- The paper presents a two-step CorrNet method that integrates MRI and histopathology to improve prostate cancer diagnosis with higher specificity and sensitivity.
- It extracts correlated features using a pre-trained VGG-16 and adapts HED models, achieving AUC scores of 0.86 (pixel-level) and 0.96 (per-lesion).
- The approach reduces false positives and offers scalable diagnostic insights, enabling accurate cancer localization even without pathology images.
CorrSigNet presents a two-step methodology aimed at enhancing computer-aided prostate cancer diagnosis by integrating information from both MRI and histopathology images. The approach centers on leveraging common representation learning to correlate MRI features with histopathology-derived signatures for more accurate cancer localization.
The primary motivation behind CorrSigNet is to address the limitations of current MRI-based predictive models that fail to incorporate pathology-derived signatures, leading to suboptimal cancer localization. The variability in radiologist interpretations and the inherent limitations of MRI inputs necessitate improved methodologies that understand cancer signatures at a deeper biological level.
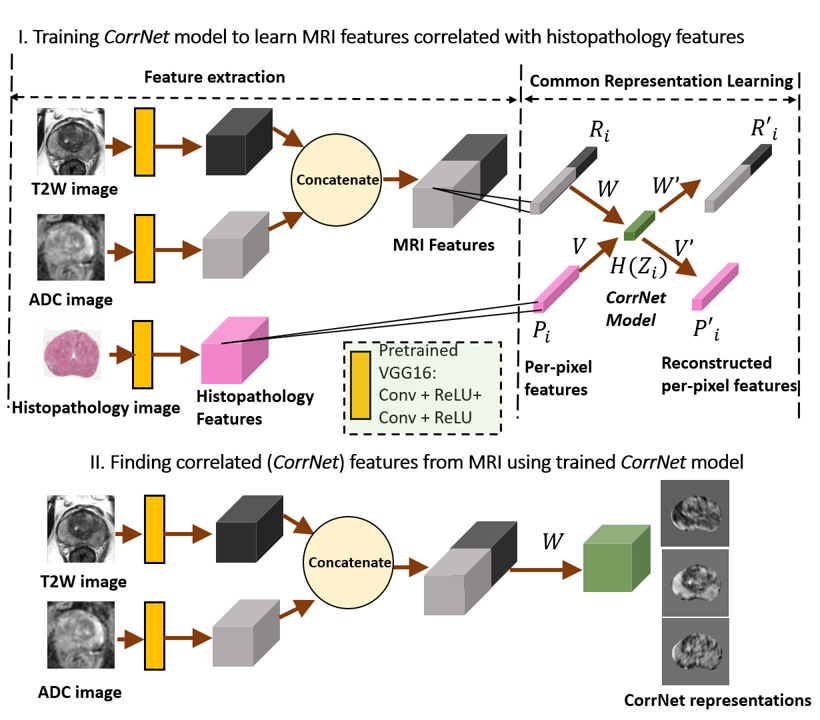
Figure 1: Learning correlated representations from spatially aligned MRI and histopathology images, and then constructing the correlated (CorrNet) representations from MRI alone using learned weights.
CorrSigNet's first stage employs a Correlational Neural Network (CorrNet) architecture to forge shared representations from MRI and histopathology images, learning correlated features per pixel that relate closely to cancerous regions. These learned representations enable subsequent prediction models to identify cancer with increased specificity and sensitivity, even in the absence of histopathology images.
Methodology
Dataset Preparation
The study's dataset comprises MRI and histopathology images of prostatectomy specimens from 95 patients. Advanced spatial alignment techniques were employed to ensure rigorous mapping between anatomical and pathological slices, creating a precise basis for correlation analysis.
Common Representation Learning
The heart of CorrSigNet lies in its CorrNet framework, tasked with extracting correlated features from spatially aligned image modalities. This architecture minimizes reconstruction errors while maximizing correlation between MRI and pathology-derived views. Features are extracted using the initial layers of a pre-trained VGG-16 model, creating a high-dimensional space accurately reflecting potential cancerous characteristics.
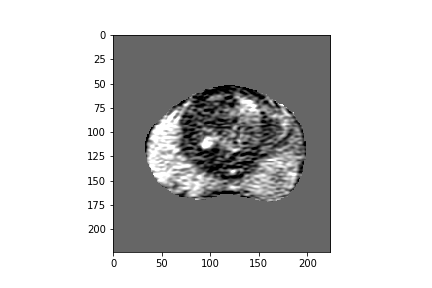
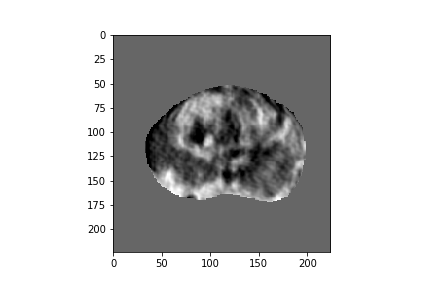
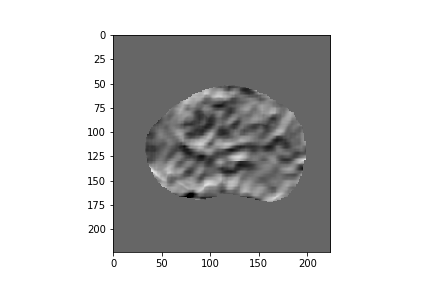
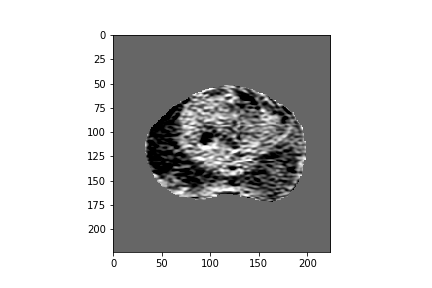
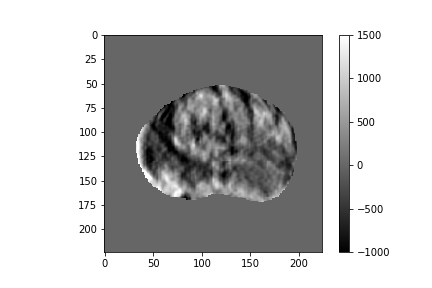
Figure 2: Five-dimensional CorrNet representations for one example MRI slice.
Utilizing a variety of hidden layer configurations, the CorrNet model projects MRI-derived features into a transformed space that aligns well with cancer signatures visible in histopathology data. These learned features subsequently inform a CNN-based predictive model that excels in predicting cancer probability maps across test cohorts.
Holistically Nested Edge Detection Adaptations
Building on this feature learning, CorrSigNet adopts variants of Holistically Nested Edge Detection (HED) models to translate correlated MRI features into actionable cancer probability maps. While HED-3 purely relies on CorrNet outputs, HED-branch-3 also harnesses original MRI intensities alongside CorrNet features for more comprehensive predictions.
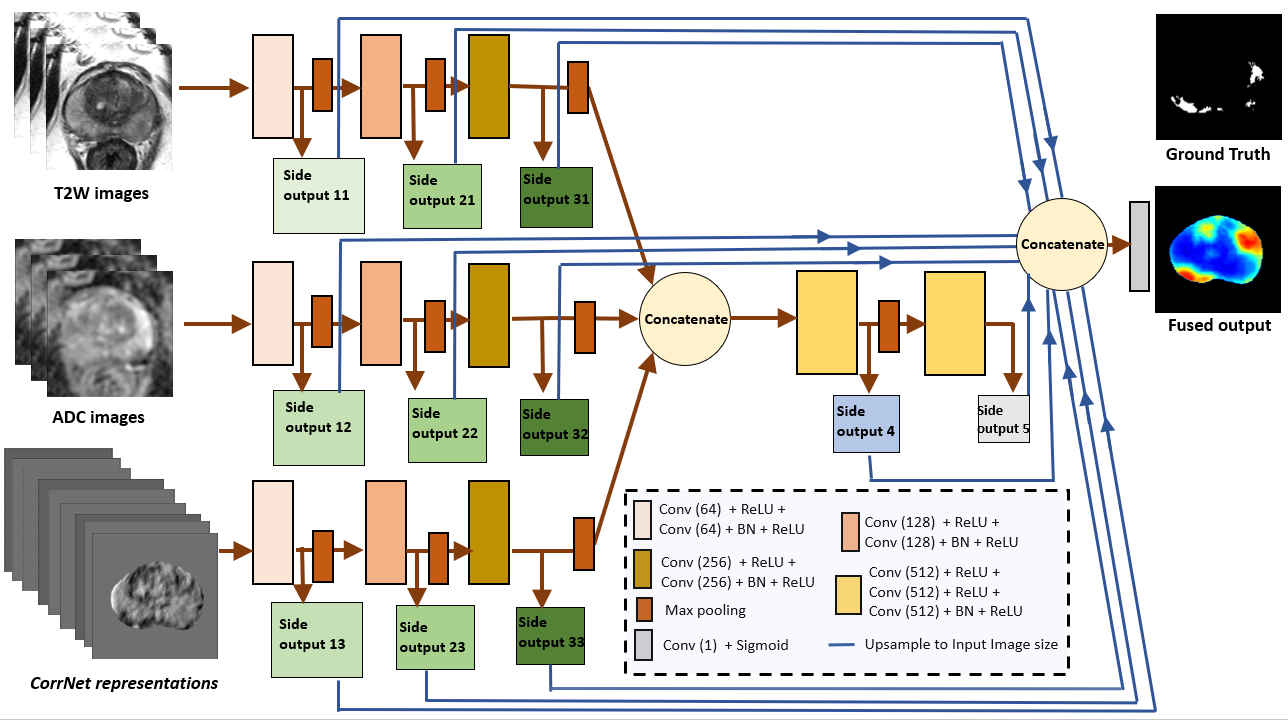
Figure 3: HED-branch-3 model for predicting cancer probability maps.
Results and Evaluation
Quantitative Analysis
CorrSigNet surpasses the current state-of-the-art in both pixel-level and lesion-level evaluations, achieving superior AUC scores (0.86 pixel-level, 0.96 per-lesion) compared to a previous benchmark at 0.80 and 0.92, respectively. This performance is manifest across various configurations, with at least 3-dimensional CorrNet features necessary for optimal model performance.
Qualitative Insights
Visualization of prediction maps demonstrates the enhanced ability of CorrSigNet to capture subtle cancer regions that current methods miss. The inclusion of correlated feature dimensions significantly reduces false positives, thus achieving a better overlap with ground truth annotations.
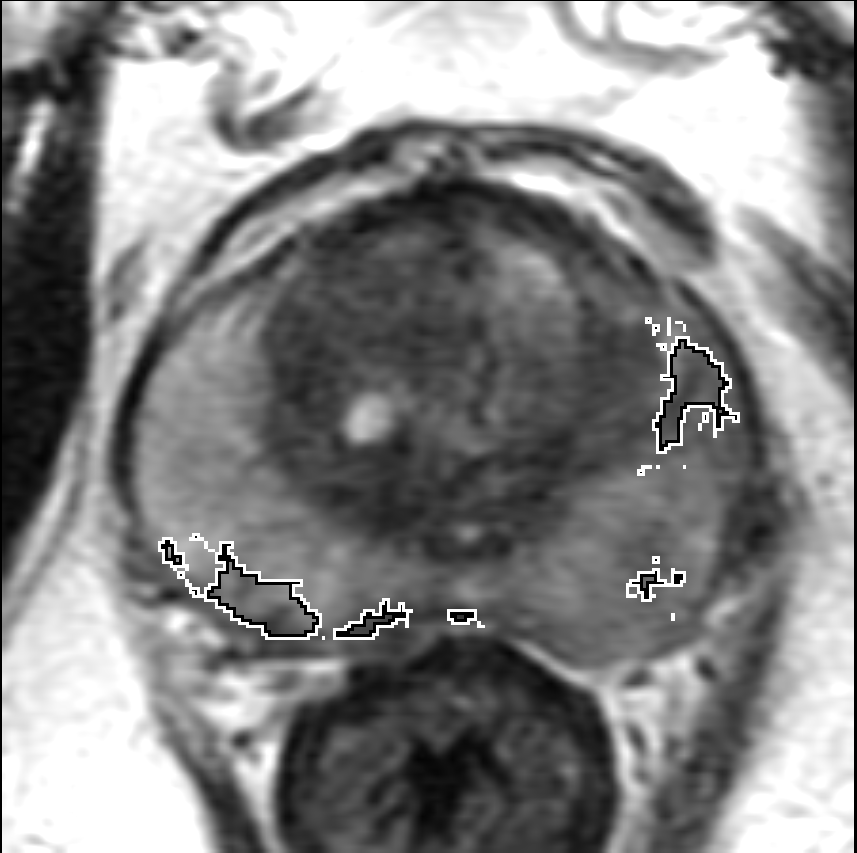
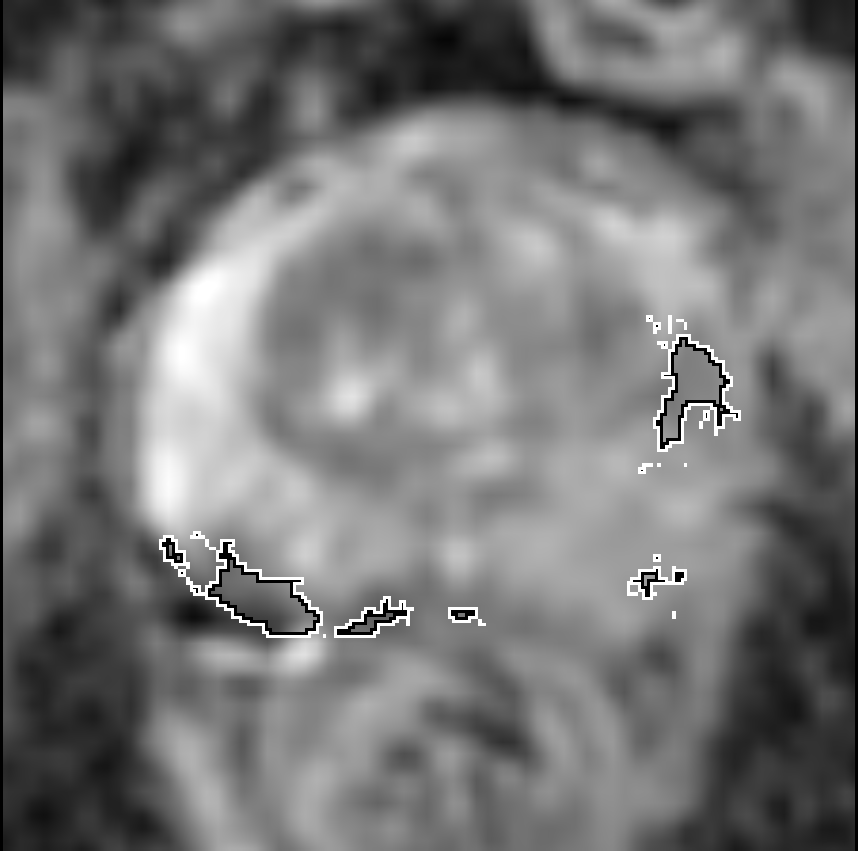
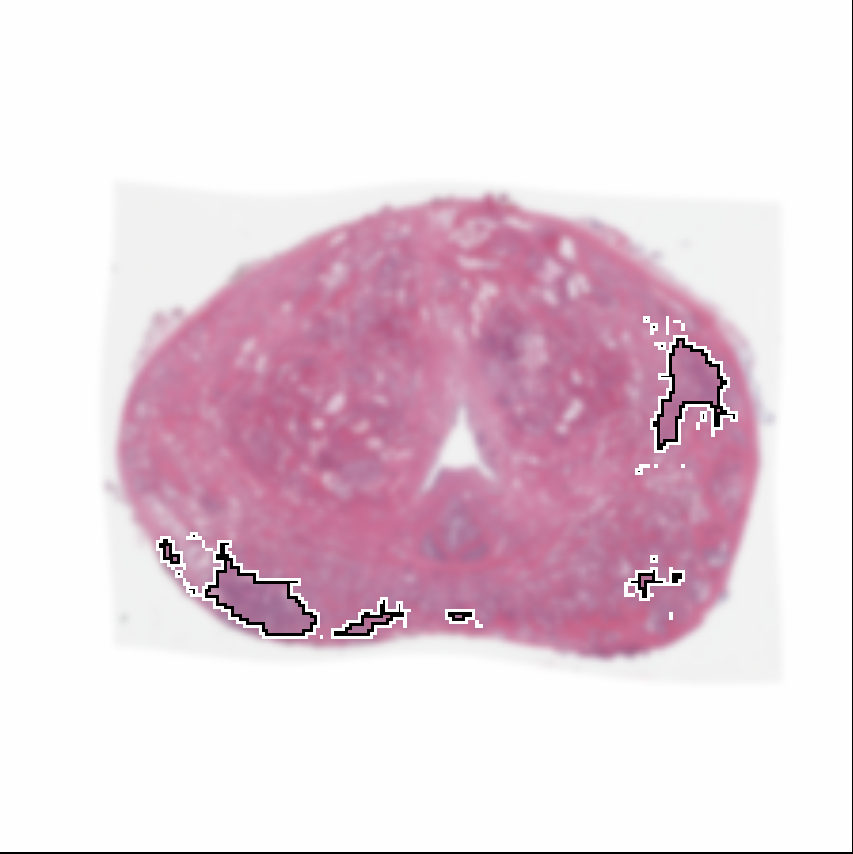

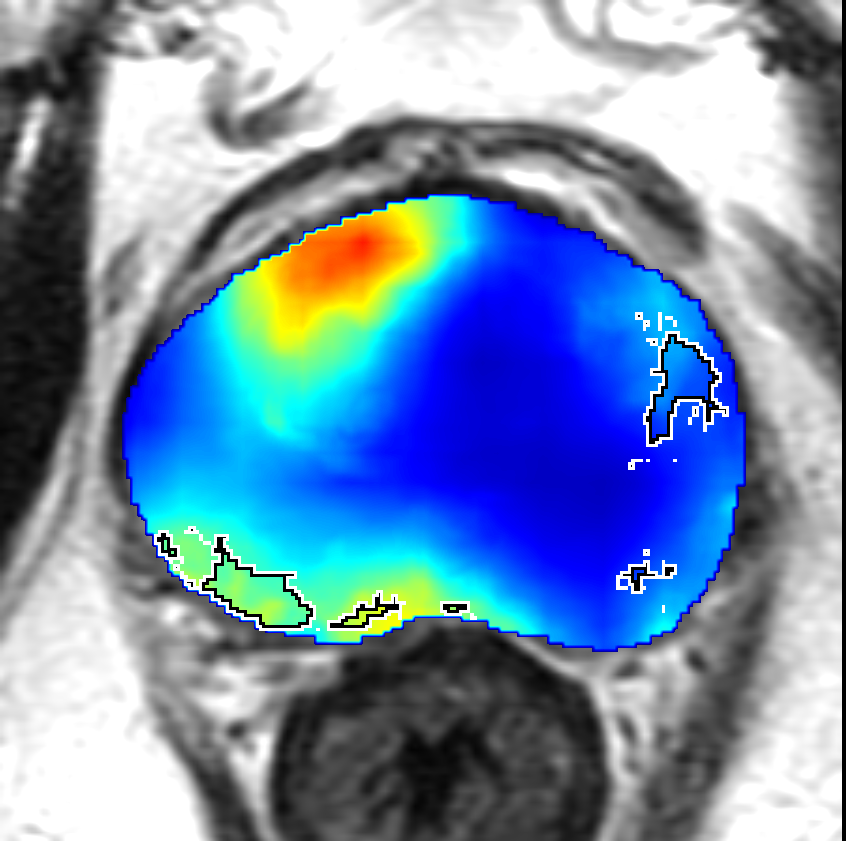
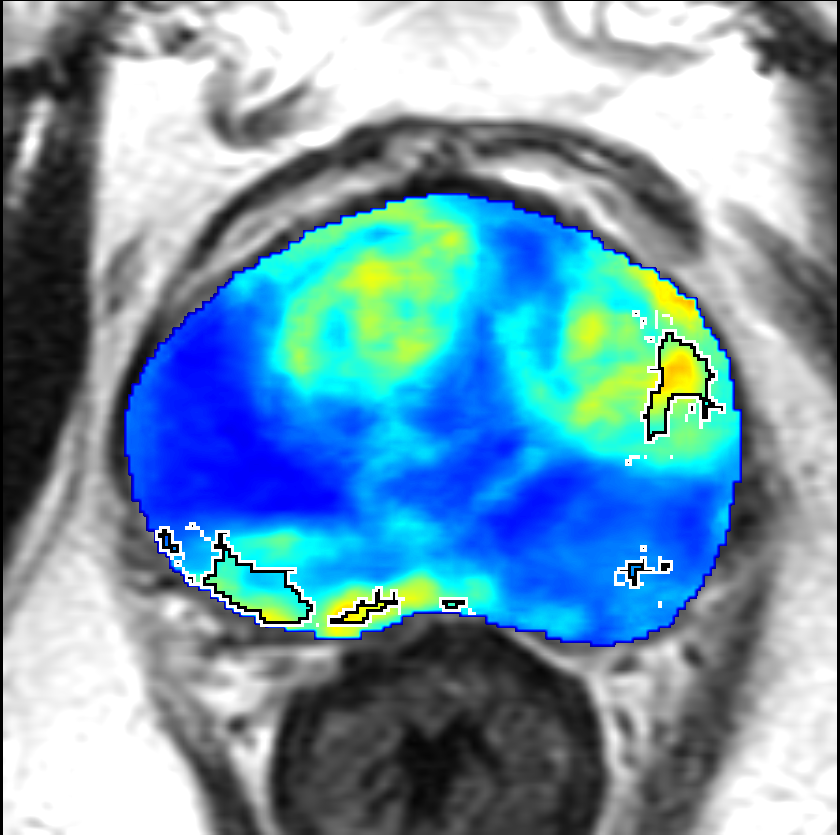
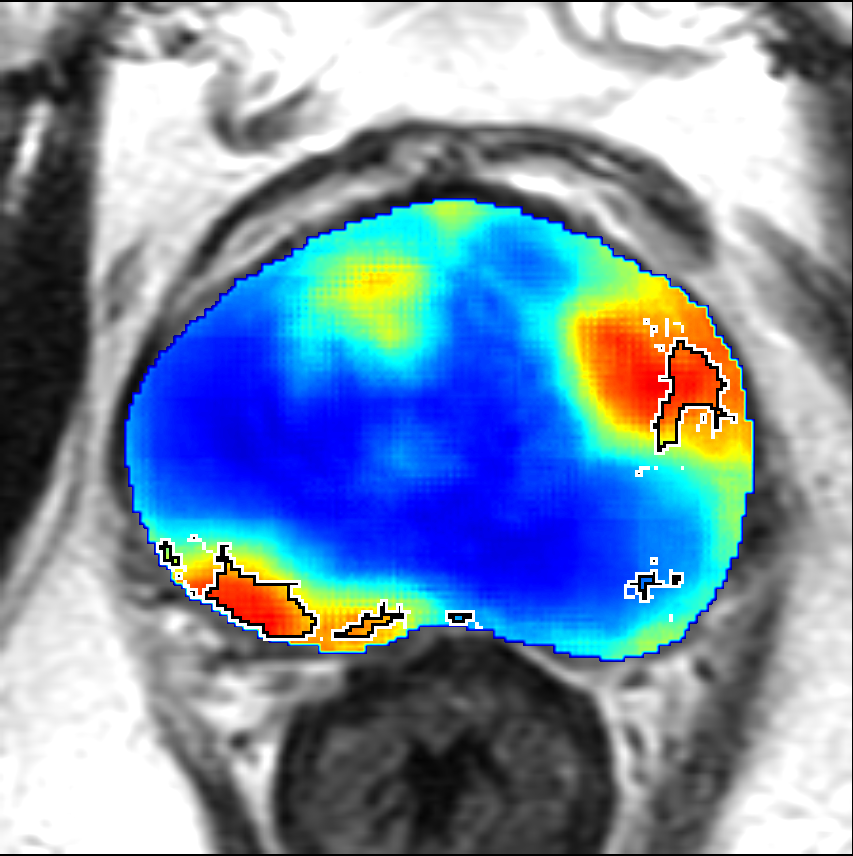

Figure 4: Prediction results using CorrSigNet models compared to current benchmarks.
Conclusion
CorrSigNet epitomizes advancements in prostate cancer imaging diagnostics by integrating cross-modality feature learning, resulting in refined diagnosis capabilities even in complex cases. The method offers a scalable approach suitable for datasets without comprehensive histopathology images, propelling future research into more nuanced, biologically-informed diagnostic tools. Further exploration with broader datasets and enhanced validation strategies can continue refining these models, bridging gaps between anatomical imaging and essential pathological insights.
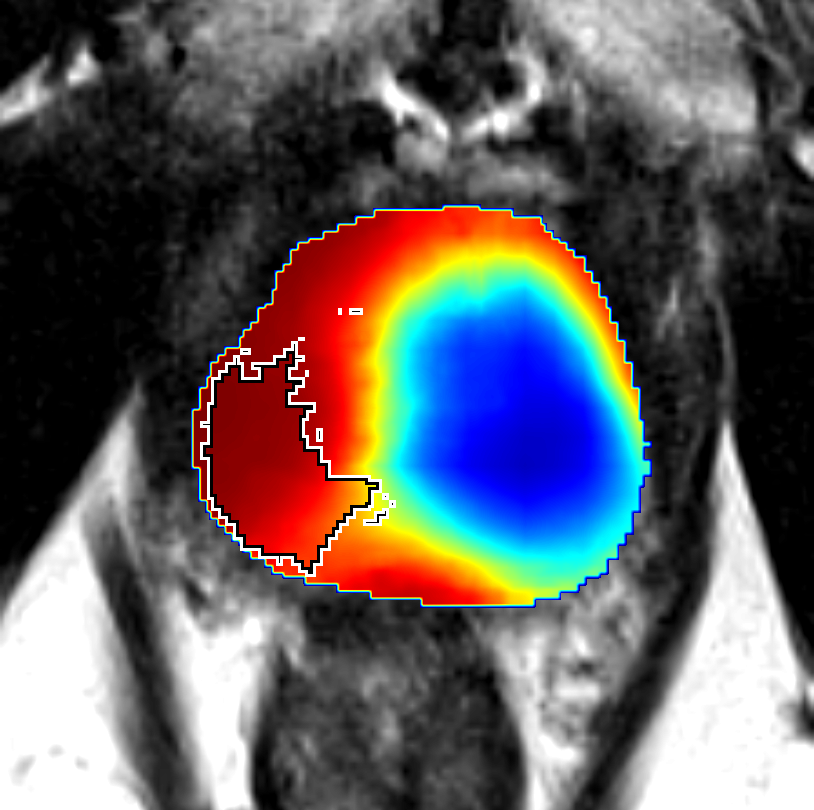
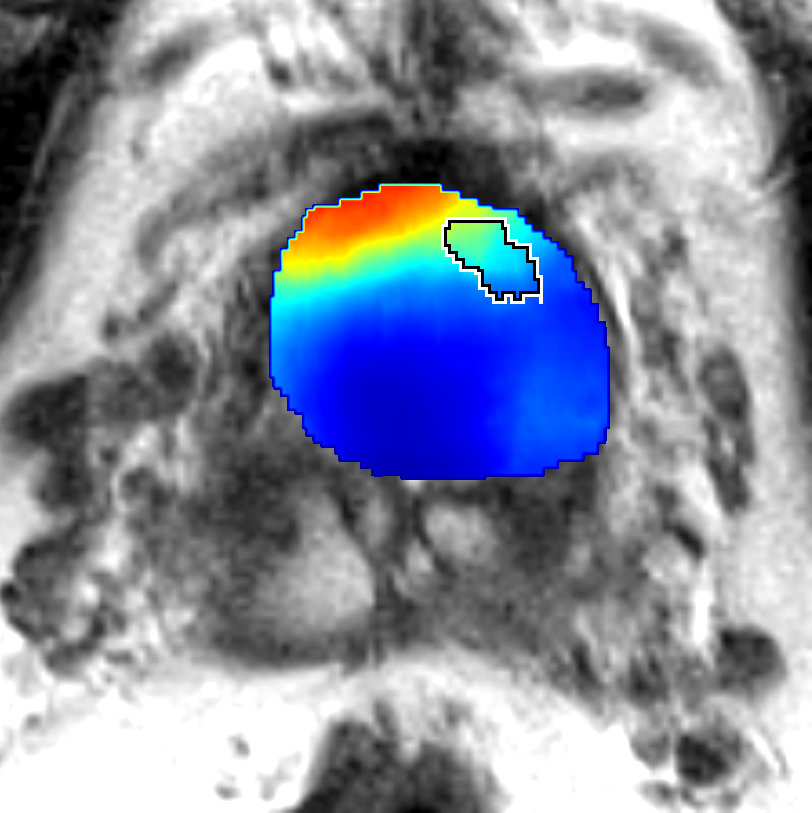
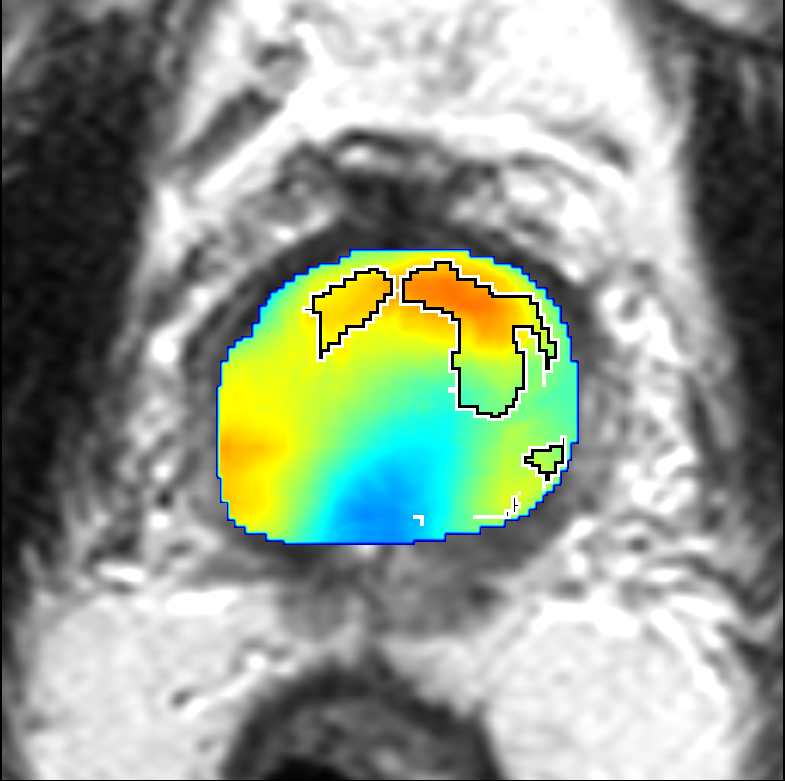
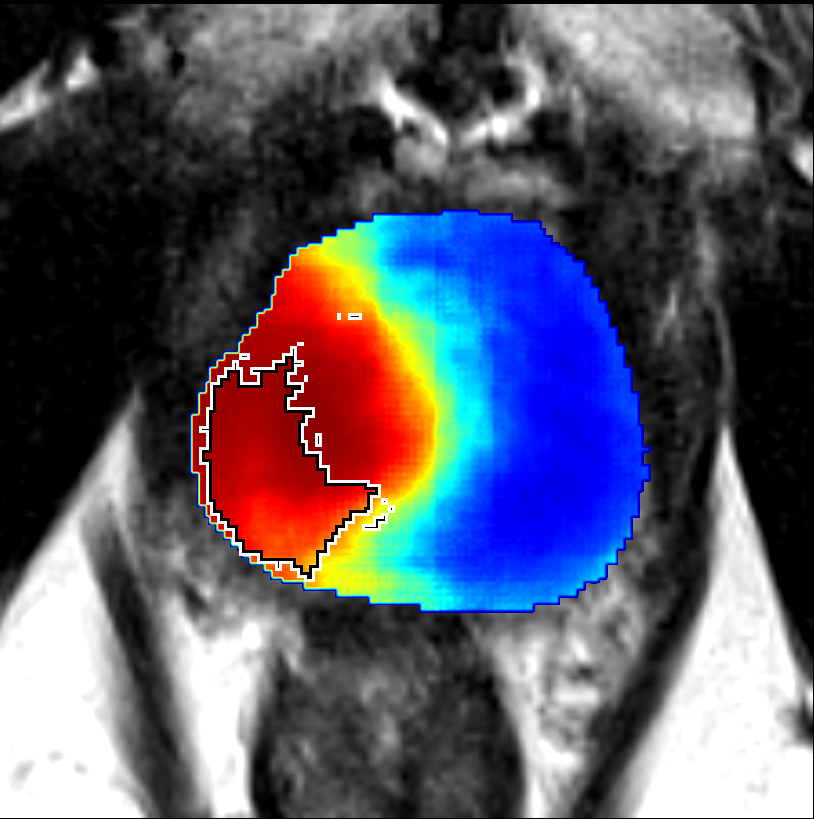
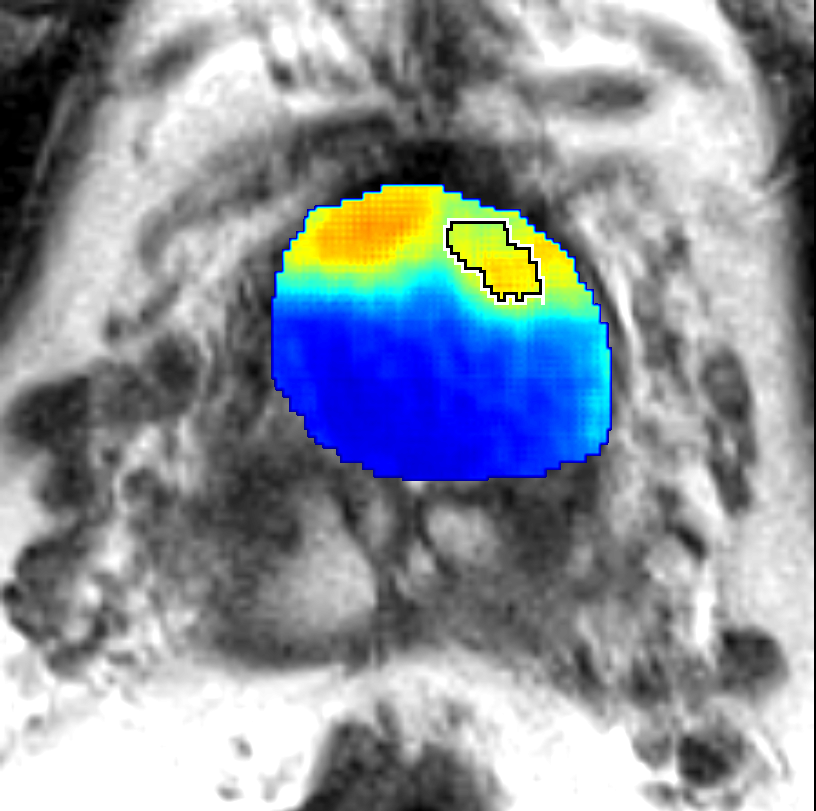
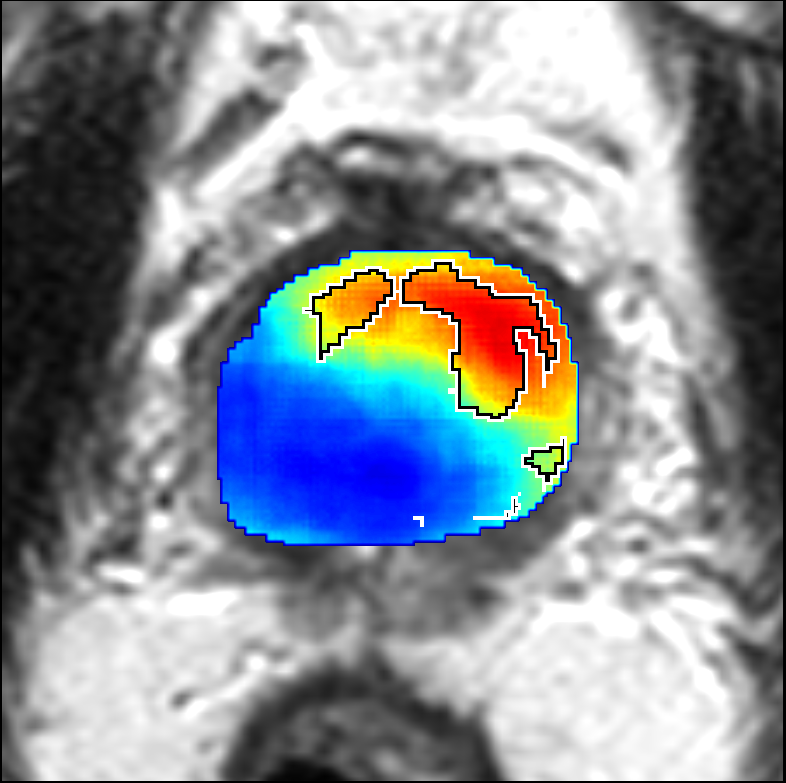
Figure 5: Comparison of CorrSigNet prediction results with existing state-of-the-art methods.





















