- The paper introduces DeepGB-TB, a framework that combines a 1D-CNN for cough audio with a LightGBM model for demographic data to enhance TB screening.
- It employs a cross-modal bidirectional attention mechanism (CM-BCA) to iteratively refine audio and demographic features, mimicking clinical reasoning.
- Experimental results demonstrate an AUROC of 0.903, underscoring its high sensitivity and on-device applicability for rapid TB diagnosis.
DeepGB-TB: A Risk-Balanced Cross-Attention Gradient-Boosted Convolutional Network for Rapid, Interpretable Tuberculosis Screening
DeepGB-TB proposes a novel framework combining deep learning with gradient-boosted decision trees to offer an efficient, interpretable method for tuberculosis (TB) screening. It utilizes acoustic features from cough audio coupled with demographic data to predict TB risk. The architecture features a Cross-Modal Bidirectional Cross-Attention (CM-BCA) mechanism that enables data from cough audio to iteratively interact with demographic features, thereby refining diagnostic interpretations in a manner akin to a clinician’s diagnostic reasoning.
Methodology
DeepGB-TB integrates a 1D-CNN for processing cough audio and a LightGBM-based model for demographic data into a Cross-Validated Probability Embedding Module (CVPEM). The CVPEM transforms raw tabular data into robust, high-dimensional vectors by employing cross-validated LightGBM predictions as features.
1D-CNN and CVPEM Integration:
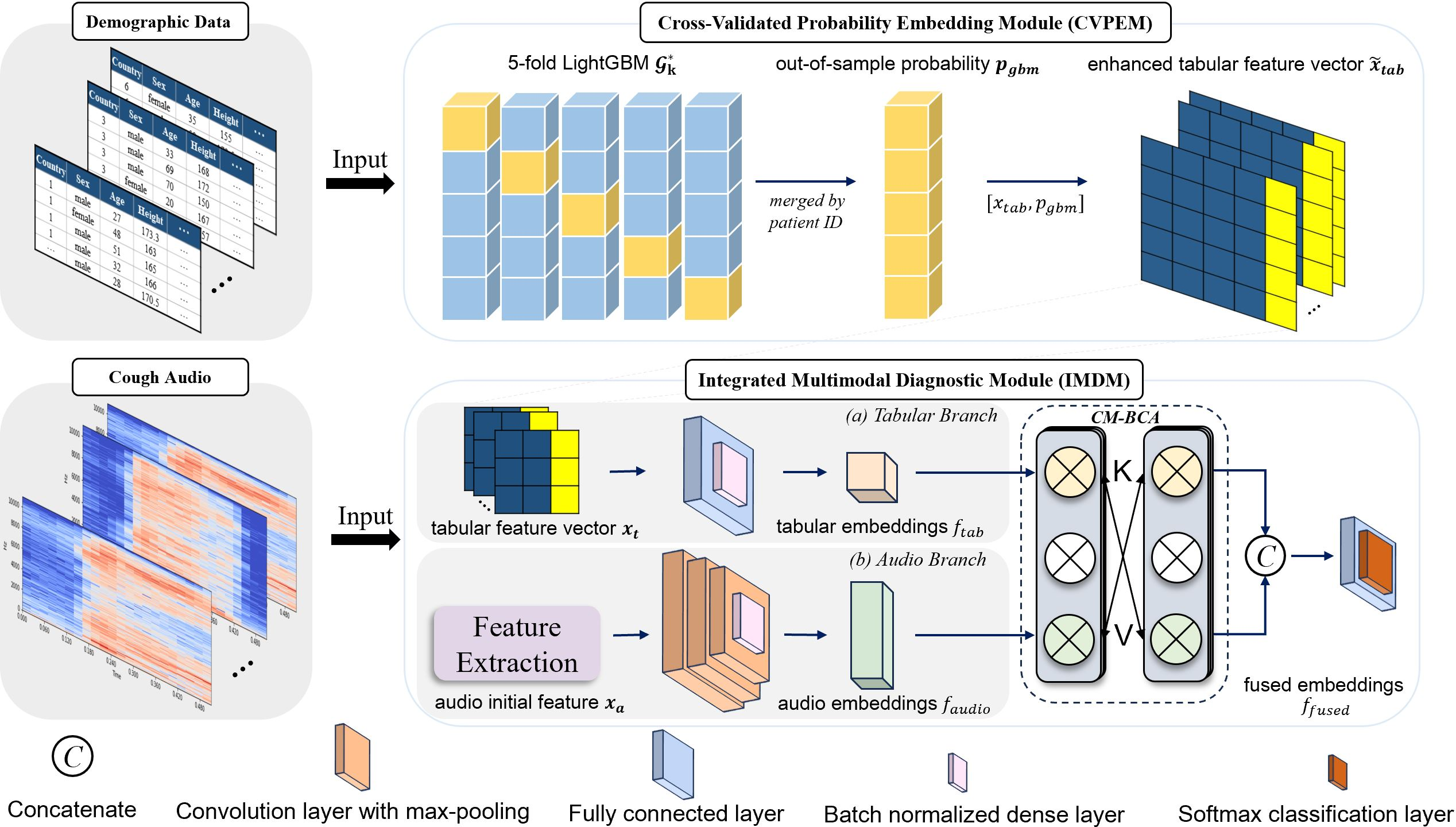
Figure 1: The architecture of DeepGB-TB.
The audio processing employs Mel-frequency cepstral coefficients (MFCCs) and supplementary features (chroma, spectral centroid) as inputs to a 1D-CNN, while a LightGBM model processes demographic data using cross-validation to yield a stable probability embedding. This configuration captures the multiplicative interplay between the modalities, effectively modeling patient profiles against audio signatures.
CM-BCA Mechanism:
CM-BCA iteratively refines the audio and tabular feature representations by cross-attending over the alternate data modality. This process, shown in (Figure 2) as a series of multi-head attention layers, relies on successive applications of attention and feed-forward layers interleaved with layer normalization. The CM-BCA provides an efficient fusion that supports the holistic assessment of patient data.
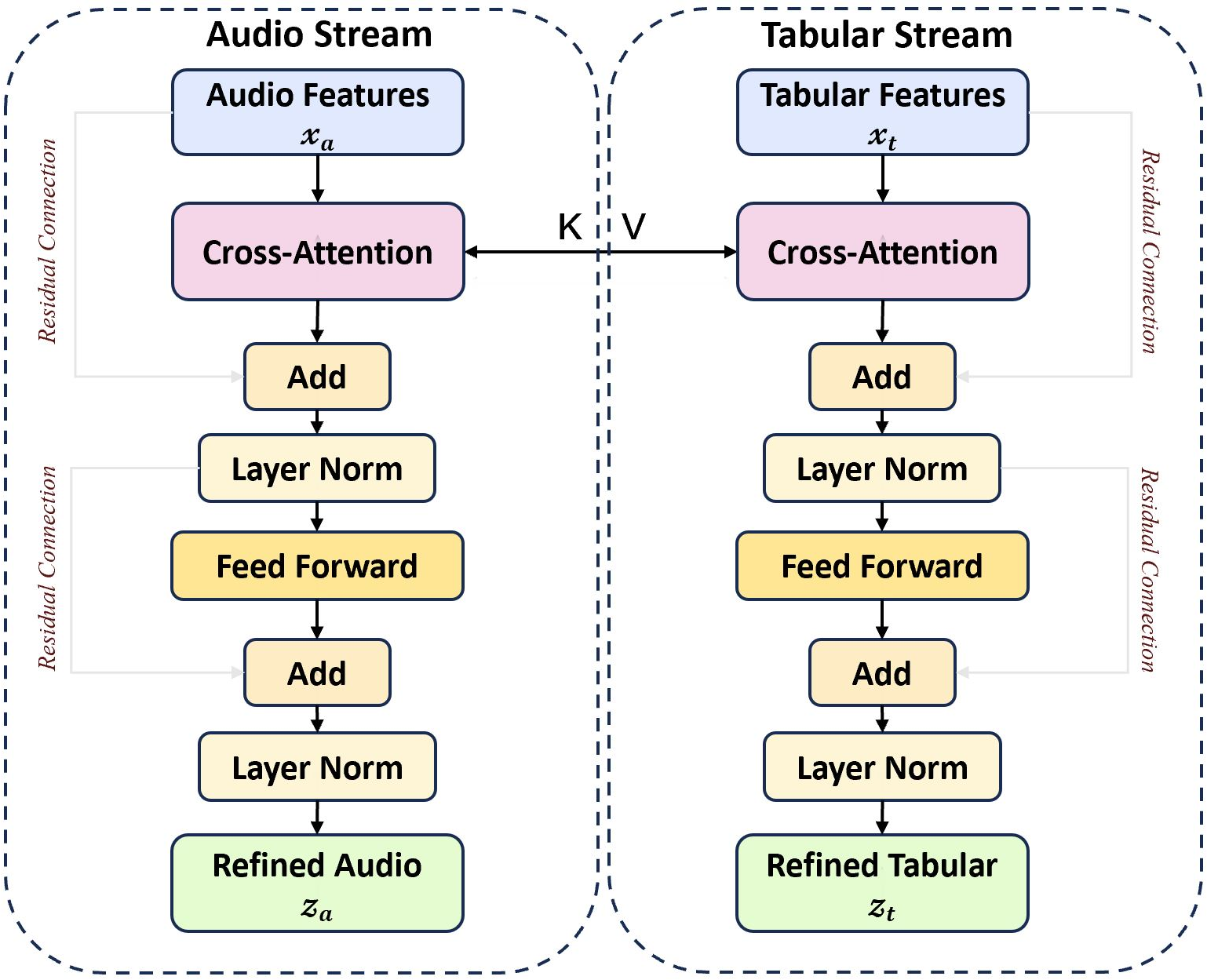
Figure 2: The process of CM-BCA.
Loss Function and Optimization:
To address the critical need for high sensitivity in TB detection, a Tuberculosis Risk-Balanced Loss (TRBL) is incorporated. This loss assigns a higher penalty to false negatives, optimizing the model for recall. The selected value of the hyperparameter λ=3 in the TRBL ensures an optimal balance between sensitivity and specificity as shown in empirical evaluations.
Experimental Results
DeepGB-TB was evaluated on a dataset comprising 1,105 participants from diverse geographic locations, featuring a balanced TB and non-TB distribution. The method achieved an impressive AUROC of 0.903, indicating robust diagnostic capability surpassing other baseline models.
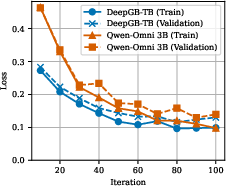
Figure 3: Comparison of Model Training and Validation Loss. The x-axis denotes epochs for DeepGB-TB and training steps for Qwen-Omni.
The ablation studies highlight the significance of each modular innovation in DeepGB-TB. Exclusion of the CM-BCA led to a significant AUROC decrease of 1.3%, underscoring its importance. On-device performance tests further validated the model's applicability in real-time settings, showing efficient processing capabilities on edge devices.
Discussion and Implications
DeepGB-TB exemplifies a tailored AI solution that effectively integrates multimodal data, achieving state-of-the-art TB screening potential through smart architectural choices like CM-BCA and TRBL. While the current results indicate high accuracy and efficiency, further prospective studies could explore the model's application in various settings to cement its clinical utility.
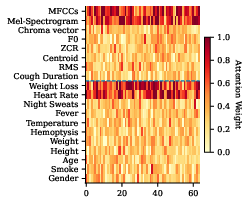
Figure 4: Attention heatmap over input features.
Given its design, the framework stands as a promising tool in overcoming the barriers to accessible TB diagnosis, especially in resource-limited settings. Future enhancements might consider expanding the dataset diversity or integrating additional modalities to bolster its diagnostic robustness.
Conclusion
The DeepGB-TB framework highlights the power of combining convolutional neural networks with decision trees and innovative fusion mechanisms, offering a compelling solution for rapid, mobile-based tuberculosis screening. Despite the methodology's inherent strengths, ongoing validation across broader datasets and settings remains essential to fully realize its potential for widespread clinical deployment.





