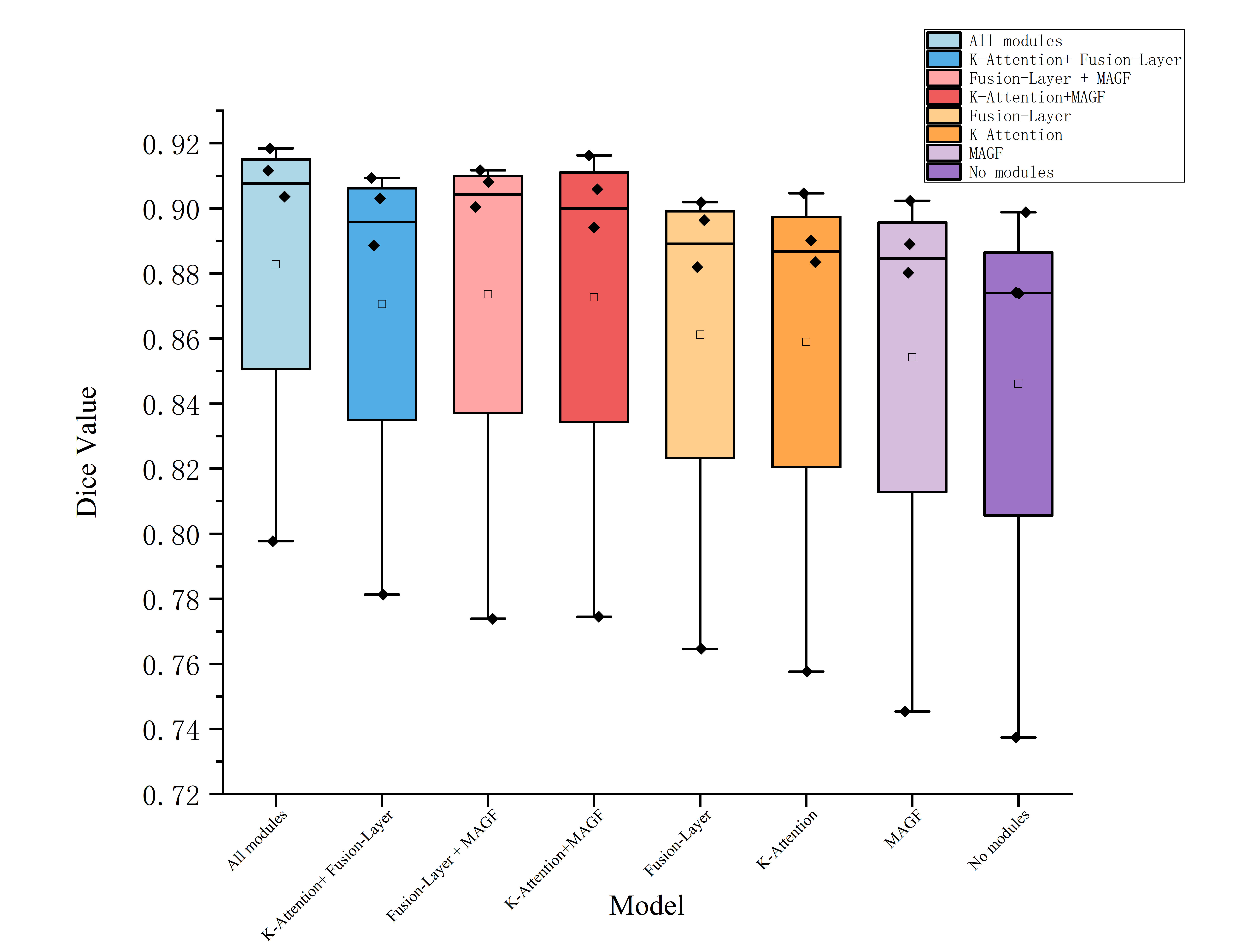- The paper introduces Rich-U-Net, a novel model that fuses spatial context with LSTM-based depth modeling via Sparse K-Attention to capture minute anatomical details.
- The paper employs a Multi-Scale Adaptive Gated Fusion module to combine multi-resolution features and improve boundary recovery in complex clinical images.
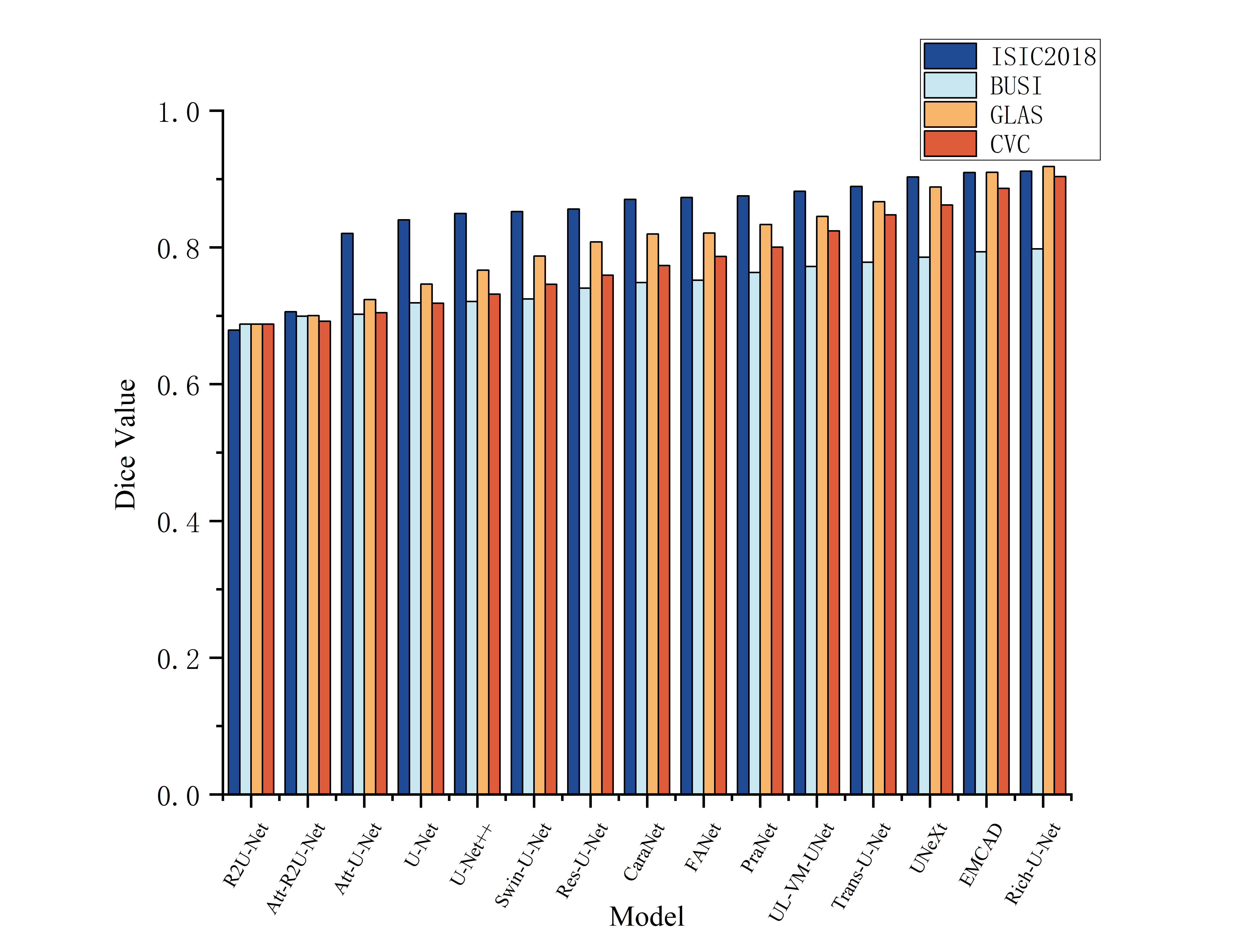
- The paper demonstrates superior performance with higher Dice scores and lower HD95 on benchmarks, outperforming current transformer-based and multi-scale methods.
Rich-U-Net: A Model for Fusing Spatial Depth Features and Capturing Structural Detail in Medical Image Segmentation
Introduction
Rich-U-Net addresses limitations common to most U-Net variants in medical image segmentation, specifically the underutilization of spatial context and the insufficient modeling of minute and complex anatomical structures. The model synthesizes multiple advanced mechanisms, including Sparse K-Nearest Neighbor Attention (K-Attention), a novel Fusion-Layer that amalgamates LSTM-based depth modeling with spatial convolutions, and a Multi-Scale Adaptive Gated Fusion (MSAGF) module, to yield state-of-the-art results across several medical segmentation benchmarks. The architecture is explicitly optimized for complex clinical images where fine-grained boundary recovery and contextual understanding are critical.
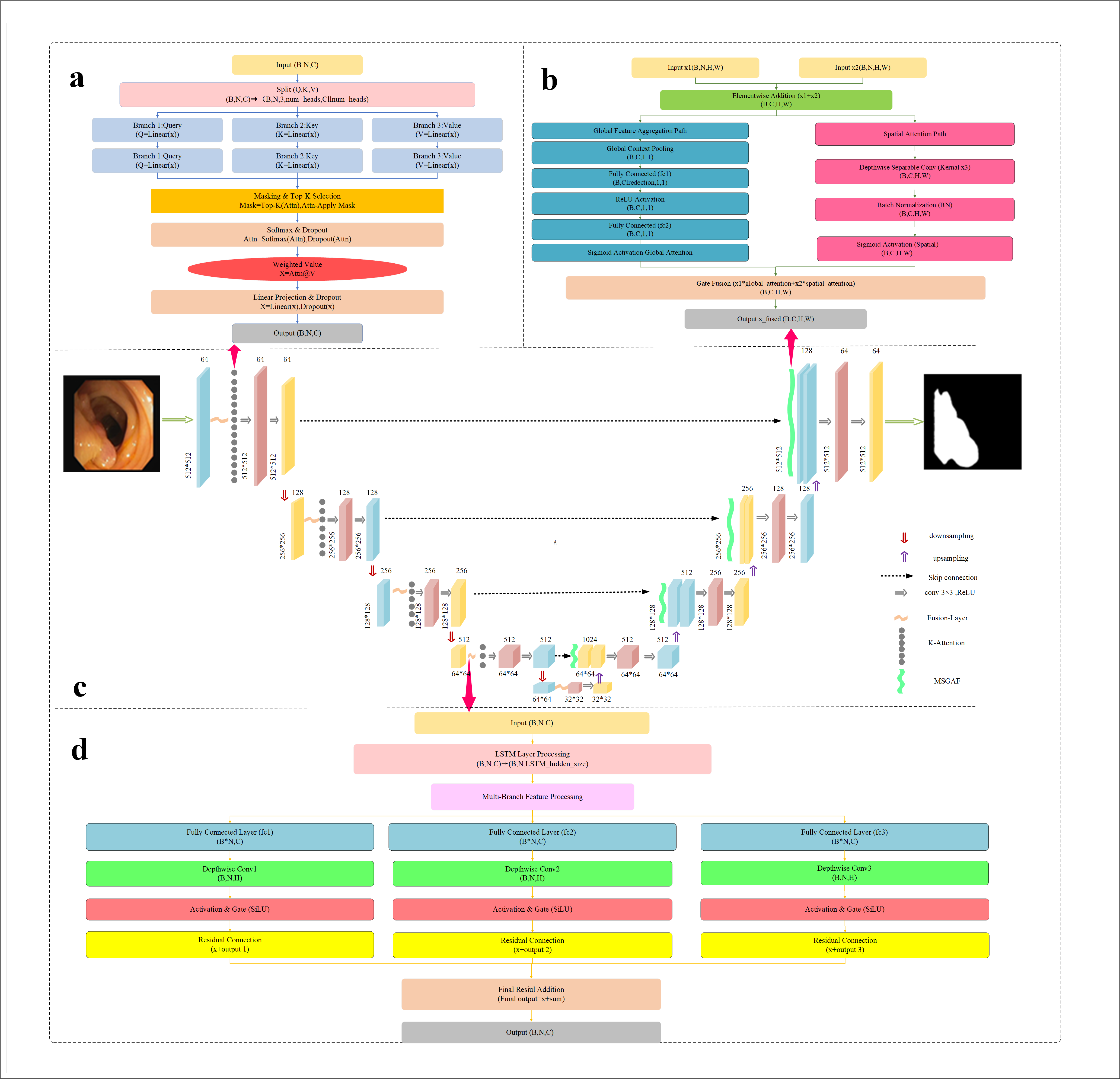
Figure 1: Rich-U-Net architectural overview, integrating K-Attention, Fusion-Layer, and MSAGF modules within a U-shaped topology.
Model Architecture
Rich-U-Net preserves the encoder-bottleneck-decoder macro-structure typical of U-Net while embedding three major architectural innovations:
- K-Attention Module: Introduced post-encoding, this mechanism sparsifies self-attention using a kNN-like top-k relationship mask, focusing computational and modeling capacity on the most relevant neighborhoods for each token. This facilitates robust local and global context modeling, particularly for boundaries and small structures with ambiguous contrast.
- Fusion-Layer: Fuses temporal and spatial dependencies using LSTM to model sequence/contextual (depth or slice-level) information and depthwise separable convolutions for spatial detail extraction. A gating mechanism dynamically weighs these components, allowing selective information flow both temporally and spatially, which is crucial for medical images where context can be heterogeneous.
- MSAGF Module: In the decoder, multi-scale features are aggregated through parallel global (channel) and spatial (pixel-wise) paths, both regulated by learned attention gates. This dual-path aggregation exploits low- and high-resolution cues for improved segmentation robustness, notably when lesion scale and appearance vary considerably.
These components are orchestrated with skip connections and residual operations to maximize feature reuse and maintain gradient flow during optimization.
Experimental Results
Experiments were conducted on ISIC2018 (skin lesions), BUSI (breast ultrasound), GLAS (colon histology), and CVC (colonoscopy polyp) benchmarks. Dice, IoU, and 95-th percentile Hausdorff Distance (HD95) were used for evaluation. Across all datasets, Rich-U-Net outperforms highly competitive baselines, including transformer-based architectures, lightweight variants, and recent multi-scale decoders.
Visualization of segmentation masks demonstrates effective delineation of minute structures, with clear superiority over contenders especially for regions of ambiguous intensity or small size.
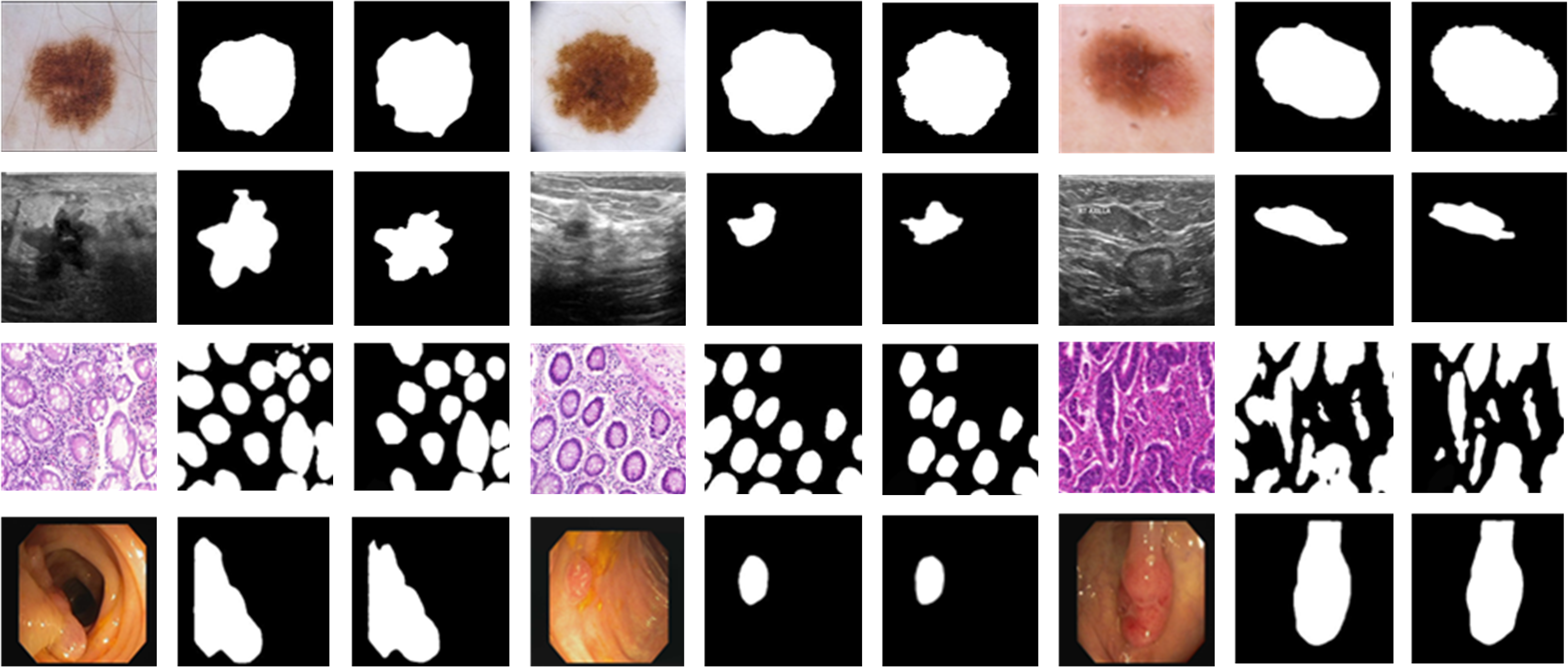
Figure 3: Qualitative Dice performance examples on multiple datasets, showcasing segmentation accuracy at various scales.
Ablation Study
Ablation experiments disassemble the contributions of K-Attention, Fusion-Layer, and MSAGF:
Methodological Implications
- K-Attention imposes structure on self-attention, reducing computational complexity from O(N2) to O(kN) and enhancing feature selectivity.
- Fusion-Layer bridges spatial and depth-wise (or inter-slice/temporal) dependencies; such integration can be readily extended to 3D volumetric data or multimodal temporal imaging.
- MSAGF generalizes the notion of multi-scale fusion, making it agnostic to resolution and thus resilient to differences in imaging protocol, scale, or anatomical context.
These findings empirically support the necessity of contextually selective attention, scale-agnostic fusion, and explicit depth modeling in clinical segmentation. The modularity facilitates incorporation into diverse frameworks, including hybrid Transformer-CNN backbones.
Future Directions
Potential research directions include:
- Fusing Rich-U-Net with unsupervised or semi-supervised objectives to leverage the wealth of unlabeled clinical imagery.
- Extending MSAGF to cross-modal fusion (e.g., MRI and CT).
- AutoML-based discovery of optimal scale fusion strategies for new imaging modalities.
- Investigating interpretability, especially which module primarily contributes to error reduction in challenging scenarios (e.g., rare pathologies).
Conclusion
Rich-U-Net sets a new performance baseline for medical image segmentation by explicitly fusing spatial and depth features and emphasizing minute anatomical structure. Its performance superiority and robust design validate the importance of sparsity-aware attention, gated multi-scale fusion, and spatio-temporal context modeling in medical vision applications. The results motivate further exploration of context-dependent modeling and adaptive fusion strategies for broader biomedical imaging challenges.