTrinity-25M contain less than 5% ultrasound content, introducing a modality gap that degrades the clinical performance of general-purpose medical vision-LLMs.
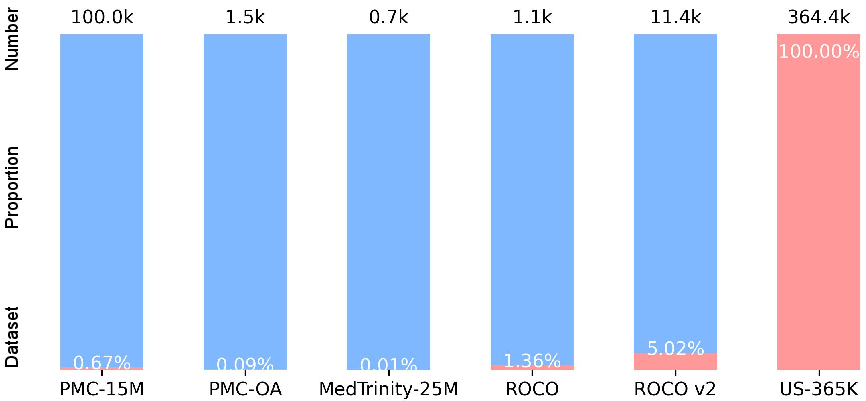
Figure 1: US-365K is the first large-scale, 100% ultrasound-focused image-text benchmark, in sharp contrast to the modality imbalance in existing datasets.
Ultrasonographic Diagnostic Taxonomy (UDT)
A critical factor limiting text-image understanding in sonography is the semantic and anatomical heterogeneity specific to ultrasound images and reports. The paper introduces the Ultrasonographic Diagnostic Taxonomy (UDT), a domain-specific hierarchical knowledge framework:
- Ultrasonographic Hierarchical Anatomical Taxonomy (UHAT): Encodes 9 body systems and 52 organs, forming a two-level classification spanning the coverage of clinical ultrasound.
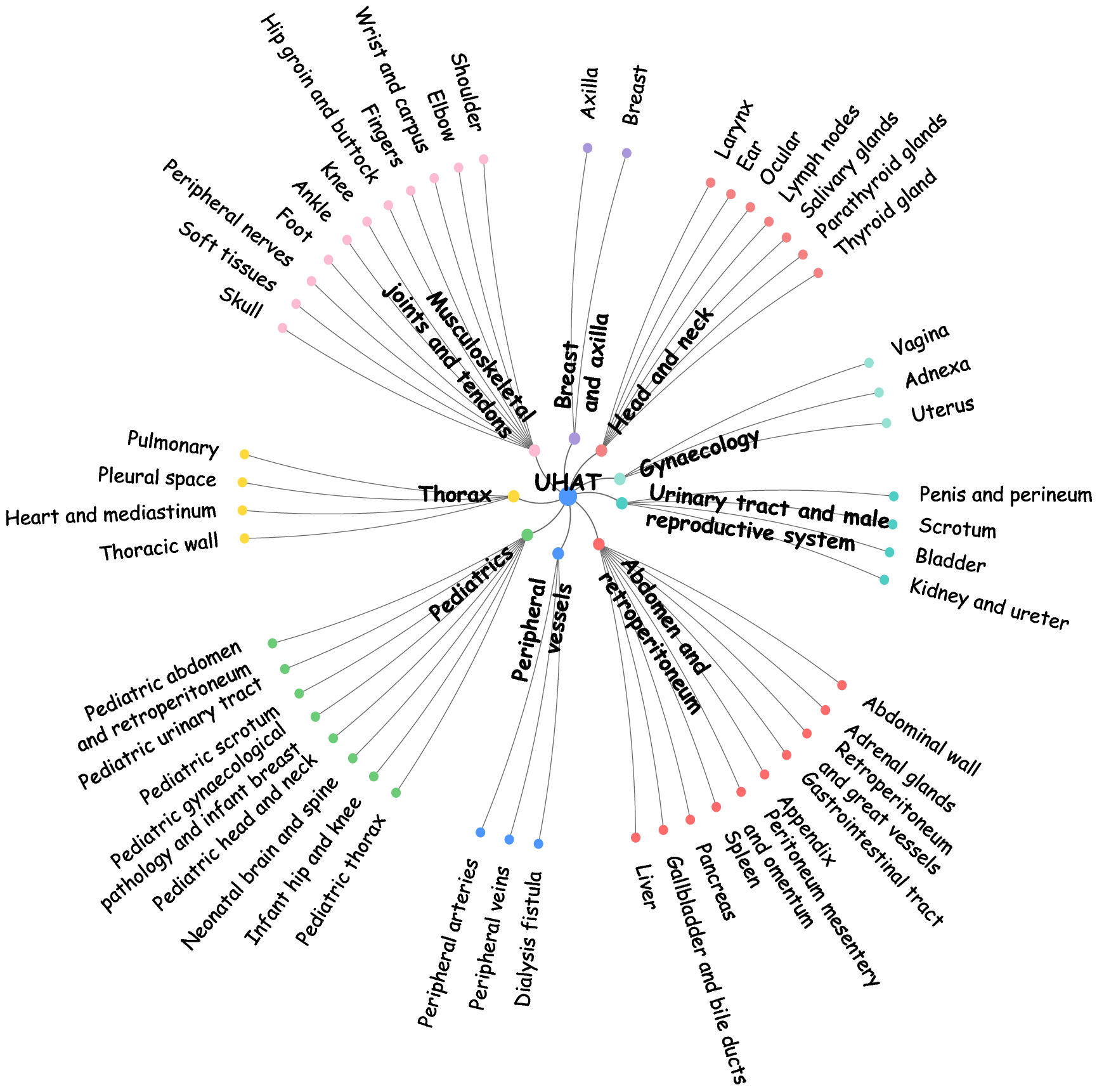
Figure 2: Visualization of UHAT anatomical hierarchy spanning major body systems and organs in US-365K.
- Ultrasonographic Diagnostic Attribute Framework (UDAF): Formalizes nine standardized diagnostic dimensions (e.g., diagnosis, shape, margins, echogenicity, vascularity), each with a clinically curated vocabulary.
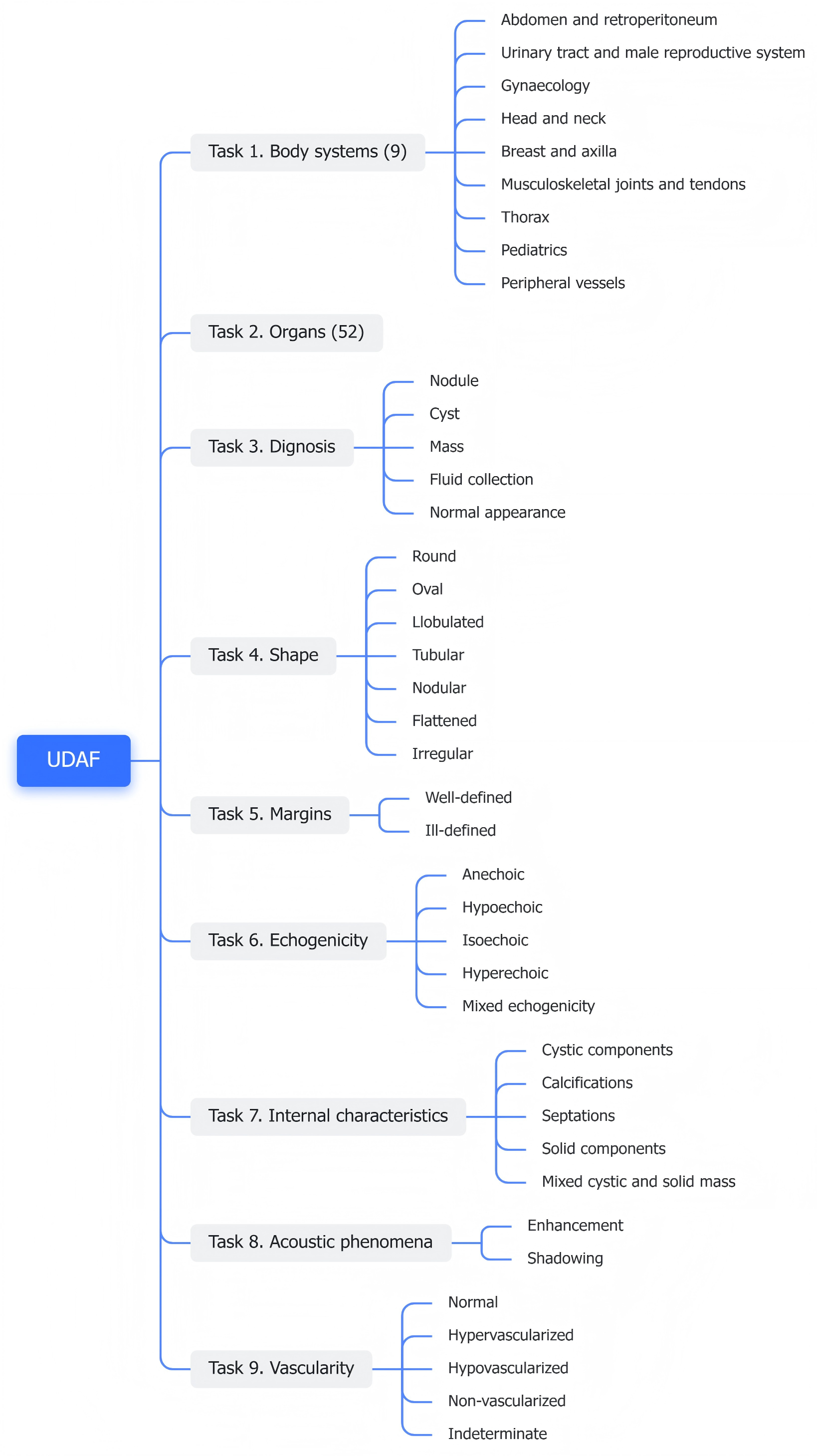
Figure 3: UDAF schema decomposes the diagnostic report into nine structured dimensions, supporting both normalization and structured label construction.
UDT serves as the semantic backbone for both data annotation and model construction, enabling standardization across highly variable clinical narratives.
US-365K: Dataset Construction and Analysis
The US-365K dataset is curated from five complementary open-access resources, resulting in 364,365 image-text pairs after strict anatomical QA, de-duplication, and expert verification. The dataset encompasses:
- Broad coverage: 9 systems, 52 organs (defined by UHAT).
- Diagnostic diversity: Long-tail distribution across major and minor pathologies.
- Linguistic heterogeneity: Captions normalized using UDAF for robust semantic representation, yet reflecting the variability of natural-language clinical reporting.
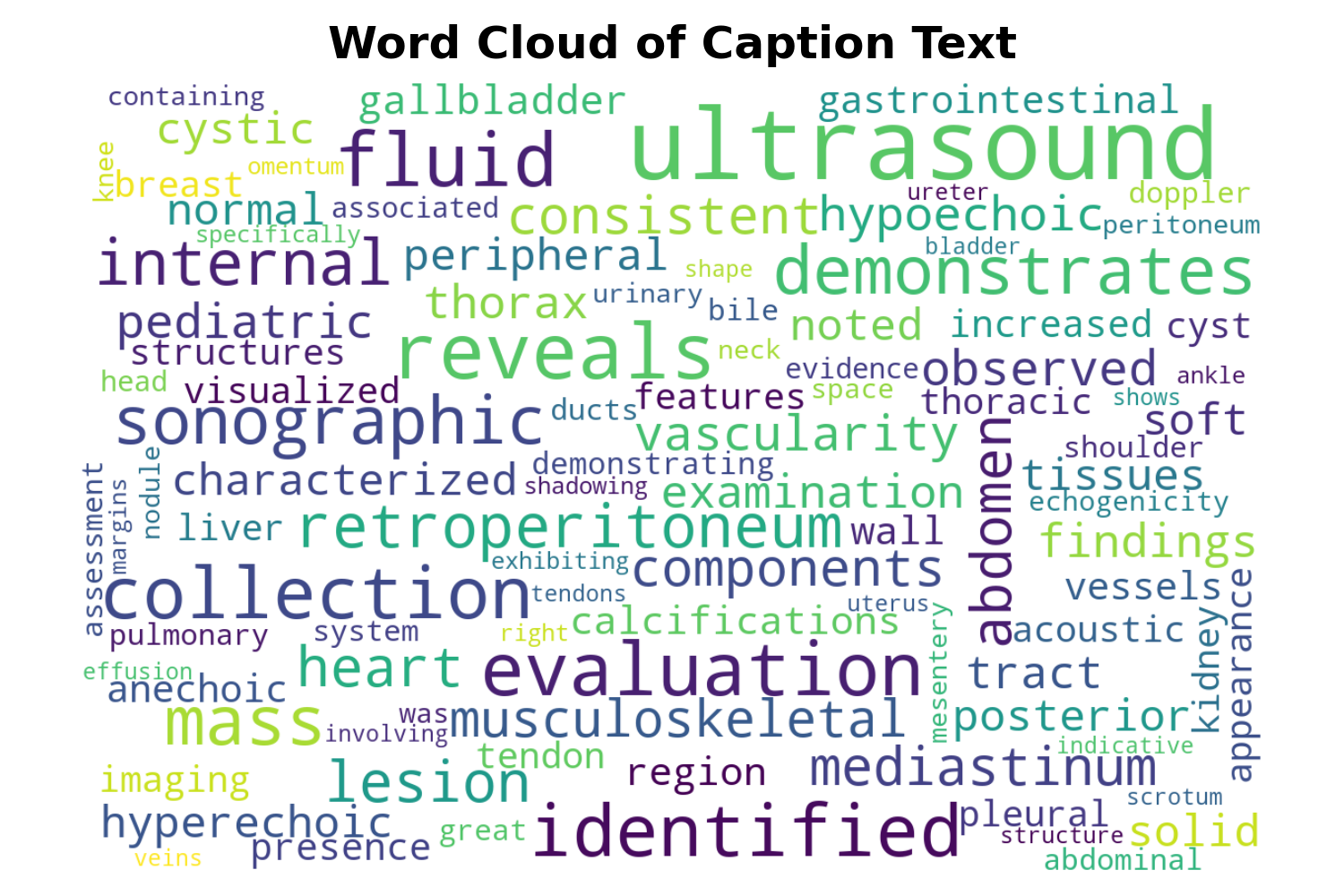
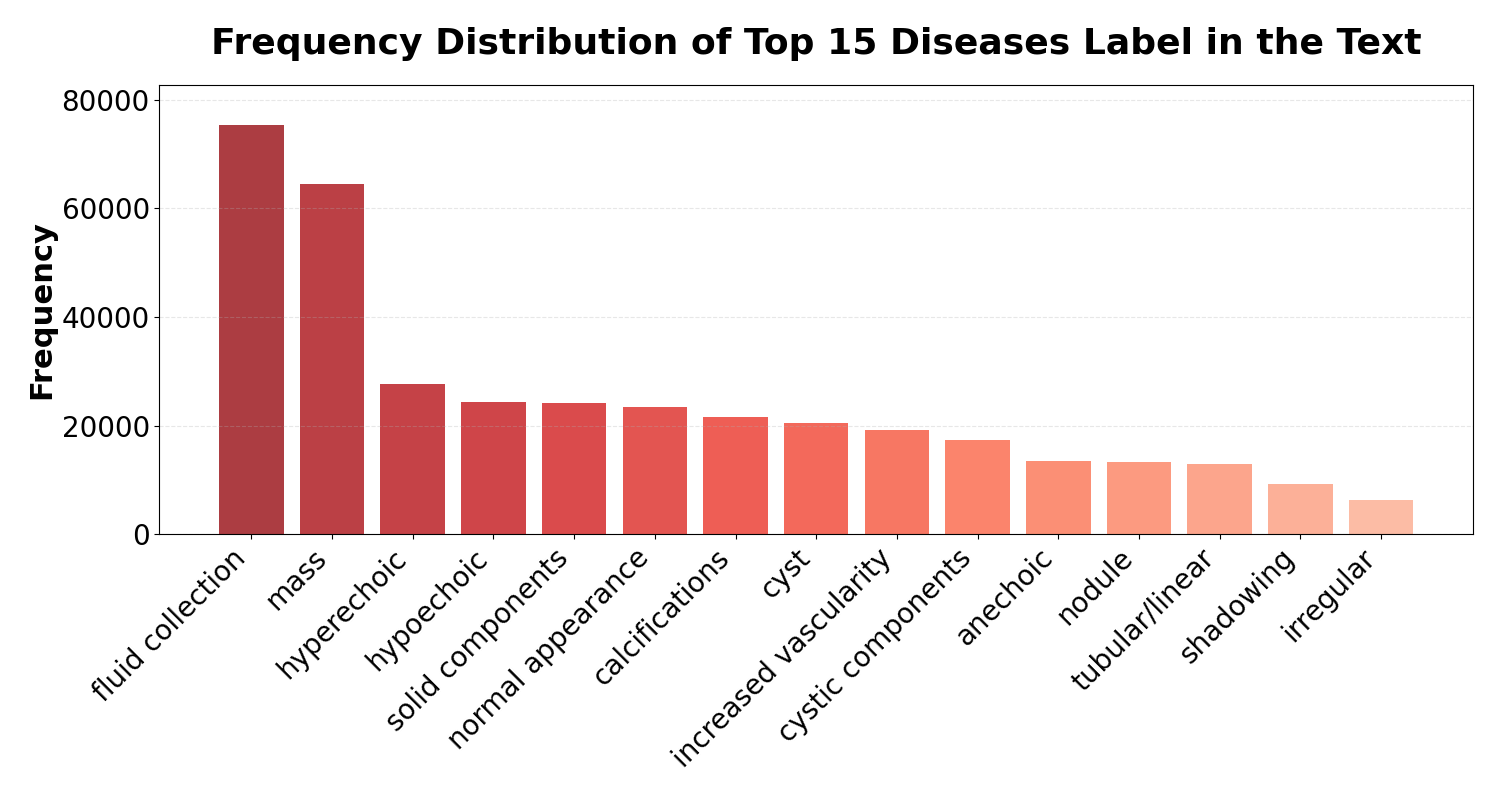
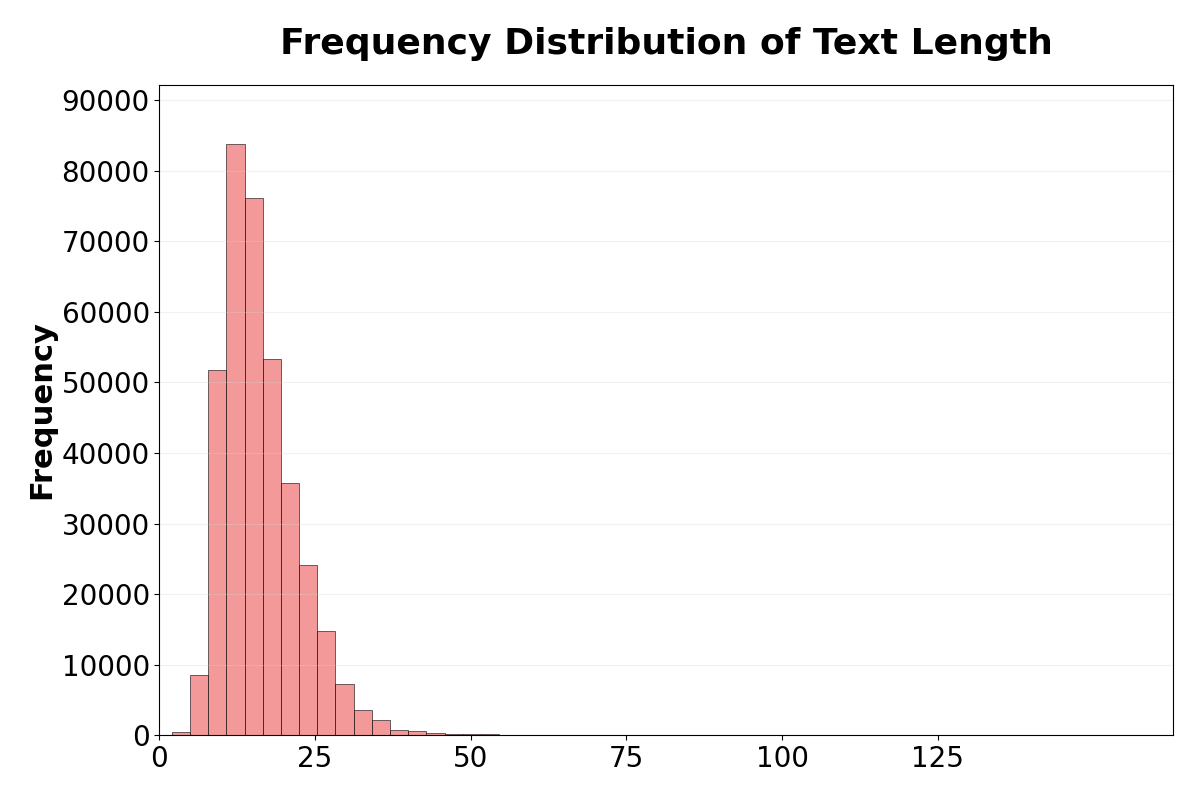

Figure 4: (a) Frequency distribution of findings; (b) Caption vocabulary word cloud; (c) Caption length distribution, confirming both conciseness and diversity in real clinical text.
The data construction pipeline leverages prompt-based LLM annotation for reliability and compositional consistency under the UDT framework.
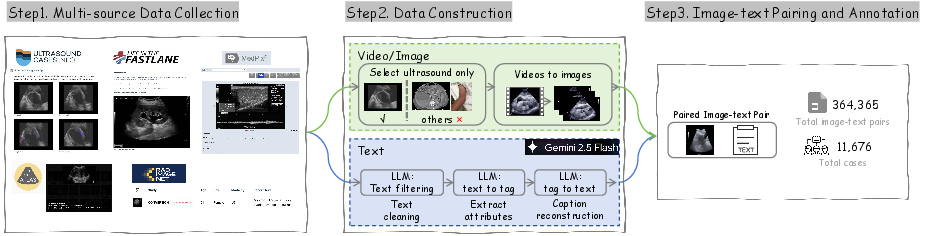
Figure 5: US-365K data processing pipeline with multi-source ingestion, attribute extraction, and fine-grained text-image alignment.
Ultrasound-CLIP: Model Architecture
Structural Overview
Ultrasound-CLIP introduces an architecture specifically optimized for ultrasound data:
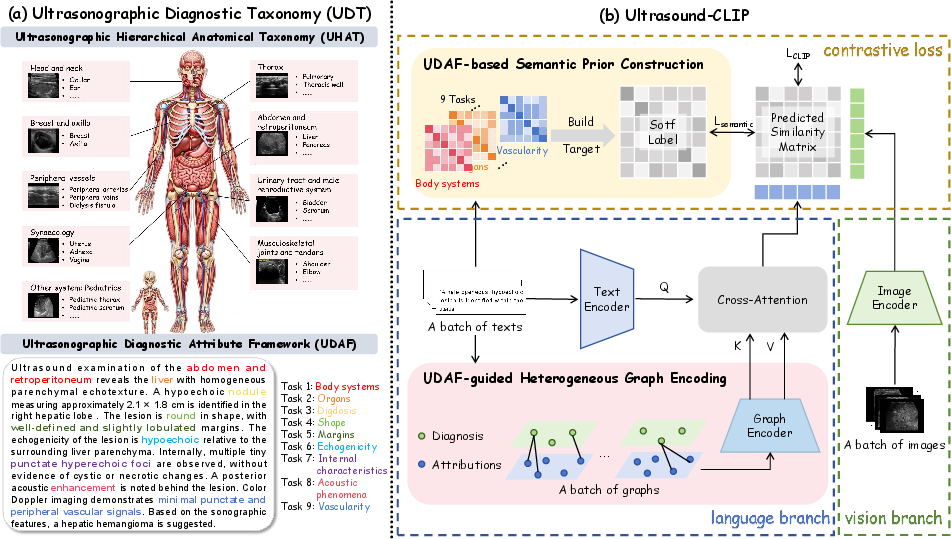
Figure 6: The UDT serves as semantic foundation, while Ultrasound-CLIP leverages both semantic priors and a UDAF-guided graph encoder for robust, structured multimodal reasoning.
The framework consists of:
- Dual Encoder (CLIP-style): ViT for images, BioClinical-BERT for text.
- Semantic Priors: Soft label matrices (UDAF-guided) encode non-binary similarity in batch training, mitigating semantic ambiguity.
- Heterogeneous Graph Encoder: Converts UDAF-aligned diagnostic attributes into sample-specific heterogeneous graphs that model intra-text dependencies; graph embeddings are fused with text representation via multi-head cross-attention.
Optimization Strategy
Learning is driven by a dual-objective:
- Contrastive Loss LCLIP: Standard CLIP-style bidirectional alignment.
- Semantic Alignment Loss Lsemantic: MSE and KL-regularized soft alignment with UDAF-based priors.
This dual mechanism enables Ultrasound-CLIP to resolve annotation ambiguity and enforce semantically consistent cross-modal clustering at finer clinical granularity.
Empirical Results: Robustness, Generalization, and Interpretability
On US-365K, Ultrasound-CLIP demonstrates strong and consistent improvements over both generalist (e.g., CLIP, MetaCLIP, SigLIP) and medical VLP models (PMC-CLIP, UniMed-CLIP, BiomedCLIP):
- Multi-attribute Diagnostic Classification: AvgAcc = 59.61%, outperforming BiomedCLIP (33.81%) by 25+ points; particularly strong on fine-grained diagnostic dimensions ("Diagnosis": 64.05% vs. 39.40%).
- Image-Text Retrieval: R@10 (I2T): 0.3745 vs. 0.3011 (best baseline). Both attribute-aware semantic loss and graph encoder offer substantial complementary gains.
Generalization: Downstream Task Transfer
Across four challenging downstream datasets (BUSBRA, GIST514-DB, BreastMNIST, Breast), the model achieves superior or competitive accuracy in zero-shot (ZS), linear-probe (LP), and fine-tuning (FT) settings:
|
ZS |
LP |
FT |
| Avg.% |
54.42 |
75.40 |
84.23 |
These results establish clear domain transfer advantages attributed to the ultrasound-specific pre-training and structured supervision, with negligible reliance on model parameter scaling (Ultrasound-CLIP is smaller and faster than SigLIP).
Ablation Study
Removal of either semantic loss or the graph encoder yields substantial performance drops; the full model significantly outpaces reduced variants, empirically verifying the necessity of both structured priors and explicit attribute reasoning for ultrasound.
Interpretability and Case-Level Clinical Reasoning
- t-SNE Visualization: UDAF-guided graph enhancement yields more semantically coherent text embedding clusters compared to non-graph baselines.
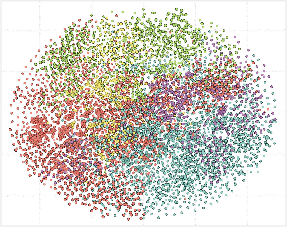
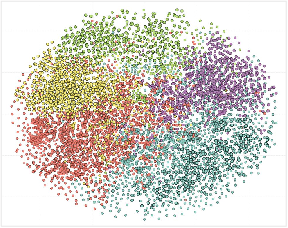
Figure 7: t-SNE of text embeddings: clusters after graph encoding are more distinct and diagnosis-specific.
- Case Studies (Clinical Coherence and Probabilistic Diagnosis): The model captures both primary and secondary diagnostic hypotheses, reflecting clinical uncertainty. Supporting attribute predictions are consistently aligned with ground truth, indicating learned interpretability and robust reasoning.
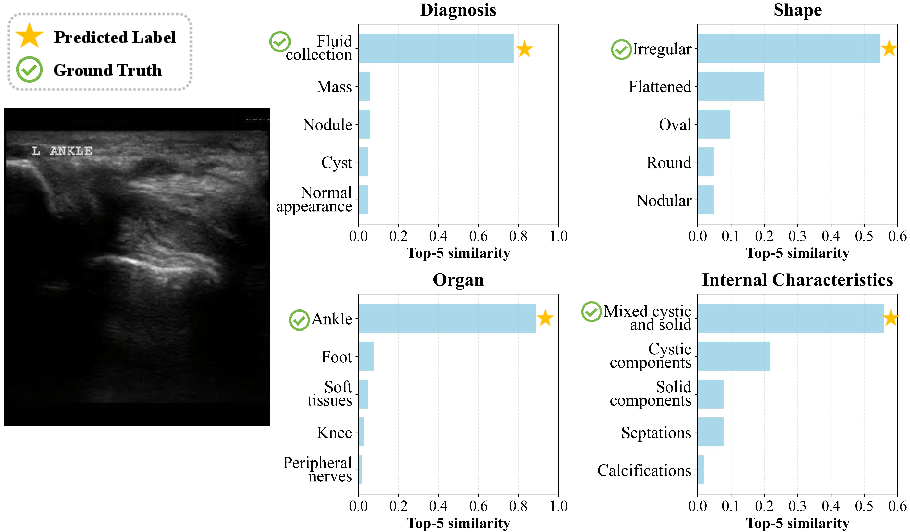
Figure 8: Visualization of model predictions for left ankle ultrasound; outputs reflect high clinical interpretability.
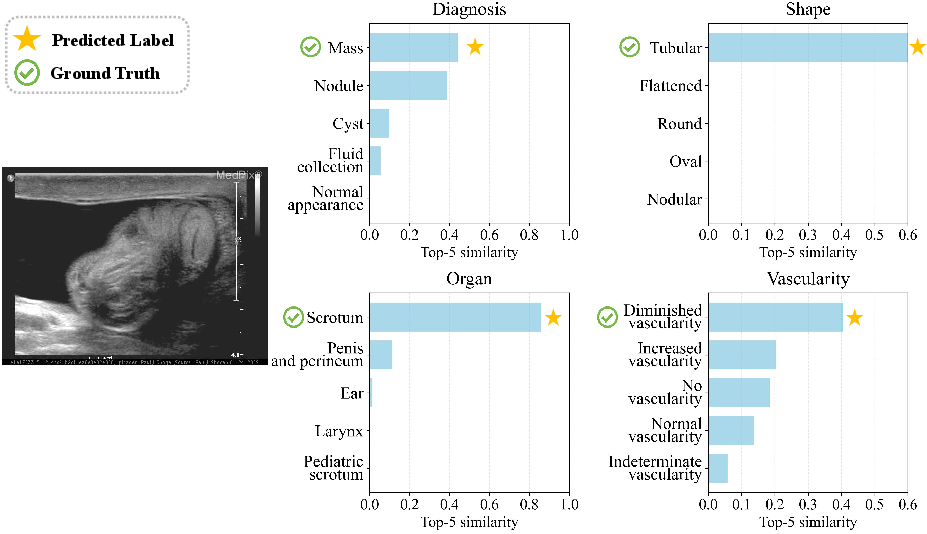
Figure 9: Model aligns with probabilistic evidence in ambiguous or overlapping diagnoses.
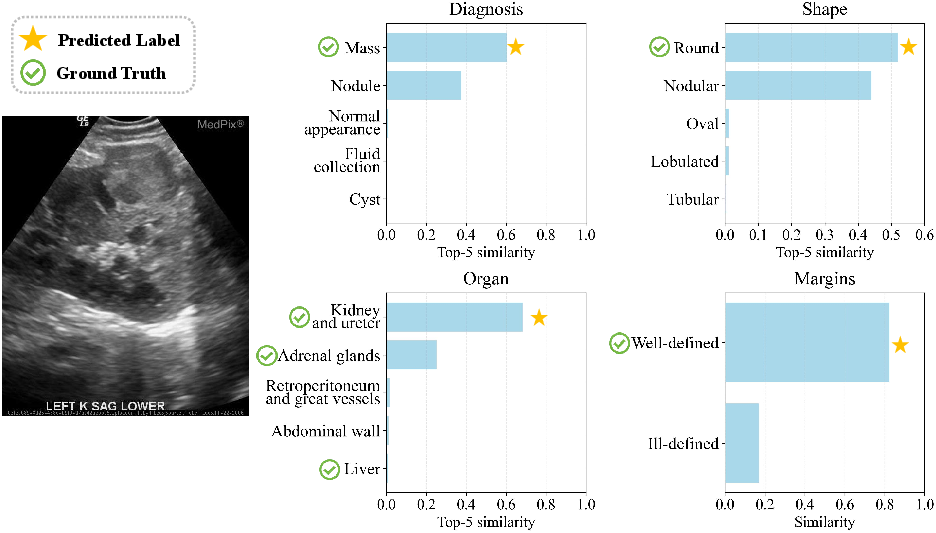
Figure 10: Multi-label insight—ranked outputs cover clinically plausible alternatives in complex lesions.
Theoretical and Practical Implications
The integration of domain-driven semantic priors and structured attribute graphs into the contrastive paradigm yields new directions for modality-specific VLP research. The authors demonstrate that anatomical and diagnostic taxonomies, if faithfully encoded into both data and model, can overcome the limits of vanilla contrastive learning in high-ambiguity, high-domain-shift contexts.
Practically, this approach enables:
- Significantly improved cross-site, cross-population diagnostic inference in ultrasound—a common clinical bottleneck.
- Superior performance in low-annotation and zero-shot application regimes, critical for rare findings and global deployments.
- Enhanced interpretability and attribute-level reasoning, crucial for clinical acceptance and trust.
Future Directions
Possible extensions include:
- Extension of UDT/UDAF to support other heterogeneous, specialist modalities (e.g., echo, fetal, vascular).
- Joint graph-based modeling of image and text, with tighter coupling at the feature and token level.
- Integration with LLM-based report generation for more naturalistic diagnostic output.
- Exploration of multi-label outputs for multi-pathology findings and richer clinical reporting.
Conclusion
Ultrasound-CLIP (2604.01749) establishes a new benchmark for ultrasound-centered image-text understanding through the fusion of ontology-driven supervision, a large-scale, modality-specific dataset, and a semantic-aware CLIP-style model. By addressing modality gap, annotation ambiguity, and attribute-structure representation, this work delineates a principled path for future clinically robust VLP systems in sonography and broader medical imaging.














