- The paper demonstrates a unified approach that interleaves HTML tags, cell text, and discretized coordinates into a single sequence for table extraction.
- It employs a lightweight CNN backbone with a transformer encoder and a structure-prior head to ensure precise alignment of logical structure and spatial geometry.
- Empirical results show state-of-the-art performance on benchmarks with improved decoding speed via multi-token prediction and robust generalization across datasets.
TableSeq: Unified Generation of Structure, Content, and Layout
Introduction and Problem Statement
"TableSeq: Unified Generation of Structure, Content, and Layout" (2604.16070) addresses image-based table structure recognition (TSR), cell content extraction (TCR), and cell localization via a unified, sequence generation-based architecture. Table understanding from documents is non-trivial due to visually complex layouts, missing or weak visual separators, misaligned headers, merged cells (spans), and annotation inconsistencies. Existing approaches typically fragment the problem into separate modules or stages, hindering end-to-end optimization and increasing engineering complexity.
TableSeq advances the field by producing an interleaved token stream comprising HTML markup, cell textual content, and discretized cell coordinates. The method operates exclusively on table images, without reliance on external OCR engines or multiple decoders, and produces globally aligned logical structure, content, and cell geometry within a single autoregressive sequence.
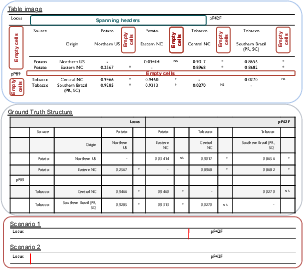
Figure 1: PubTabNet sample with missing column separators, weak line cues, and header misalignment, illustrating layout ambiguity and challenge for TSR.
Methodology and Model Architecture
The TableSeq architecture features a lightweight CNN backbone (FCN-H16), augmented by a minimal structure-prior head and a single-layer transformer encoder. The CNN maintains fine vertical resolution (H′/16) critical for document tables, filtering and extracting local structural features. The structure-prior head predicts soft geometric cues: row separators, column separators, and cell corners. These cues bias the decoder's cross-attention via a key-only additive mechanism during decoding, facilitating attention alignment to plausible table grid locations.
A BART-style autoregressive decoder generates a sequence that interleaves HTML table tags, cell text content, and discretized coordinate tokens, providing direct alignment between logical structure and physical localization. Positional embedding is implemented using 2D RoPE, ensuring spatial information is preserved throughout the sequence modeling. The model’s design is strikingly minimalist but empirically robust, in contrast to common multi-decoder or pointer-heavy alternatives.
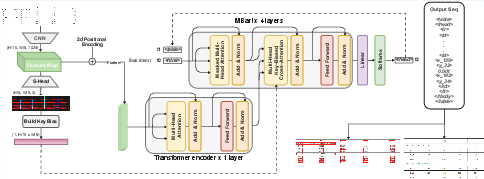
Figure 2: TableSeq's end-to-end pipeline: input images are encoded by a CNN and single-layer transformer; the structure-prior head biases decoding; a single decoder produces HTML tags, cell contents, and coordinates as one sequence.
Supervision interleaves HTML tags (e.g., <thead>, <tr>, <td>), cell textual content, and quantized position tokens (<x_k>, <y_k>, with 5\,px granularity). The embedding analysis (via UMAP/PCA) reveals that colspan and rowspan tokens are encoded with clear semantic and cardinality separation, indicating the model's internal consistency regarding span semantics.
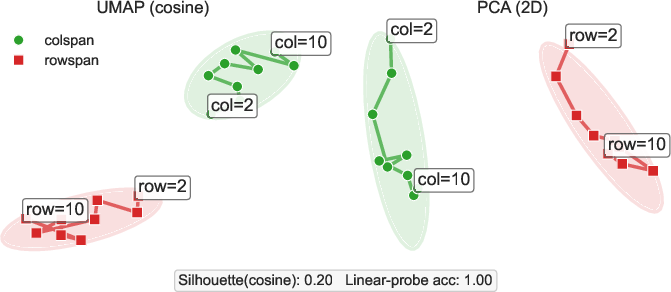
Figure 3: Span token embeddings exhibit distinct clusters and monotonic trajectories for increasing span (k), confirming robust disentanglement of column/row spans in the learned representation.
Training Protocol and Synthetic Data
The training objective combines token-level cross-entropy for sequence prediction with auxiliary losses (binary cross-entropy + Dice) for the structure-prior head, promoting geometric attention alignment. Training utilizes a synthetic-to-real ratio annealing schedule: initially, 100% synthetic tables are used (with structure/content augmentation and in-place editing to diversify fonts, backgrounds, and layouts) and the proportion is gradually reduced.
The synthetic pipeline edits table images and HTML in situ, retaining visual fidelity and preserving layout realism. Explicit structure-aware operations (span expansion/insertion, nested headers, groupings) increase generalization to complex real-world tables.
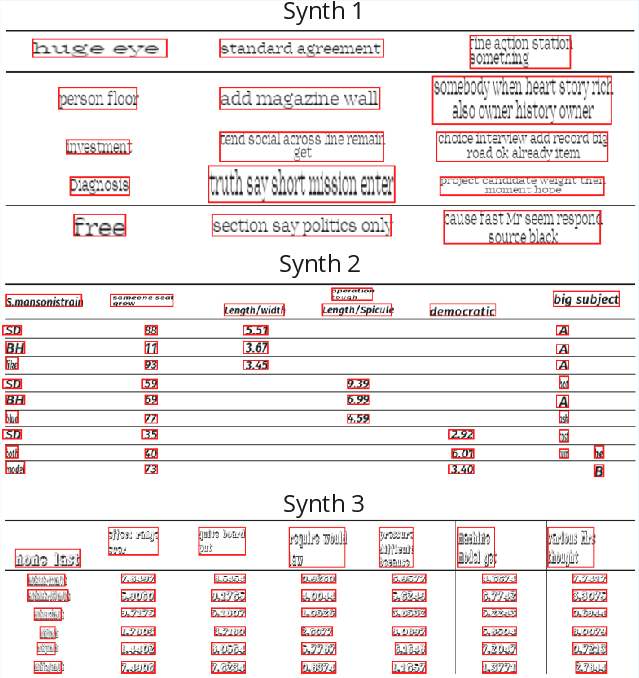

Figure 4: Synthetic, in-place table edits diversify content, formatting, and layout while preserving ruling lines and page background.
Structure Prior and Supervision
Training supervision for structure-prior maps is derived from ground-truth HTML and bounding boxes, ensuring predicted geometric cues (row/col/corner maps) are tightly aligned with underlying structure. At inference, these priors act as additive key-biases to the decoder cross-attention, incentivizing geometric consistency during sequence generation.
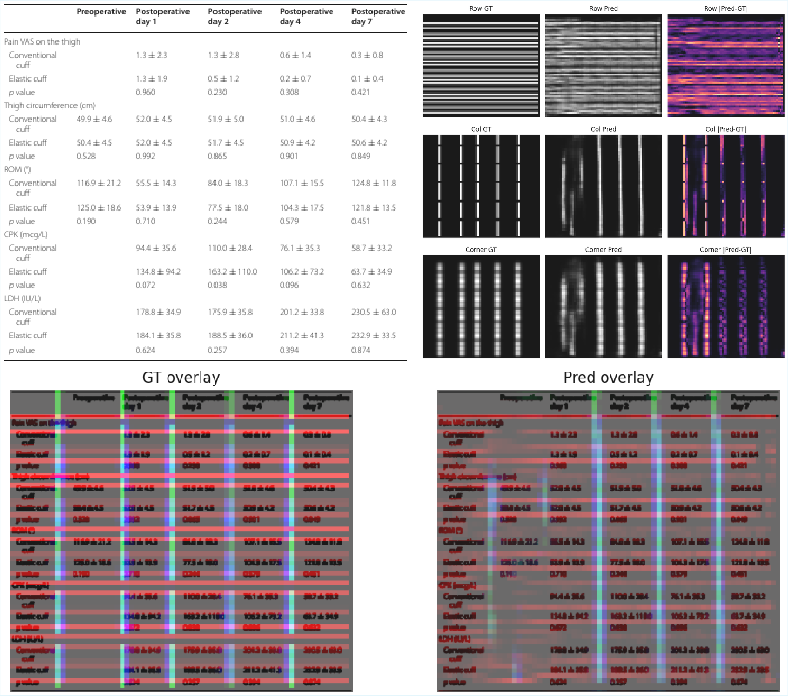

Figure 5: Example of predicted structure prior maps (rows, columns, corners) and corresponding supervision targets, showing the geometric alignment cues injected during decoding.
Experimental Results
TableSeq is evaluated on PubTabNet, FinTabNet, SciTSR, and PubTables-1M using TEDS, S-TEDS, CAR, and GriTS metrics. On SciTSR, TableSeq achieves state-of-the-art precision and F1 (99.79/99.66), with the second-highest recall, outperforming both image-to-sequence and modular approaches. On FinTabNet, TableSeq delivers the best S-TEDS in the image-to-sequence category.
In cross-dataset evaluation (zero-shot to ICDAR 2013) and under grid-based measures (GriTS) on PubTables-1M, TableSeq ranks at or near the top in structure/content and localization fidelity, demonstrating transferability and robustness. For cell localization (AP50), TableSeq obtains the highest reported accuracy (96.5%).
Representative predictions on scientific, financial, and grouped-row tables validate qualitative robustness and expose residual limitations on ambiguous span allocation and regionally empty structures.
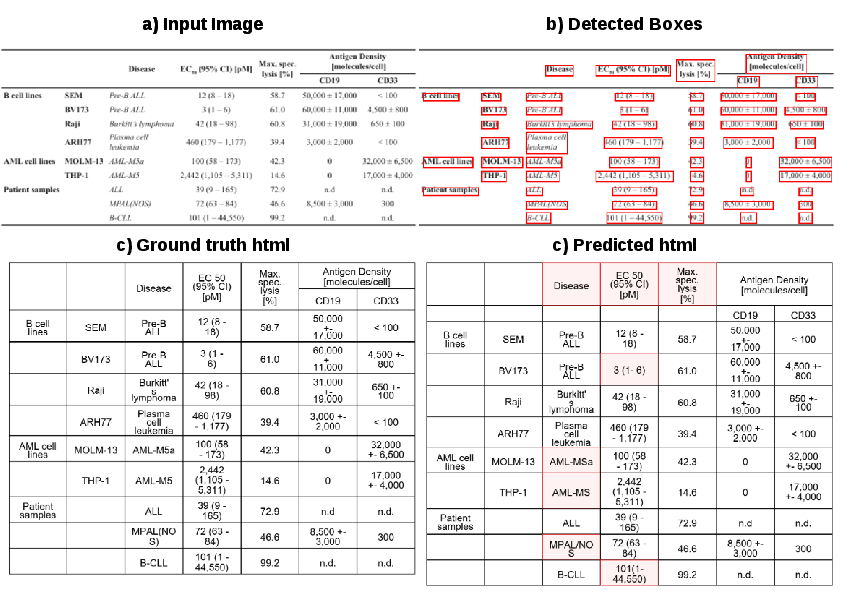
Figure 6: Prediction on a dense scientific table; TableSeq accurately recovers structure and content except in regions with empty cells or complex groupings.
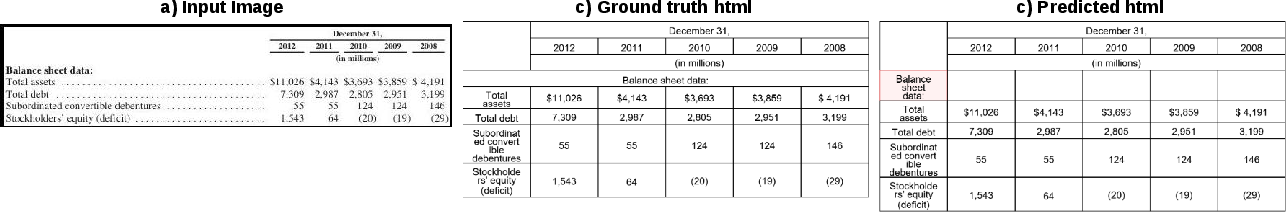
Figure 7: Financial table prediction; headers and values are well predicted, but the stub and blank header regions remain partially ambiguous.

Figure 8: Prediction on grouped-row table; global structure is coherent, but row-span/group allocation is not fully resolved due to ambiguity.
Sequence Interface, Multi-token Decoding, and Generalization
Notably, TableSeq's autoregressive emission of a single sequence unifies structure and content, facilitating index-based querying without architectural modification. On index-based data extraction tasks (IRDR, ICR, ICDR), TableSeq substantially surpasses large vision–language foundation models (e.g., Llama-3.2, GPT-4o, Gemini) on IRDR and matches or outperforms for ICR/ICDR, demonstrating the competitive advantage of a targeted, structure-aware image-to-sequence model.
The paper introduces multi-token prediction (MTP), analogous to blockwise LLM generation, yielding up to 47% reduction in decoding time with less than 1.4 S-TEDS point degradation at 4 tokens/step—providing a practical trade-off between throughput and quality.
Ablations and Architectural Insights
A comprehensive ablation study demonstrates that a compact FCN-H16 backbone outperforms parameter-matched shallow Vision Transformer (ViT) models for this sequence formulation, underlining the importance of high-resolution local features. The impact of vertical sampling, the structure-prior head, transformer encoder depth, biasing mechanisms, and synthetic data is systematically quantified, with each component contributing nontrivial accuracy gains.
Additionally, TableSeq's discretized tokenization of coordinates introduces a favorable speed–accuracy trade-off; finer bins marginally improve localization but increase sequence length and inference cost. Key-biased cross-attention yields consistent benefits and is robust to hyperparameter variation.
Practical and Theoretical Implications
TableSeq bridges logical and spatial table understanding in a single-stage, image-only architecture without external OCR or post-processing, reducing engineering overhead and improving reproducibility. Practically, this design is advantageous in domains with complex table layouts (biomedical, financial, technical) and provides a blueprint for unified markup/geometric extraction tasks.
Theoretically, the results endorse the sufficiency of sequence models—when equipped with bias-aware attention and structure-aligned supervision—for complex document parsing. TableSeq's ability to generalize to index-based queries implicates broader utility for document intelligence tasks beyond table recognition.
Limitations and Future Directions
Autoregressive inference remains a limitation; while MTP accelerates decoding, sequence length still constrains latency. Coordinate quantization, while simplifying alignment, may miss subtle boundary nuances. TableSeq does not explicitly address arbitrarily rotated or polygonal (non-rectangular) tables, which would require further architectural extensions.
The evaluation scope is bounded by available benchmarks; larger, more diverse, or low-resource script settings are not exhaustively tested. Extensions leveraging stronger visual pretraining, polygonal geometry modeling, and integration of global document context represent promising directions.
Conclusion
TableSeq establishes a strong single-stream baseline for end-to-end table structure, content, and layout extraction. The method's simplicity, generalizability, and competitive empirical performance confirm the effectiveness of unified autoregressive modeling for table recognition tasks. The model’s architectural principles—compact high-resolution encoding, lightweight transformer integration, structure-guided cross-attention, and single-sequence target design—mark a significant progression for unified document intelligence applications.









