- The paper introduces a novel framework that jointly generates network topology and weights using latent Gaussian fields on compact, multiply-connected manifolds.
- The methodology employs an inhomogeneous Poisson process and geodesic proximity to assign neuron positions and connectivity, ensuring sparsity and local field affinity.
- Empirical results on a torus demonstrate robust uncertainty quantification and data-efficient learning with competitive predictive metrics.
Supervised Learning of Random Neural Architectures Structured by Latent Random Fields on Compact Boundaryless Multiply-Connected Manifolds
Introduction and Motivation
This work establishes a formalized probabilistic framework for supervised learning in neural systems, uniquely characterized by architectures that are not fixed, but instead stochastically generated through latent Gaussian random fields defined on compact, boundaryless, multiply-connected manifolds. The formulation explicitly aims to address complex systems where output random variables are strongly non-Gaussian, and model uncertainty cannot be relegated to simple output noise. Rather than merely varying weights on a predetermined structure, the topology, connectivity, and weights of the neural architecture all emerge jointly from a reduced-order random field.
At its core, the network geometry is sampled via an inhomogeneous Poisson process modulated by the latent field, and input/output neurons are determined by extremal field values. Connectivity is determined through geodesic proximity on the manifold and local field affinity. The entire pipeline is conceptualized as a generative process: structural and parameteric uncertainty are encoded at the architectural level, not merely as Bayesian priors on weights.
Geometry-Aware Stochastic Neural Architecture
The stochastic architecture construction begins with a latent, real-valued, anisotropic Gaussian field defined on a compact, multiply-connected 2-manifold M⊂R3, constructed via the solution to an SPDE:
(τ0−⟨∇,[K]∇⟩)U(x)=W(x)
where [K(x)] is a position-dependent anisotropic tensor enforcing local geometry, and W(x) is Gaussian white noise. This model admits efficient reduced-order representations (m≪n0) via finite-element-based Karhunen-Loève expansions, enabling scalable sampling of the field and its covariance structure.
The manifold is discretized to a mesh for numerics.
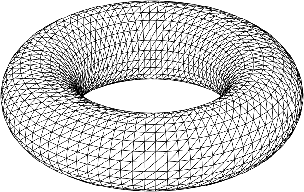
Figure 1: Structured finite-element mesh approximating the compact, boundaryless, multiply-connected manifold.
The Gaussian field's reduced covariance spectrum and associated approximation error are critical in controlling field variability and consequently the stochastic network's expressivity.
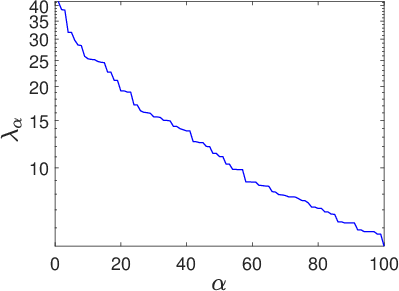
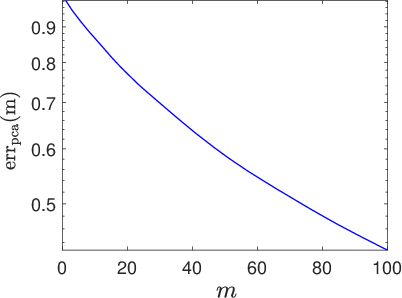
Figure 2: Eigenvalues of the latent field covariance matrix [C] on a log10 scale, quantifying retained variability in reduced-order representations.
Neuron locations are then determined by an inhomogeneous Poisson process whose spatial intensity is proportional to the pointwise variance of the latent field, explicitly concentrating computational resources in regions of high uncertainty.
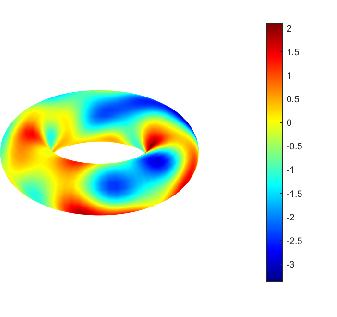
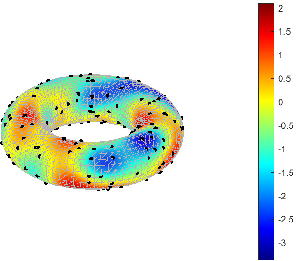
Figure 3: Realization of the anisotropic latent random field Um used to drive neuron placement and connectivity.
Random Graph Construction and Functional Differentiation
Neural connectivity is established by a geometry-informed binary mask. Using local geodesic distances on the manifold (interpolated via the mesh), a self-tuning diffusion kernel produces affinity scores, followed by percentile-based thresholding for scalable sparsification. The network thus explicitly encodes both geometric proximity and manifold topology.
Synaptic weights are further modulated by field-derived similarity: weights are high for geodesically proximal neurons with similar field values, enforcing both structural and feature-level locality. This process ensures a sparse, geometry- and field-aware, non-Gaussian random connectivity.
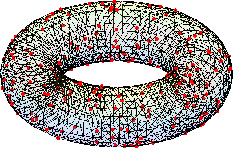

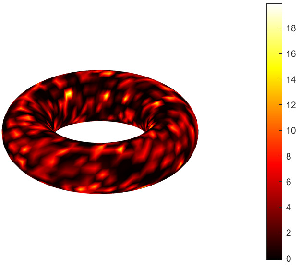
Figure 4: Realization of the random sparse connectivity graph derived from the field and manifold geometry.
Input and output neurons are not fixed, but are stochastically assigned according to the most extreme latent field values (least and greatest, respectively), introducing both architectural and functional randomness and obviating arbitrary input-output designation.
Supervised Likelihood-Based Hyperparameter Inference
Learning is cast as statistical inference on the hyperparameters of the generative process (h1,h2,ζs,β). Rather than direct optimization of weights, the framework defines a negative log-likelihood loss estimated via Monte Carlo sampling: for each deterministic input, the stochastic neural architecture samples a random function mapping, producing a sample output. Thus, even with a single observation per input, the network internalizes architectural uncertainty, estimating the conditional output law by explicit sampling.
Optimization employs Adam on the hyperparameters, with a hybrid global/local schema (grid search plus gradient steps), leveraging finite differences due to the model's implicit and intractable analytical gradients.
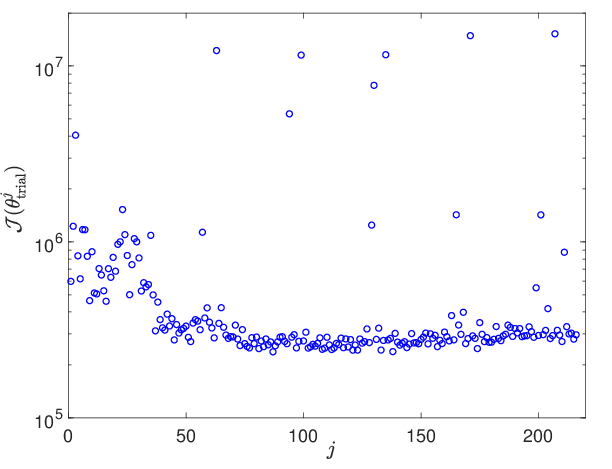
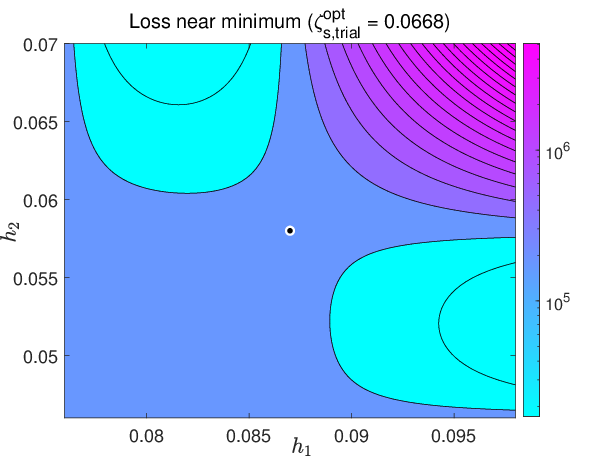
Figure 5: Loss function values across the hyperparameter trial grid—used for initialization of local search—demonstrating smoothness and suitable regularity for gradient-based methods.
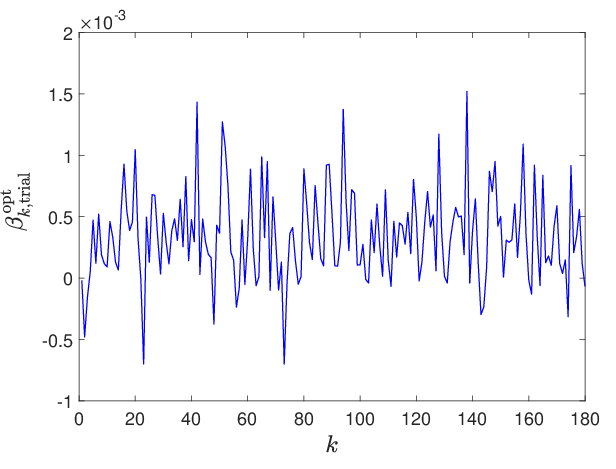
Figure 6: Evolution of the trial-based optimal parameter components k↦β∗,k, revealing regularization essential for robust learning.
Numerical Experimentation: Geometry-Driven Random Architectures on the Torus
The paper presents a comprehensive numerical illustration on the torus, a canonical compact boundaryless manifold, triangulated into ~2000 nodes. For N=200 neurons, nin=20, nout=100, and nd=3000 training samples, networks are constructed as described. Notably, only 25% of the possible connections are activated due to sparsification, and the network captures complex stochastic mappings with as few as 100 basis functions in the latent field representation.
The induced neural system accurately models the distributional (strongly non-Gaussian) outputs for complex mappings, achieving Competitive Continuous Ranked Probability Score (CRPS) and minimal overfitting as measured by normalized NLL comparisons:
- For N=200, nout=100, training and test NLLs are comparable (difference ratio <0.03), and the absolute CRPS aligns closely with a GKDE reference.
- Statistical diagnostics confirm accurate per-component confidence coverage and minimal local misfit.
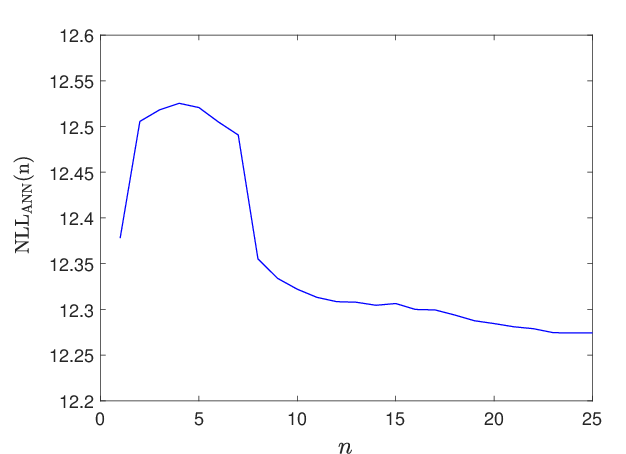
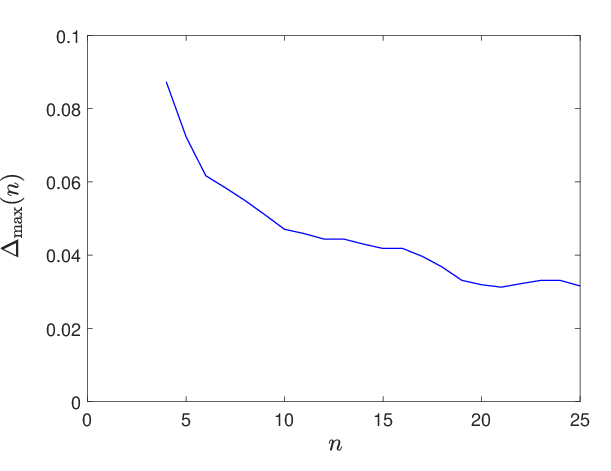
Figure 7: Loss convergence as a function of Adam/gradient descent iterations shows effective convergence despite high stochasticity.
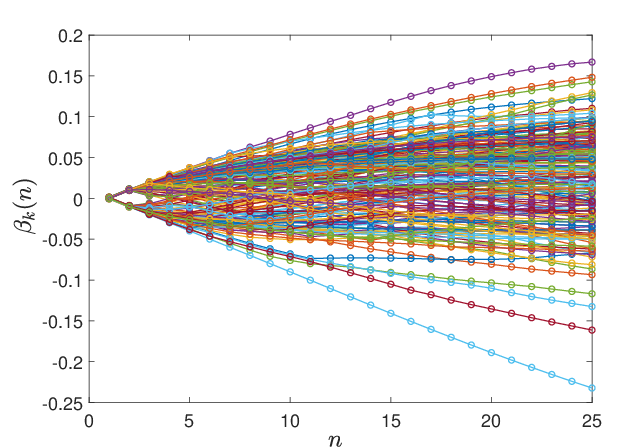
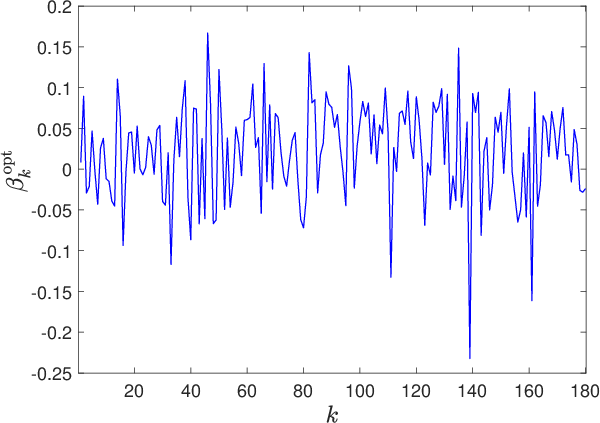
Figure 8: Evolution of hyperparameter components βk(n) over optimization, revealing smooth transition and absence of abrupt parameter jumps.
(Figure 9, 11, 12)
Figure 9: Accurate resolution and stabilization of learned parameter trajectories for different nh (field expansion basis) values.
- The model's expressivity and generalization are robust to mesh refinements, increasing latent basis size (nh), and changes in neuron number, indicating intrinsic architectural regularization from the geometric prior.
Mathematical Analysis and Theoretical Expressive Power
The paper delivers a theoretical footing for the expressive stochastic capacity of these models:
- The latent random field, parameterized via SPDE and low-rank expansions, is proven dense in the class of Gaussian fields with nontrivial diagonal structure, guaranteeing diverse architectural sampling.
- The mapping from hyperparameters to random neural graph is shown measurable and differentiable wherever required for gradient-based optimization.
- The resulting function class is dense in Hilbert-space—valued mappings from input to random output, suggesting that, with adequate basis and sufficiently flexible field parameterization, these stochastic architectures serve as universal approximators at the distributional level.
Implications and Future Directions
Contrary to Bayesian deep learning, where weight priors and function-space posteriors are anchored to fixed graphs, this framework induces uncertainty and regularization at the architectural (structural) level. This geometric prior provides a discipline to the learning capacity, preventing overfitting even in small-data, high-dimensional, non-Gaussian settings.
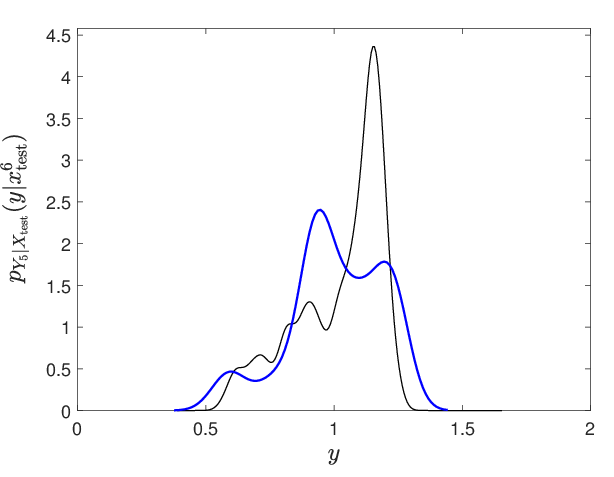
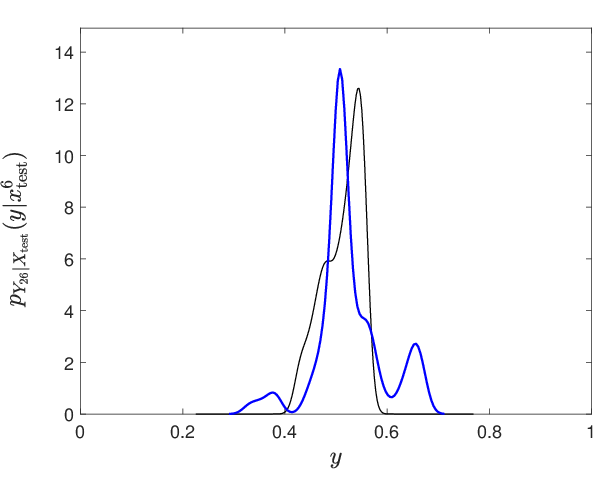
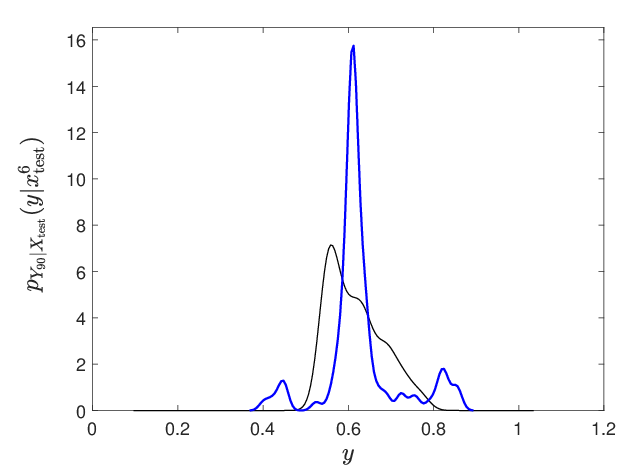
Figure 10: Representative conditional probability density estimation for a selected output neuron, demonstrating accurate capture of complex, non-Gaussian output laws.
Practical implications include:
- Physics-Informed Neural Networks (PINNs): The ability to encode physical geometry and boundary (or absence thereof) directly in the architecture supports applications in stochastic PDEs and physical sciences, aligning neural priors with problem topology.
- Uncertainty Quantification: The generative nature allows direct propagation of architectural and parameteric uncertainty to outputs, with empirical validation against high-quality nonparametric benchmarks.
- Data-Efficient Learning: The model provides robust estimates with a minimal number of neurons in severely data-limited examples, due to field-driven adaptivity.
Potential avenues include extension to deep/hierarchical architectures, hybridization with operator learning (e.g., neural ODEs on random graphs), and architecture adaption for domains with boundaries or evolving topologies, as well as refined inference and scalable learning algorithms.
Conclusion
The paper establishes a rigorous, geometry-driven foundation for supervised learning of random neural architectures generated by latent random fields on compact, multiply-connected manifolds. The approach introduces architectural uncertainty as a principled inductive bias, unifying network topology, weights, and role assignment in a single stochastic process, and demonstrates substantial empirical capacity with strong regularization and uncertainty quantification properties.
Theoretical exposition confirms expressive capacity and foundational well-posedness, positioning the model as an inverse-problem surrogate or uncertainty quantification tool in structured random systems, with significant implications for both stochastic machine learning and physics-informed computation.
(2512.10407)
















