- The paper presents EndoASR, a novel ASR system featuring a two-stage domain adaptation with synthetic data and noise augmentation that significantly improves transcription accuracy.
- It demonstrates a reduction in character error rate from 20.52% to around 14.14% on clinical data and boosts medical term accuracy from 54.3% to 87.6%, outperforming general ASR models.
- The study validates EndoASR in multi-center clinical trials, showcasing its low-latency performance and robust human-AI teaming in real-world gastrointestinal endoscopy workflows.
Domain-Adaptive Speech Recognition for Human-AI Teaming in Real-World Gastrointestinal Endoscopy
Introduction and Motivation
This work presents EndoASR, a domain-adapted automatic speech recognition (ASR) system tailored for real-time deployment in gastrointestinal endoscopy workflows. The paper addresses limitations of general-purpose ASR models in clinical environments, emphasizing specialized medical terminology, high acoustic variability, and latency constraints in procedural medicine. The authors argue that ASR is a critical interface for effective human-AI teaming, particularly in multi-task, hands-busy workflows such as endoscopy, where natural language voice control is superior to alternative interaction modalities.
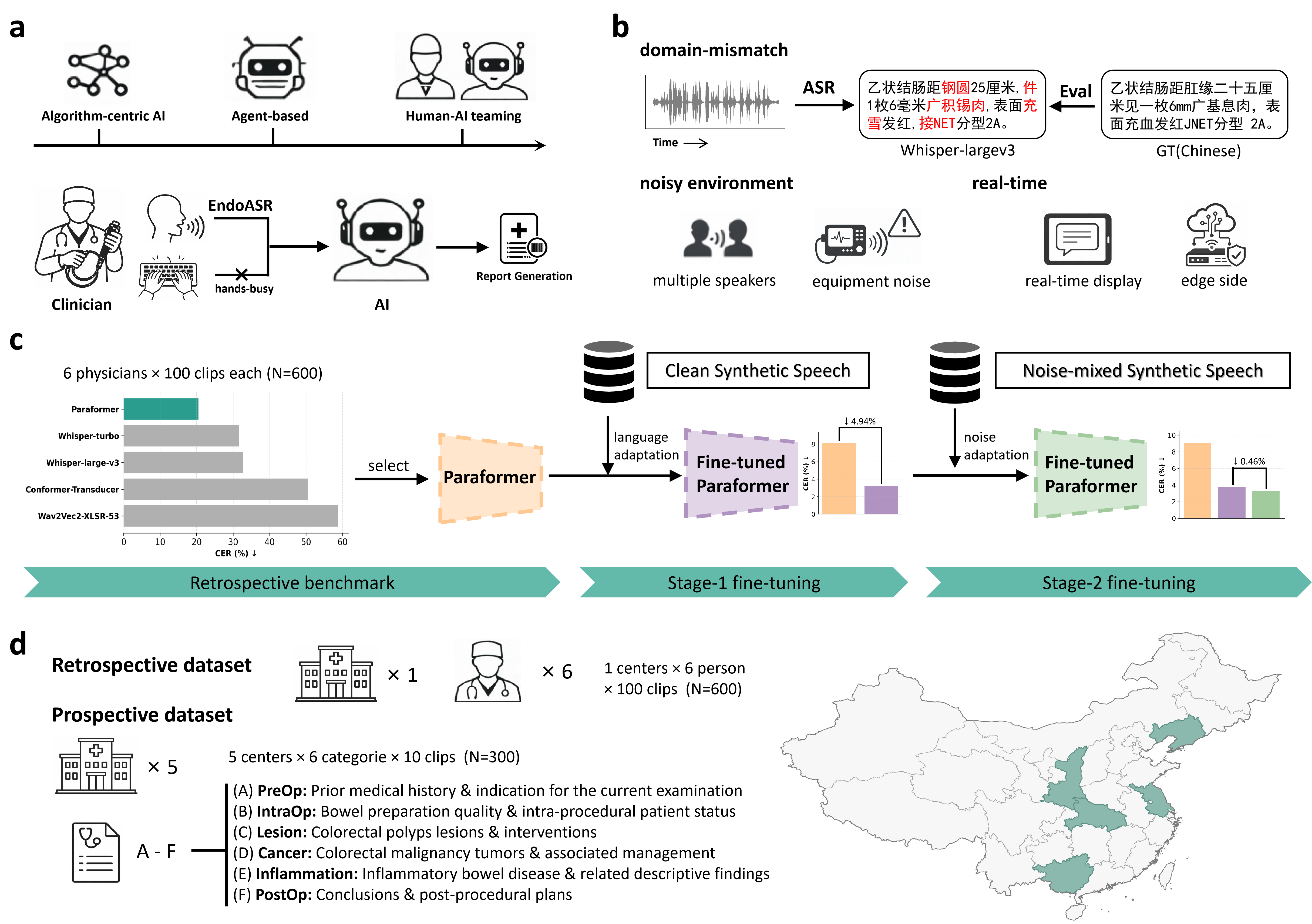
Figure 1: The EndoASR framework, illustrating a methodological pipeline from human–AI teaming, domain-specific challenges, a two-stage adaptation strategy, to progressive clinical validation.
Methodological Design
The EndoASR system is based on Seaco-Paraformer, a non-autoregressive end-to-end ASR model. The core methodological innovation is a two-stage domain adaptation pipeline designed to address linguistic and acoustic mismatch:
- Stage 1 – Domain-Specific Language Adaptation: The base model is fine-tuned on synthetic speech generated from structured endoscopy reports, optimizing for recognition of professional terminology and formulaic reporting styles.
- Stage 2 – Noise-Robustness Enhancement: The model is further fine-tuned with noise-augmented synthetic data, where real-world endoscopy room recordings are mixed into the synthetic speech, thus modeling challenging, heterogeneous acoustic backgrounds.
The complete framework supports both controlled (retrospective) and real-world (prospective, multi-center) clinical validation, allowing quantification of both algorithmic advances and practical deployment robustness.
Retrospective Benchmarking
General-purpose ASR models, including Whisper-large-v3 and wav2vec2-XLSR-53, were systematically benchmarked against Paraformer on 600 intraoperative recordings from six endoscopists, with no domain adaptation applied. Results demonstrate that Paraformer outperformed both large-scale and smaller models, achieving a CER of 20.52% and superior semantic metrics, providing the backbone for EndoASR development.
Model comparison revealed that state-of-the-art general ASR systems exhibit high error rates and notable deficiencies in medical term transcription under authentic clinical acoustics. Paraformer, with a favorable efficiency–accuracy trade-off, was selected as the adaptation substrate.
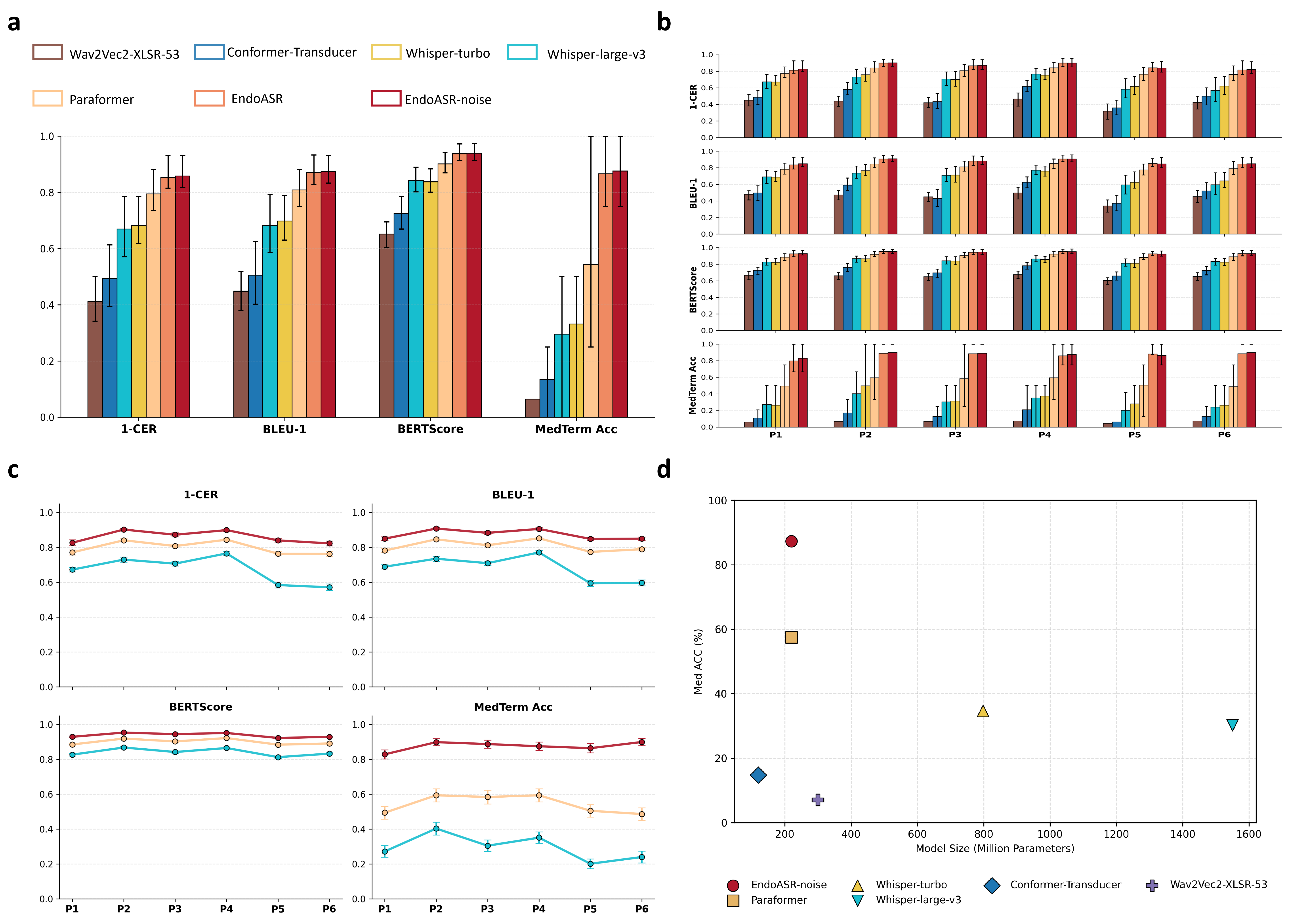
Figure 2: Retrospective performance across metrics, speaker- and parameter-dependent variability, and the medical terminology recognition/efficiency frontier.
Two-Stage Adaptation Strategy and Controlled Evaluation
Fine-tuning Paraformer on synthetic domain-specific speech (Stage 1) immediately reduced CER to 3.71% on synthetic data and 14.72% on clinical data; noise-robust fine-tuning (Stage 2) brought further improvements (CER 14.14%). Medical term accuracy (Med ACC) improved from 54.3% (baseline) to 87.6% post adaptation—confirming the effectiveness of domain-adaptive techniques.
Speaker-wise breakdown demonstrates that both language adaptation and noise robustness generalize over diverse individual speaking styles, though gains are predominantly driven by linguistic adaptation due to greater inter-speaker than inter-noise variability.
Prospective Multi-Center Validation
For deployment robustness, prospective evaluations were conducted on real-world data from five endoscopy centers, each characterized by diverse recording environments and clinical content types. t-SNE visualizations of MFCC-derived acoustic embeddings confirmed pronounced inter-center heterogeneity.
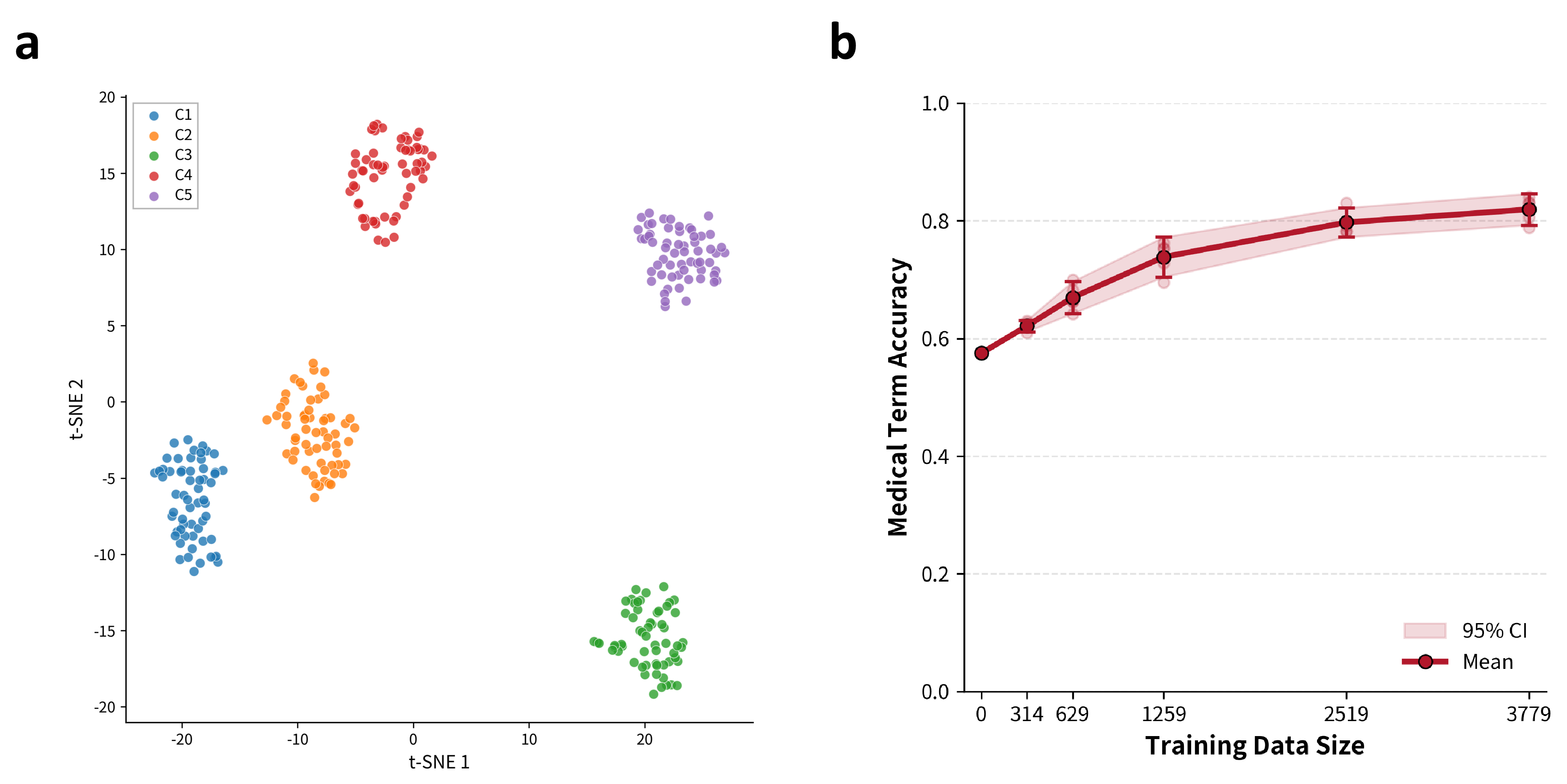
Figure 3: t-SNE embedding of multi-center speech segments, displaying acoustic clustering by center, and the monotonically increasing trend of Med ACC with synthetic data scale.
EndoASR and EndoASR-noise consistently outperformed Paraformer across all metrics (mean CER reduced from 16.2% to 14.97%; Med ACC from 61.6% to 84.2%). Importantly, terminology accuracy gains proved most significant in clinically critical workflow segments (lesion description, intra-operative communication) and showed only modest saturation as synthetic data scale increased.
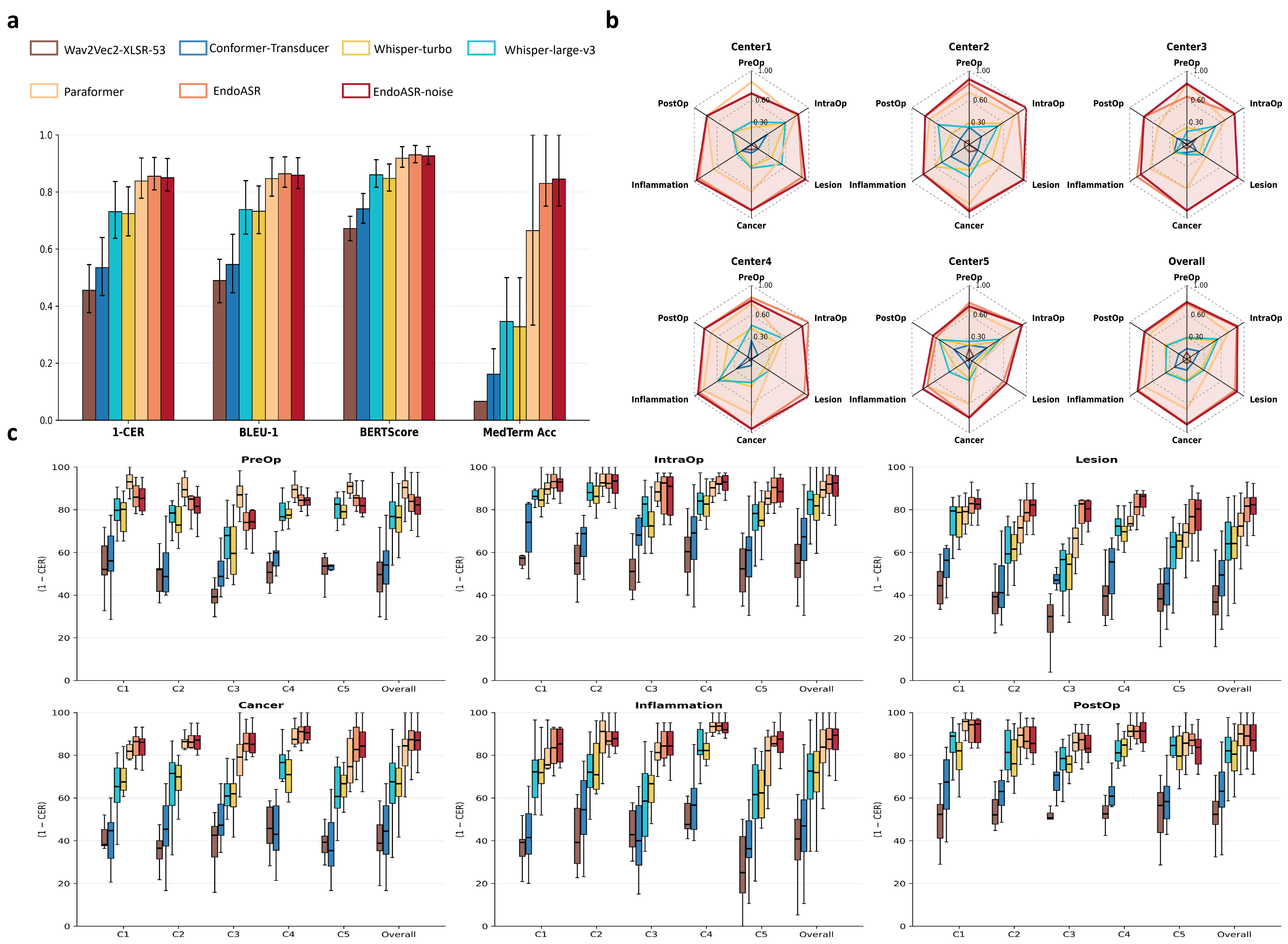
Figure 4: Multi-center breakdown of CER and Med ACC by clinical content category and institution, demonstrating robust cross-domain performance of EndoASR models.
Further analysis uncovered that noise-robust fine-tuning is most impactful when clinical acoustic conditions closely match training augmentation statistics; linguistic adaptation alone explains the bulk of cross-center generalization gains in current deployment scenarios.
Downstream Integration and Systemic Impact
Embedding EndoASR outputs in LLM-powered clinical information extraction pipelines demonstrated that higher ASR fidelity translates directly to error reduction in downstream structured documentation and automated report generation. EndoASR produced near-real-time, high-accuracy medical transcripts, facilitating new forms of clinician–AI interaction essential for next-generation clinical workflow automation.
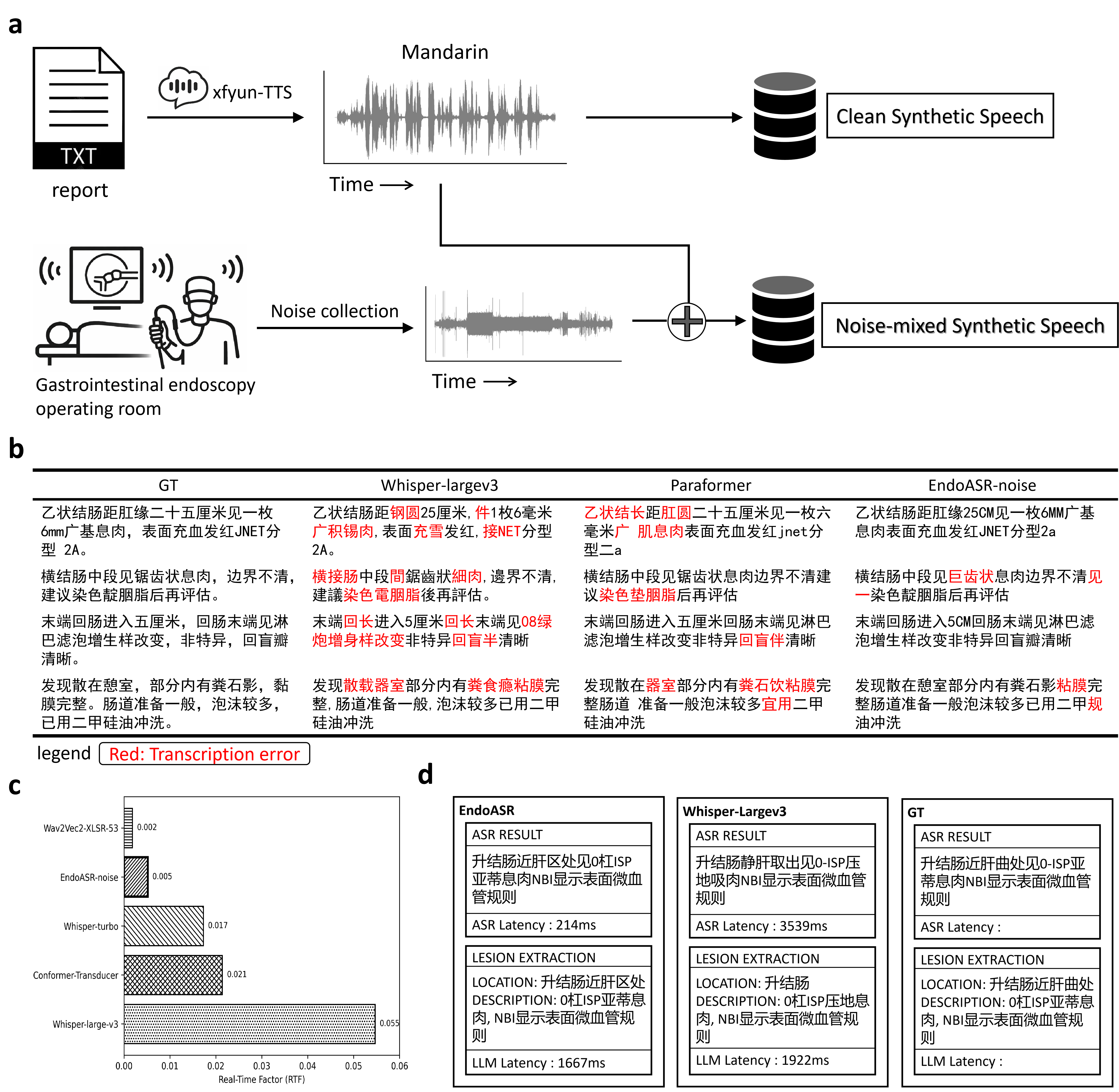
Figure 5: Upstream training data construction and augmentation, transcription quality head-to-head, RTF profiling, and downstream LLM-based report extraction comparison between EndoASR and Whisper-Large.
EndoASR's real-time factor (RTF) of 0.005 far outpaces Whisper-large-v3 and is competitive even against smaller models, supporting its suitability for low-latency, edge-side deployment in high-acuity clinical settings.
Implications and Future Directions
The study provides compelling evidence that domain-adaptive strategies, leveraging even moderate-scale synthetic data and center-specific noise augmentation, can robustly bridge the OOD gap between generalist ASR models and high-demand, specialized medical speech applications. Gains in terminology recognition yield downstream improvement in automated reporting, clinical decision support, and auditability in human–AI teaming.
The theoretical implications include a demonstration that tailored linguistic fine-tuning dominates ASR adaptation returns within highly-structured specialized domains; practical implications involve enabling real-world trials and scaled uptake of conversational AI agents in medical environments.
Future research should incorporate multi-accent and spontaneous dialogue data to further increase inclusivity and robustness. Center-agnostic or dynamic noise modeling may further enhance out-of-distribution generalization. Integrating ASR outputs with agentic LLMs could unlock higher-level workflow intelligence, with feedback between perception modules (such as EndoASR) and reasoning components as an avenue for adaptive improvement.
Conclusion
This paper establishes that domain-adapted ASR, achieved with a scalable two-stage adaptation mechanism, produces marked and clinically salient advances in speech-driven human–AI interaction for gastrointestinal endoscopy. The EndoASR system demonstrates strong generalization and efficiency, validated across heterogeneous multi-center cohorts, and paves the way for reliable, workflow-integrated AI agents in procedural clinical practice. This approach and its methodology are directly extensible to other complex, terminology-intensive medical and technical domains.






