- The paper introduces a novel weakly-supervised learning approach leveraging negative mp-MRI scans to synthesize normal prostate images for cancer inference.
- The CNN-based U-Net model utilizes prevalence maps and distance metrics like ADC Increment to achieve an AUC of 0.84 and 77.0% detection sensitivity.
- The methodology offers an annotation-free, scalable diagnostic tool with potential for multi-institutional use and application to other imaging domains.
Prostate Cancer Inference via Negative MRI and Weakly-Supervised Learning
Introduction
This paper focuses on developing a novel machine learning approach for prostate cancer (PCa) inference through weakly-supervised learning leveraging negative multi-parametric MRI (mp-MRI) scans. It tackles the challenge of limited lesion annotation availability, which restricts traditional machine learning models in medical imaging. By using a large dataset of radiology-confirmed negative MRI cases, the authors propose a model that synthesizes normal prostate appearances, facilitating pixel-wise cancer suspiciousness inference by comparing synthetic and original images.
Baseline MRI Model Development
The baseline MRI model, a convolutional neural network (CNN) structured as a U-Net, is trained to capture the normal prostate mp-MRI appearance using only negative scans. This model synthesizes partially-obstructed regions via weakly-supervised learning, avoiding the need for positive lesion annotations (Figure 1).
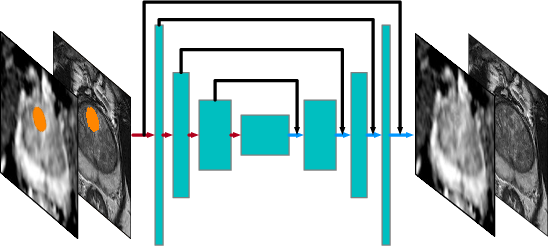
Figure 1: The baseline MRI model synthesizes for the partially obstructed region M (shown in orange) using an unobstructed image as input.
The structured encoder-decoder design enhances anatomical summarization, while skip connections simplify training and partial convolutional layers compensate for zeroed-out inputs. The prevalent appearances are captured through ROI candidates, summarized in a prevalence map, aiding in synthesizing the suspected regions (Figure 2).
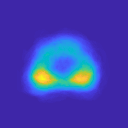
Figure 2: The prevalence map, P, constructed by the collection of ROI candidates, accounting for the common locations and shapes of prostate cancer.
Cancer Inference Method
In the inference stage, the trained model predicts pixel-wise cancer suspiciousness for each testing image. Suspicious regions are highlighted where synthesized images differ significantly from originals, indicating potential PCa locations. The paper proposes two distance metrics, T2-weighted SSIM and ADC Increment, utilizing structural and intensity variations to compute suspiciousness maps (Figure 3).
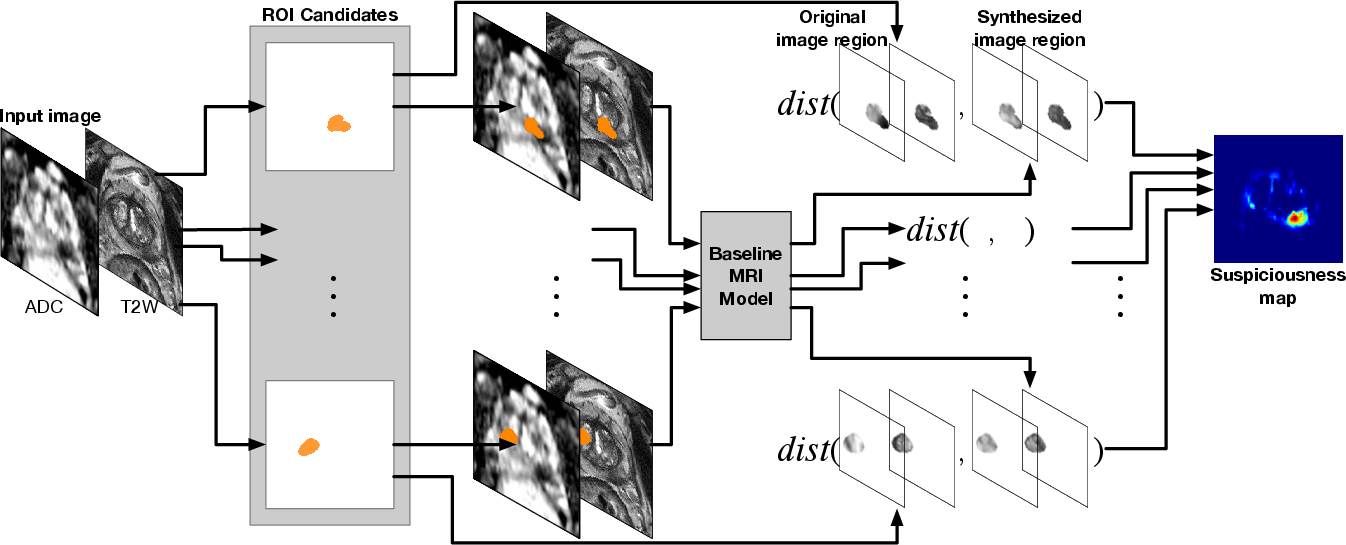
Figure 3: The inference of the PCa suspiciousness map using the trained baseline MRI model given an input testing image. The baseline MRI model synthesizes regions specified from the collection of ROI candidates.
For each imaging parameter (T2w and ADC), differences between real and synthesized versions are evaluated, producing maps indicative of suspicious areas. Strong results were particularly observed with ADC Increment metrics, showing high PCa predictability (Figure 4).
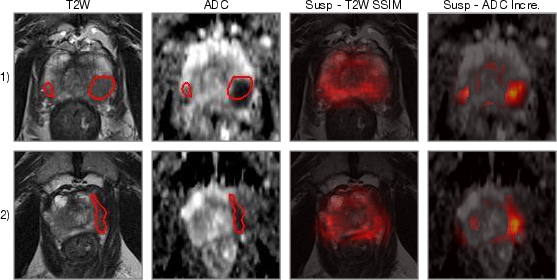
Figure 4: The PCa suspiciousness maps with different distance functions for testing images. The red contours on T2w and ADC are the ground truth ROIs.
Experimental Results
Evaluation with receiver operating characteristic (ROC) and free-response receiver operating characteristic (FROC) analyses was conducted on a 232-case dataset consisting of positive and negative MRI scans. The model achieved a remarkable area under curve (AUC) of 0.84 through ADC Increment, outperforming traditional models requiring extensive annotated training data (Figure 5).
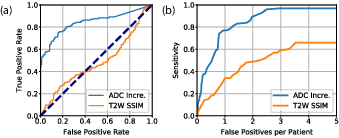
Figure 5: (a) ROC analysis for the classification between PCa lesions and non-PCa regions. (b) FROC analysis for lesion localization performance.
With a detection sensitivity of 77.0% at one false positive per patient in FROC analysis, the approach competes with fully-supervised learning models, demonstrating the potential viability of weakly supervised learning for scalable medical image analysis.
Discussion
This method introduces a scalable, annotation-free strategy in medical imaging, relying solely on negative mp-MRI scans for training. Such negative scan abundance proffers practical advantages, facilitating multi-institutional collaborations without inconsistency concerns inherent in annotated datasets. The discriminative synthesis approach marks a significant shift from annotation-dependent frameworks, stressing the innovation potential in deep learning applications in healthcare.
Future work might explore diverse distance functions to enhance prediction accuracy further and address potential data biases. Moreover, extending this framework to other anatomical or pathological domains maintains promising prospects for non-invasive diagnostics.
Conclusion
Prostate cancer inference through weakly-supervised learning utilizing negative MRI successfully distinguishes between PCa and non-PCa regions without annotated lesions. The robust baseline MRI model presents a scalable alternative, boasting competitive diagnostic performance, with significant implications for future deployments in medical image analysis. The model's effectiveness suggests an opportunity for advancing non-invasive cancer diagnostics, emphasizing the integration of innovative imaging and inferential techniques.






