- The paper presents ProstNFound+, a medical foundation model adapted for μUS-based csPCa detection, achieving up to 79.1% AUROC in prospective validation.
- It employs adapter tuning, clinical metadata prompting, and a multi-head architecture to enhance lesion localization and risk scoring.
- The study demonstrates robust generalization with minimal performance drop across time and different clinical sites.
ProstNFound+: Prospective Validation of Medical Foundation Models for Prostate Cancer Detection
Introduction and Motivation
Prostate cancer (PCa) remains a leading cause of cancer-related mortality in men, with early detection of clinically significant PCa (csPCa) being critical for optimal patient outcomes. While multiparametric MRI (mpMRI) and the PI-RADS protocol are established for risk assessment, their high cost and limited accessibility motivate the search for scalable alternatives. High-resolution micro-ultrasound (μUS) with the PRI-MUS scoring system offers a promising, cost-effective imaging modality, but its reliance on operator expertise and associated inter-observer variability limit its generalizability. Deep learning approaches, particularly those leveraging medical foundation models (FMs), have demonstrated strong performance in medical imaging tasks, but their application to μUS-based PCa detection and, crucially, their prospective clinical validation, have been lacking.
Model Architecture and Methodology
ProstNFound+ is an adaptation of a medical FM for μUS-based PCa detection, integrating several key innovations: adapter tuning, a custom prompt encoder for embedding clinical biomarkers, and a multi-head output module for simultaneous cancer localization and risk scoring.
The model architecture is depicted in (Figure 1).
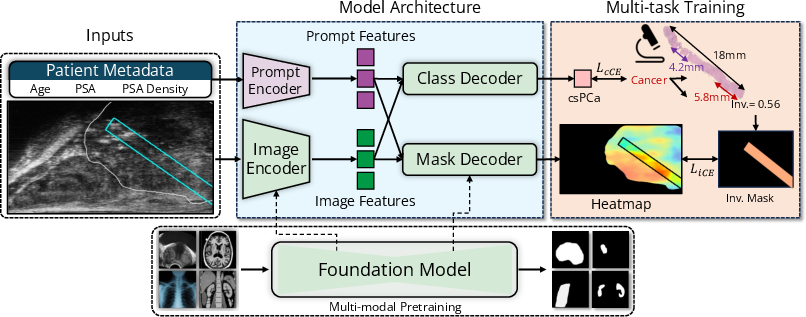
Figure 1: ProstNFound+ integrates a B-mode image encoder with conditional prompting using clinical metadata. The resulting embeddings are used by the mask decoder to generate a heatmap of cancer likelihood, and by the class decoder to output an image-level score representing the likelihood of clinically significant prostate cancer (csPCa).
The image encoder and mask decoder are derived from MedSAM, a vision transformer-based FM. The encoder processes 256×256 B-mode μUS images, outputting 256×64×64 embeddings. Clinical metadata (age, PSA, PSAD) are embedded via a two-layer MLP prompt encoder, producing 256-dimensional prompt vectors. These are concatenated with image embeddings and passed to the mask decoder for heatmap generation and to a class decoder for csPCa risk scoring.
The model is trained with a multi-task loss: cross-entropy for csPCa classification and a region-based loss for heatmap accuracy, using the average activation in the annotated needle region as a surrogate for cancer involvement. The csPCa risk output is discretized into a 1–5 scale to match PRI-MUS and PI-RADS conventions, enabling direct comparison with clinical protocols.
Experimental Design
The study utilizes a retrospective multi-center dataset for model development and a temporally and geographically distinct prospective dataset for evaluation. The retrospective set comprises 693 subjects and 6607 biopsy cores, while the prospective set includes 77 subjects and 1040 cores, with ground truth provided by histopathology. The prospective data were acquired five years after the training data at a new clinical site, ensuring a robust test of generalization.
Training employs five-fold cross-validation on the retrospective data, with model selection based on average performance. Baselines include patch-based ResNet, MicroSegNet, MedSAM-UNETR, Cinepro, and other FM adaptation strategies. Ablation studies assess the impact of clinical prompting and multi-head architecture.
Retrospective and Prospective Results
ProstNFound+ achieves the highest AUROC (77.5%) among all tested methods, outperforming both prior SOTA and other FM-based approaches. Sensitivity and specificity metrics are consistently strong across thresholds. Ablation studies demonstrate that conditional prompting with clinical features (age, PSA, PSAD) yields a +6% AUROC improvement over the baseline, and the multi-head architecture enhances both localization and risk assessment.
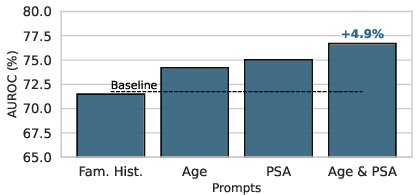
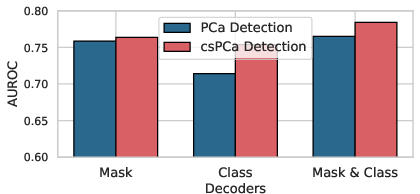
Figure 2: Summary of ablation experiments on clinical prompt combinations and model architectures, demonstrating the additive benefit of clinical metadata and multi-head design.
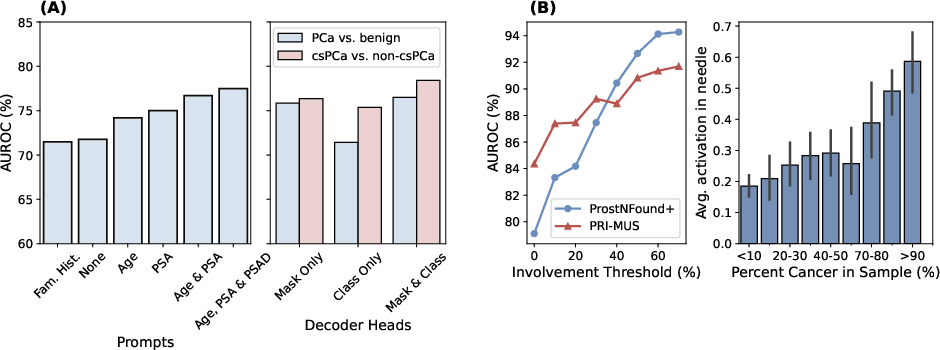
Figure 3: (A) Ablation study results. (B) csPCa detection performance and average heatmap activation by true cancer involvement, showing improved detection with higher tumor burden.
Prospective Validation
On the prospective test set, ProstNFound+ demonstrates robust generalization, with no significant performance degradation relative to retrospective evaluation. For csPCa detection, ProstNFound+ achieves an AUROC of 79.1%, compared to 84.4% for PRI-MUS and 88.1% for PI-RADS on the subset with MRI. Sensitivity is lower than PRI-MUS by 10%, but specificity is comparable. Notably, for biopsy samples with high tumor involvement (≥40%), ProstNFound+ matches or exceeds PRI-MUS performance.
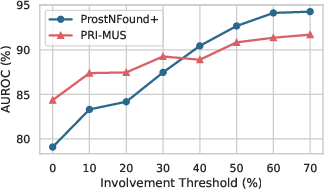
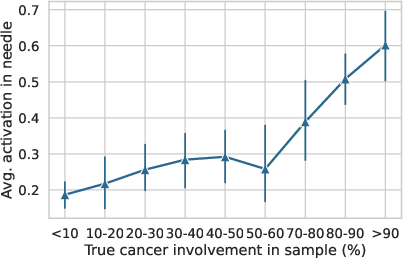
Figure 4: Left: Both PRI-MUS and ProstNFound+ show improved csPCa vs. non-csPCa classification on higher involvement cores. Right: Higher cancer involvement correlates with increased heatmap activation in the needle region.
Qualitative analysis of heatmaps confirms that ProstNFound+ localizes suspicious lesions with high activation in csPCa cases, aligning with biopsy-confirmed cancer regions.
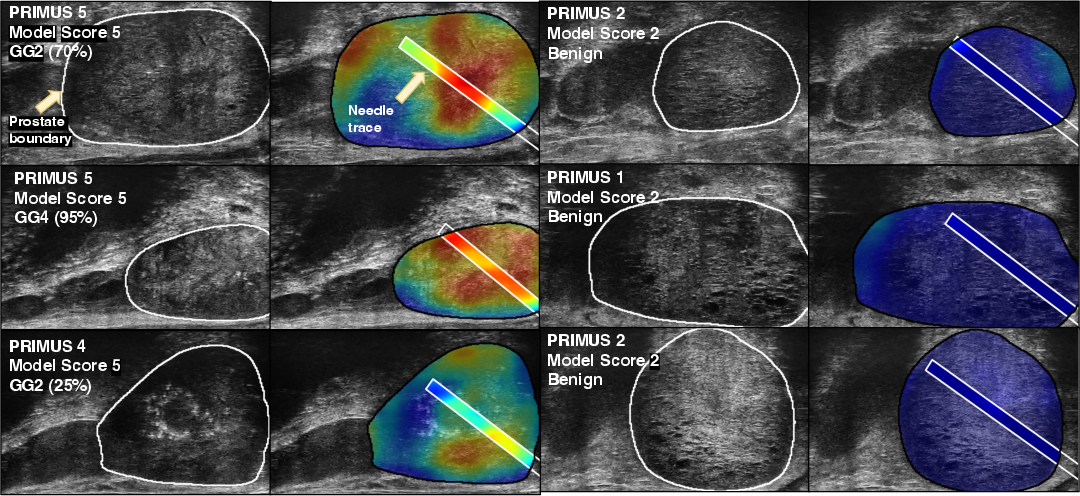
Figure 5: Example heatmaps generated by ProstNFound+, with high model and PRI-MUS scores corresponding to biopsy-confirmed cancer.
At the patient level, the highest model risk score per subject strongly correlates with csPCa diagnosis, and a score of 5 is highly specific for cancer. The model's risk scores align closely with PRI-MUS and PI-RADS, with few false positives in benign cases.
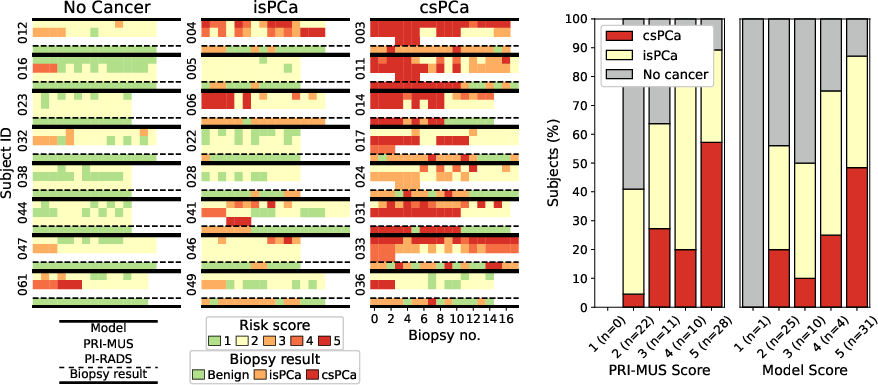
Figure 6: Left: Risk scores and results across biopsies for subjects in the prospective test set. Right: Distribution of subject-level diagnoses by highest PRI-MUS and model risk score.
Discussion and Implications
ProstNFound+ demonstrates that medical FMs, when adapted with domain-specific prompting and multi-task objectives, can approach the diagnostic performance of expert-driven visual protocols in μUS-based PCa detection. The model's generalization to a temporally and geographically distinct prospective cohort, without performance degradation, is a strong indicator of its robustness and potential for clinical deployment.
The performance gap with PRI-MUS (approximately 5% lower AUROC) is modest, especially considering the extensive training required for PRI-MUS expertise and the operator-agnostic nature of the model. For high-involvement tumors, the model achieves parity with expert scoring. The interpretable heatmaps and risk scores facilitate both targeted biopsy and global risk assessment, supporting integration into clinical workflows.
Limitations include reliance on core-level labels (limiting pixel-level localization accuracy) and analysis of 2D images in isolation, whereas clinical protocols often leverage 3D gland-wide assessment. Future work should explore training with pixel-level annotations, 3D network architectures, and integration of temporal or volumetric data. Additionally, combining model outputs with expert visual scores may further improve diagnostic accuracy.
Conclusion
ProstNFound+ provides a robust, interpretable, and scalable approach to μUS-based prostate cancer detection, validated in a prospective clinical setting. Its performance approaches that of established visual protocols, with strong generalization across time and site. The model's operator-agnostic design and interpretability position it as a promising tool for standardizing and expanding access to high-quality PCa diagnosis, particularly in settings where MRI or expert interpretation is limited. Further advances in annotation granularity and 3D modeling are likely to close the remaining performance gap and enhance clinical utility.







