- The paper presents PuckerFlow, a novel generative model using flow matching in CP space to produce accurate cyclic conformers.
- Key methodology involves a 3D equivariant graph neural network with a cyclic Fourier filter and a tailored prior to enforce ring closure and symmetry.
- Benchmarking shows PuckerFlow outperforms baselines, achieving higher coverage and lower RMSD in CP space for five- to eight-membered rings.
Introduction and Motivation
Cyclic motifs are foundational in many functional molecular systems, with a critical role in drug discovery and catalysis owing to their conformational pre-organization and restricted flexibility. Despite their prevalence, efficient and reliable generation of diverse, energetically realistic conformer ensembles for small- and medium-sized rings remains a significant challenge, primarily due to the strong coupling in internal degrees of freedom required to maintain ring closure.
"Generating Cyclic Conformers with Flow Matching in Cremer-Pople Coordinates" (2601.12859) introduces PuckerFlow, a novel generative model that performs flow matching directly in the low-dimensional, chemically-informed internal coordinate system known as Cremer-Pople (CP) space. The CP coordinate system succinctly captures the essential puckering modes governing ring conformations, offering a reduction from the full 3D Cartesian representation and enabling efficient, symmetry-adapted learning.
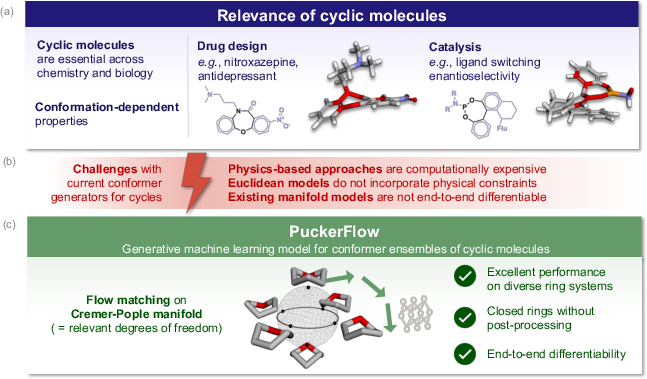
Figure 1: Cyclic molecules are critical in drug design and catalysis, but existing generative models often struggle with accurate closed-ring geometry.
Cremer-Pople Coordinates: Compact Manifold for Ring Systems
CP coordinates are derived as a discrete Fourier transform over the out-of-plane displacements of ring atoms, providing an (N−3)-dimensional manifold for an N-membered ring. This representation captures both the amplitude and phase of ring puckering modes, enabling description and comparison of chemically relevant conformational variations across different ring sizes and chemistries. For example, pseudorotational transformations are represented as phase changes at fixed amplitude within CP space.
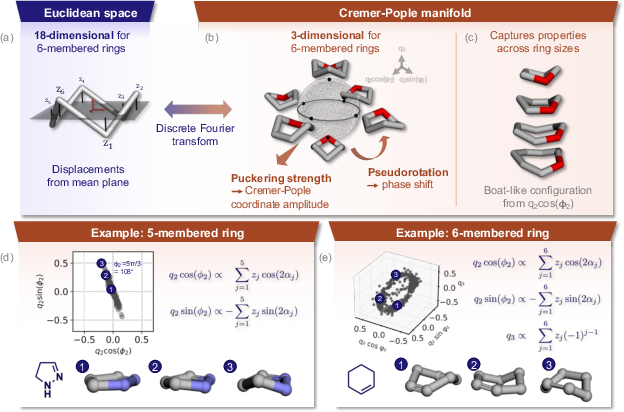
Figure 2: The CP framework captures essential ring deformations in a chemically interpretable, low-dimensional space.
The authors demonstrate that, for representative five- and six-membered rings, characteristic conformational clusters are cleanly resolved in CP space, illustrating that CP coordinates facilitate direct modeling of conformational diversity with substantial dimensionality reduction.
Methodology: PuckerFlow Architecture and Flow Matching
PuckerFlow leverages the flow matching paradigm, learning a time-dependent vector field that deterministically transports initial samples from a geometry-informed prior in CP space toward the distribution of real conformers. Crucially, this prior is amplitude-bounded and tailored to exclude geometrically invalid regions (such as infeasible bond lengths), a capability made possible by flow matching's flexibility in prior design.
The core neural architecture employs a 3D equivariant graph neural network, with atomic embeddings passed to a cyclic Fourier filter that produces outputs adapted to the symmetry and dimensionality of the relevant CP space. Model outputs respect ring size–dependent internal coordinate dimensionalities and maintain necessary (pseudo)scalar and parity symmetries.
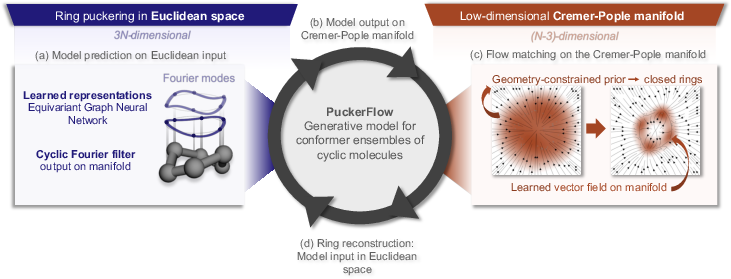
Figure 3: PuckerFlow pipeline—learned atomic embeddings are processed by the cyclic Fourier filter, yielding CP space updates, reconstructed into 3D structures.
To reconstruct full molecular geometry from CP coordinates, the method employs ring-size and chemistry-dependent dictionaries of bond lengths and angles, ensuring consistent reparameterization and closure of generated rings upon conversion to Cartesian coordinates.
Benchmarking and Numerical Results
PuckerFlow is evaluated on the ring puckering dataset from Folmsbee et al., focusing on five- to eight-membered monocyclic systems drawn from diverse chemical sources. The model is benchmarked against RDKit's ETKDG (including the small-cycles extension), and two generative baselines—GeoDiff and MCF.
Quantitative metrics include Average Minimum RMSD (AMR) and coverage within a 0.1 Å threshold, both overall and specifically on the CP (puckering) degrees of freedom. PuckerFlow outperforms all baselines in both precision and recall, with particularly strong improvements over Euclidean models such as MCF and GeoDiff, even after geometry relaxation with MMFF94.
For example, without relaxation, PuckerFlow attains a coverage of 67.5% (vs. 46.2% for MCF and 51.4% for RDKit Small Cycles), and an AMR of 0.13 Å in CP space (vs. 0.16 Å for MCF). Performance gains persist or increase after force field minimization, and the method achieves comparable quality with as few as 2–5 inference steps, well below that required by competing models.
Visualization of generated conformer distributions in CP space for test set rings shows that PuckerFlow reproduces both the amplitude and angular characteristics of ground-truth ensembles, capturing characteristic multimodality and pseudorotational pathways.
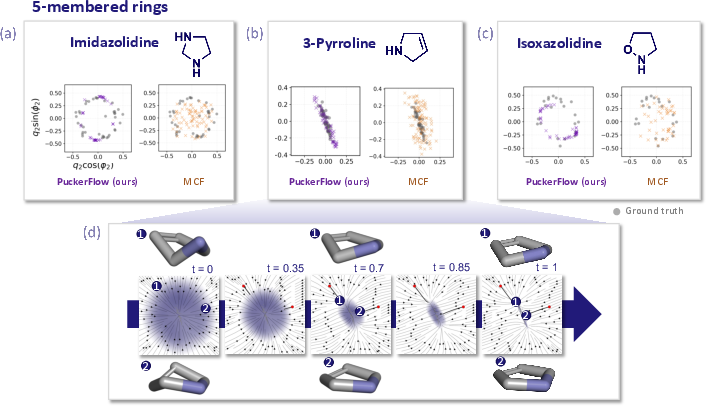
Figure 4: Conformer spaces of five-membered rings (e.g., imidazolidine) generated by PuckerFlow (violet) versus MCF (gold); PuckerFlow closely aligns with the ground-truth (grey).
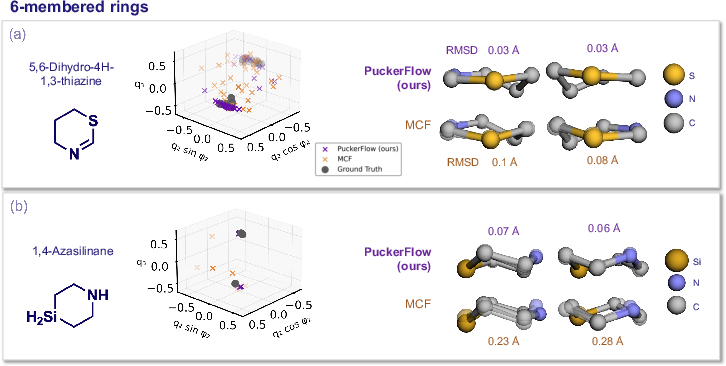
Figure 5: For six-membered rings, PuckerFlow correctly samples bi-modal distributions reflecting different chair forms, outperforming MCF.
Beyond monocyclic rings, PuckerFlow generalizes to seven- and eight-membered rings with strong performance, and, owing to modular internal coordinate generation, can be integrated with exocyclic substituent generators (e.g., RDKit ETKDG or torsional diffusion) for whole-molecule conformer sampling.
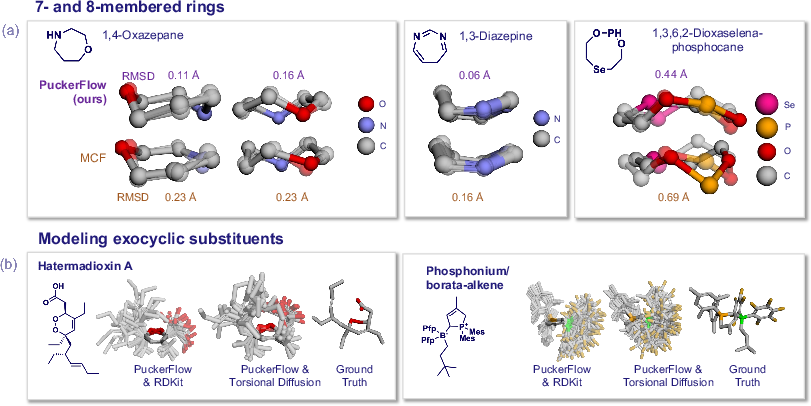
Figure 6: PuckerFlow generates high-quality conformers for larger rings and supports workflow integration with existing exocyclic modeling tools.
Technical Implications and Future Directions
PuckerFlow demonstrates that applying generative modeling to the chemically relevant internal coordinates, rather than standard Cartesian space, confers substantial improvements in both efficiency and reliability for ring conformer generation. This approach explicitly enforces ring closure, end-to-end differentiability, and enables parameter-efficient learning with rapid sampling.
Immediate implications include use in high-throughput virtual screening campaigns, where the fidelity of cyclic conformer libraries is critical, particularly for computational drug discovery pipelines and catalyst design. By providing accurate ensembles directly in CP space, PuckerFlow facilities better structure-property modeling and can serve as a drop-in replacement or enhancement to widely adopted tools.
The model's differentiable, internal-coordinate-based architecture opens several avenues for future research:
- Joint modeling of ring and exocyclic torsions: Integration with torsional diffusion or similar models for full-molecule, manifold-aware conformer generators.
- Scaling to macrocycles: Extension to larger, more flexible rings will require addressing non-convexity and sparse data.
- Property-conditioned generation: PuckerFlow's differentiability makes it amenable to gradient-based fine-tuning for property optimization or binding affinity-guided conformer generation, potentially within docking models such as DiffDock.
- End-to-end differentiable docking: Integration into ligand-binding pose prediction frameworks to improve ring geometry sampling in molecular docking.
Conclusion
PuckerFlow represents a substantial advance in the generative modeling of cyclic conformers, leveraging the Cremer-Pople internal coordinate system for efficient, precise, and symmetry-respecting generation. It surpasses established rule-based and deep generative baselines across all tested metrics and efficiently captures the structural diversity relevant for key applications in chemical and pharmaceutical research. By enforcing chemical validity by design and supporting end-to-end differentiable workflows, PuckerFlow sets a robust foundation for future developments in conformer generation and property-driven molecular design.







