- The paper introduces VoxelFM, a 3D vision transformer pretrained via self-distillation (DINO) to generate robust CT visual features without any language supervision.
- It demonstrates superior performance on tasks such as classification, regression, segmentation, and report generation using frozen backbone embeddings with lightweight probes.
- Extensive evaluation shows its efficiency under label scarcity and variability, outperforming recent CT foundation models across multiple clinical benchmarks.
Robust Visual Feature Learning in CT: VoxelFM for Efficient Transfer Across Clinical Tasks
Introduction and Motivation
Computed tomography (CT) imaging is central to diverse diagnostic, quantification, and prognostic tasks in clinical medicine. Recent CT foundation models have trended towards vision-language paradigms, leveraging CLIP-style objectives to align image and report embeddings. However, paired CT-report data are limited, constraining the efficacy of vision-language alignment. In most existing models, adaptation to new downstream tasks requires full or partial backbone fine-tuning, a process that demands substantial computational resources.
This work introduces VoxelFM, a 3D vision transformer (ViT) foundation model, pre-trained on over 137,000 CT studies via self-distillation using the DINO framework, entirely without language supervision. The core assertion is that robust, generalizable visual representations learned in a language-free paradigm can enable efficient transfer to downstream clinical tasks using only lightweight probes, with no backbone adaptation, and minimal task-specific labelled data requirements.
Pre-training Paradigm and Architecture
VoxelFM employs a 3D ViT backbone with rotary positional encodings, trained with a mix of DINO and iBOT self-distillation losses, targeting both class and patch token consistency. The training augmentation regime encompasses heavy spatial, intensity, and slice-order perturbations, fostering invariance to scanner protocol and anatomical variability. Global (class token) and local (patch token) representations are learned simultaneously, supporting both holistic and spatially resolved downstream tasks.
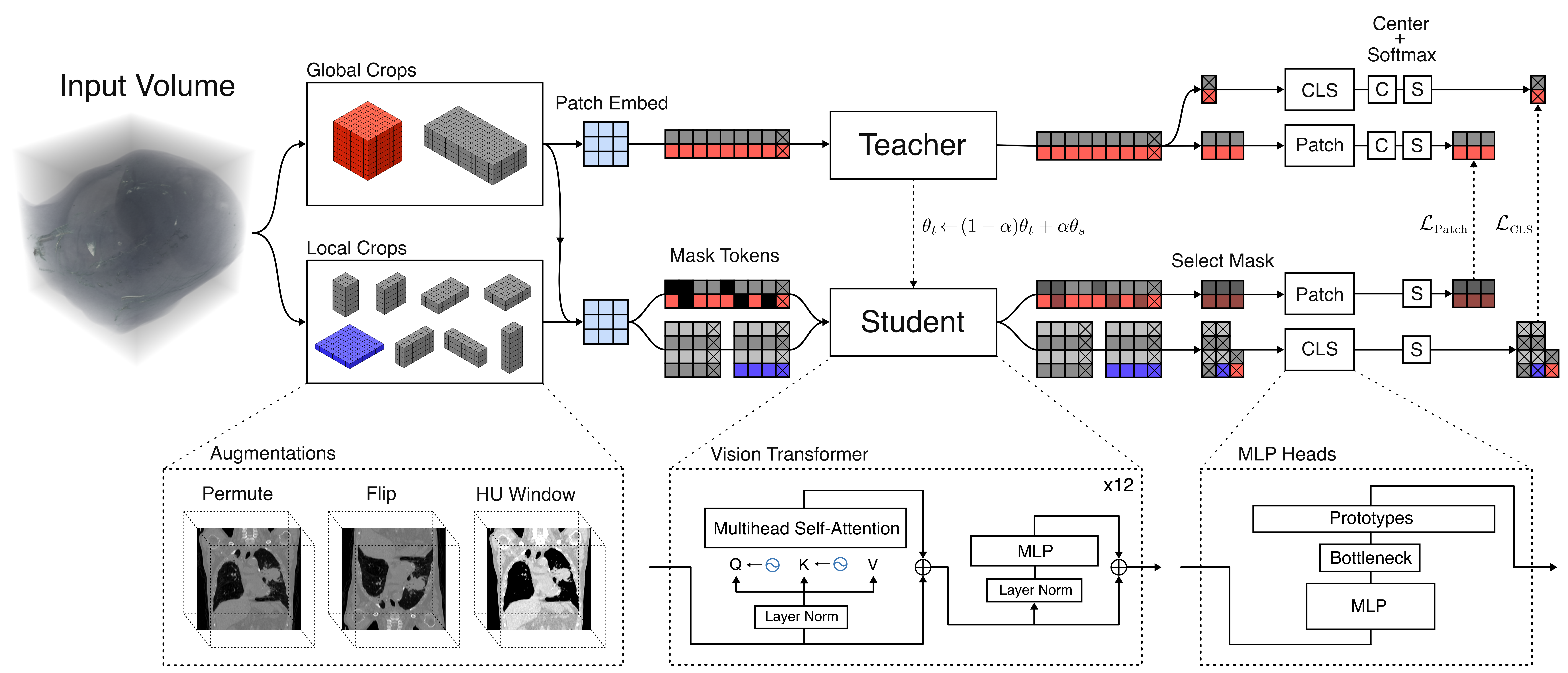
Figure 1: Overview of the VoxelFM DINO-based pre-training: a student-teacher ViT operates on multi-scale global and local crops with hybrid class and patch-wise distillation objectives.
Evaluation Protocol
The evaluation strategy uniformly exploits frozen VoxelFM backbone embeddings across seven clinically relevant task categories:
- Classification (various disease or abnormality detection benchmarks)
- Regression (quantitative biomarker estimation)
- Survival analysis (prognostication)
- Instance retrieval (matching scans by clinical phenotype)
- Localisation (lesion centroid prediction)
- Segmentation (multi-organ and lesion delineation)
- Report generation (vision-language performance via multimodal LLM adaptation)
For each task, only lightweight probe heads (MLPs or Q-Formers) are trained, keeping VoxelFM weights fixed. This isolates the utility of pre-trained representations from confounding by backbone fine-tuning.
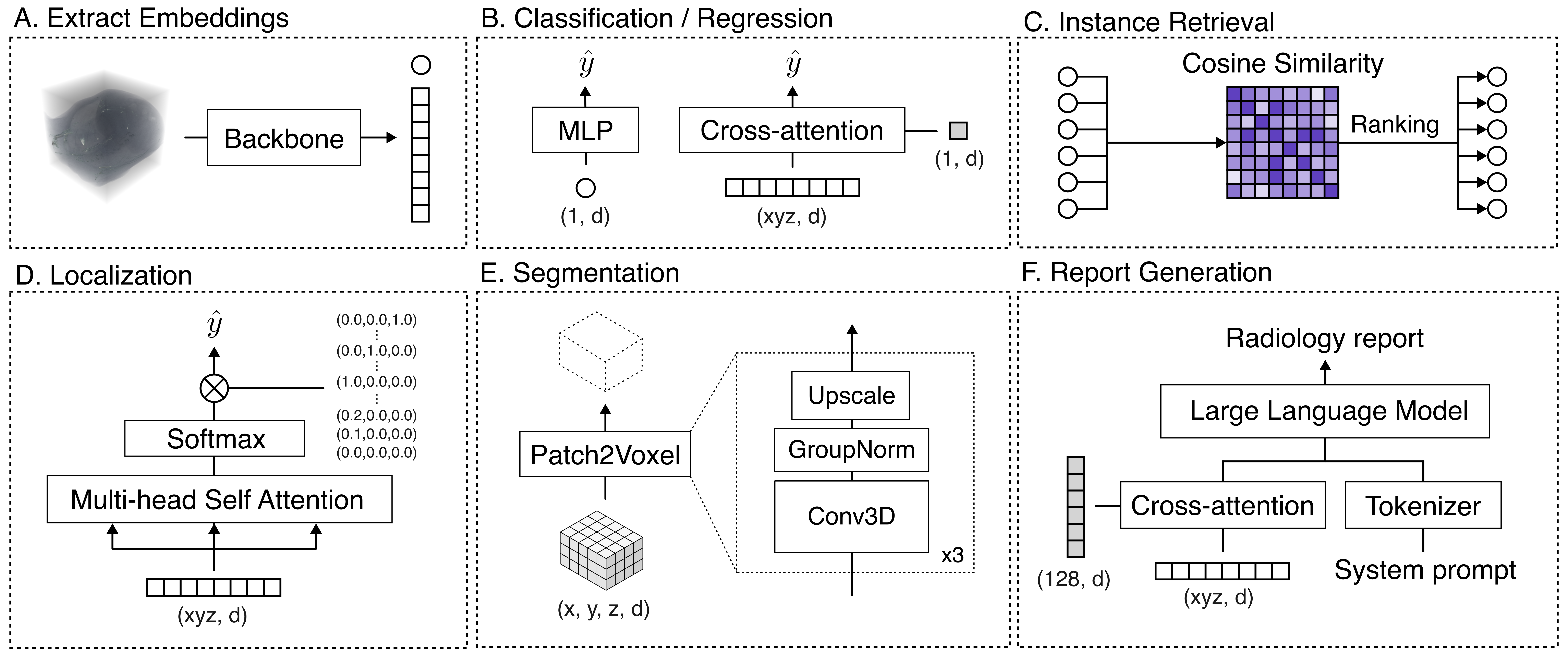
Figure 2: The evaluation protocol decouples feature extraction from task adaptation, covering classification, regression, survival, retrieval, localisation, segmentation, and report generation with specialized frozen-probe decoders.
Main Results
VoxelFM demonstrates statistically significant superior or matched performance on all downstream tasks compared to four recent CT foundation models (Merlin, RadFM, M3D, CT-CLIP), across a heterogeneous suite of clinical datasets. Notably:
- Classification: VoxelFM achieves the highest AUROC on 5/6 benchmarks; e.g., 0.870 on CT-RATE, vs. Merlin’s 0.798 (p < 10⁻¹⁷), and 0.760 on RSNA-STR pulmonary embolism (next best: 0.597).
- Regression: Outperforms competitors on OSIC forced vital capacity (MAE 0.591 vs. 0.693).
- Survival Analysis: The only model distinguishing survival above chance (C-index 0.602 vs. 0.533 for Merlin), with all other baselines performing at or below chance.
- Segmentation: Highest DICE scores and especially strong performance on macro-averaged DICE, indicating improved detection of rare/low-prevalence classes.
- Localisation: Substantially reduced MAE for lung nodule centroid prediction (0.100 vs. Merlin's 0.234).
- Report Generation: Outperforms vision-language supervision methods (macro-F1 0.432 vs. Merlin 0.327) despite no language supervision during pre-training.
- Instance Retrieval: All models perform near chance, indicating current global embeddings are suboptimal for direct clinical search.
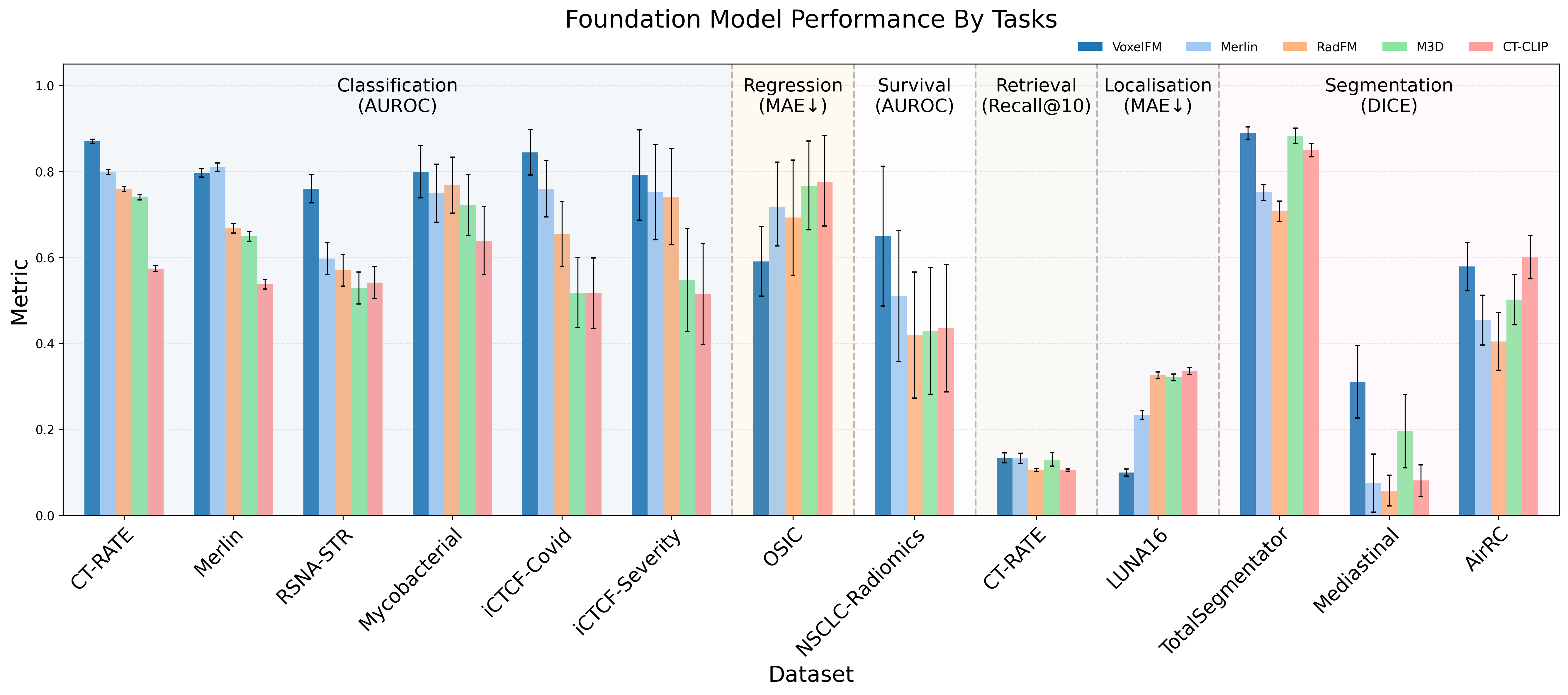
Figure 3: Bar chart summarizing model performance across multiple tasks, with VoxelFM leading in AUROC, DICE, and F1 metrics.
Ablation Studies: Token Types, Dataset Size, and Volumetric Strategies
Empirical ablation reveals that class token MLP probes excel with small training sets, while patch-token Q-Formers require larger datasets to outperform due to their higher representational capacity. VoxelFM maintains robust classification performance even with severe reduction of training labels (e.g., AUROC 0.74 with only 20% training data for iCTCF-Covid).
The model’s feature extractor is robust to full-volume inference and chunked “2.5D” strategies, confirming the capacity to generalize across clinical acquisition protocols and memory constraints.
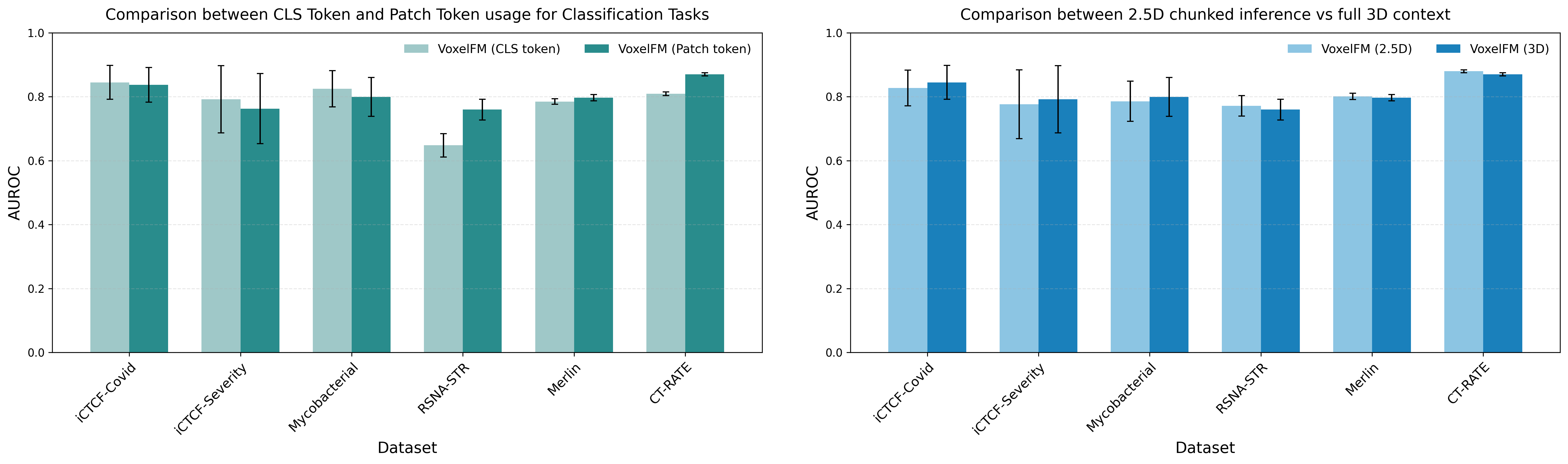
Figure 4: Comparative impact of probe type (class token MLP vs. patch token Q-Former) and processing scheme (chunked 2.5D vs. full 3D) on downstream classification accuracy.
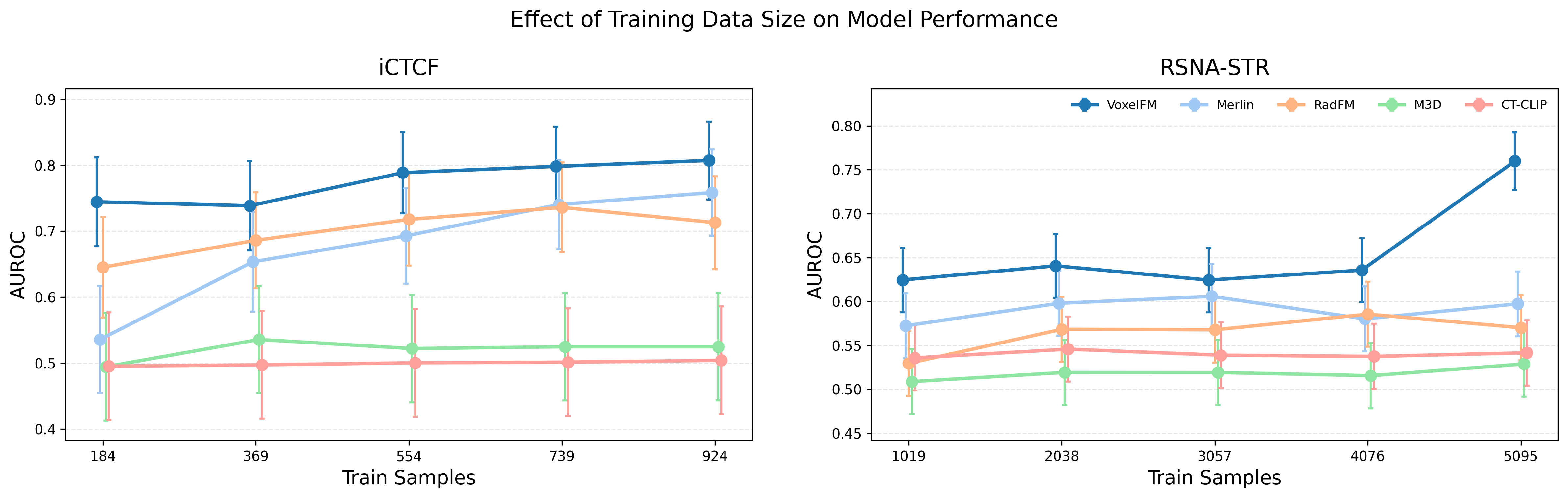
Figure 5: Downstream performance as a function of available supervised data; VoxelFM maintains high accuracy even in low-data regimes.
Detailed Downstream Analysis
Classification and Regression
Strong overall classification performance is observed across diverse anatomical sites and pathologies, with pronounced gains in challenging contexts (e.g., pulmonary embolism detection). Regression tasks benefit similarly, with quantitative biomarker predictions surpassing language-supervised alternatives.
Survival and Prognostication
Critically, only VoxelFM produces meaningful risk stratification on NSCLC-Radiomics (C-index > 0.6). Competing methods—based on language supervision or hybrid strategies—yield random assignment, highlighting a limitations of current language-aligned pipelines for complex, weakly supervised prognostic tasks.
Fine-Grained Tasks: Localisation and Segmentation
Patch-level representations acquired via DINO self-distillation enable competitive spatial encoding for both nodule localisation and multi-organ segmentation, even for rare tissues (macro-DICE). The robustness across organs illustrates the task-agnostic semantic capacity of learned features.
Report Generation and Vision-Language Transfer
Remarkably, VoxelFM outperforms all competing vision-LLMs in automated report generation metrics, even though it never saw text during pre-training. Importantly, in all cases, classification probes trained on frozen visual features decisively outperform report generation pipelines for structured label extraction from CT, reinforcing the view that current CT vision-language alignments are inferior to specialized discriminative adaptation.
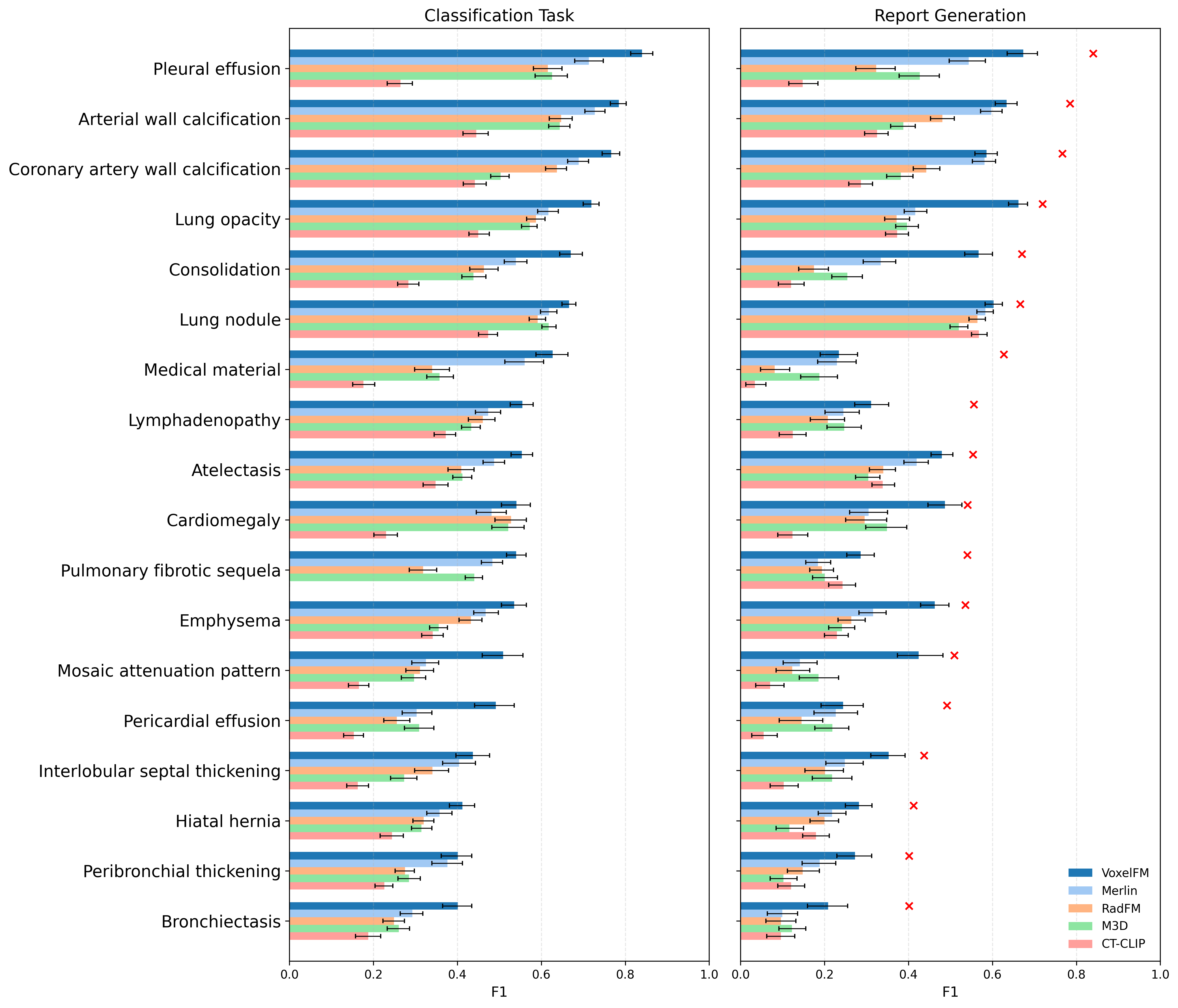
Figure 6: Per-abnormality comparison of simple classifier and report generation F1 scores, with binary classifiers consistently outperforming generative models for all 18 labels.
Discussion and Implications
The results strongly support the thesis that self-supervised pre-training with DINO yields visual features in the CT domain that are semantically rich, transfer-efficient, and outperform existing language-alignment frameworks across structurally diverse clinical tasks. There is clear evidence that feature-based transfer—using frozen backbones and lightweight probes—is substantially more practical and effective than end-to-end re-training or reliance on vision-language transfer schemes, given current data regimes and compute accessibility.
No advantage is observed for instance retrieval or open-ended report generation tasks with existing vision-language foundation models, due in part to the limited scale and structural mismatch of available paired data. Instead, domain-general visual features learned via self-distillation appear to be the primary performance bottleneck, as supported by the systematic gains demonstrated by VoxelFM.
From a practical perspective, VoxelFM’s robustness to label scarcity and data variability marks a significant step toward democratization of foundation model utility in medical imaging, where resource constraints and annotation bottlenecks are standard.
Limitations and Future Directions
Limitations include the absence of full backbone tuning comparisons for all baselines (which may slightly underestimate the ceiling of language-supervised models with abundant labels) and incomplete pre-training data distribution auditing, potentially masking distribution shifts in extreme-case deployment.
Potential future directions comprise:
- Hybrid self-distillation and language-supervised pre-training at increased scale (e.g., integrating the gram loss from DINOv3 or language fusion objectives).
- Expansion towards multi-modal or longitudinal CT data.
- Revisiting vision-language strategies as paired data scales to 10⁵–10⁶ annotated samples.
Conclusion
VoxelFM conclusively demonstrates that current CT foundation models deliver stronger, more robust transfer using self-supervised visual features with frozen backbones, compared to vision-language approaches at prevailing data scales. The present data regime in CT does not support generalist report generation: task-specific discriminative probes are far more reliable and efficient. The practical implication is a shift in paradigm—CT foundation models should focus on scalable, task-agnostic feature learning, with language supervision as an adjunct only when data and compute scale accordingly.

Figure 1: The VoxelFM pre-training framework combining multi-scale cropping, self-distillation, and robust transformer-based feature extraction for universal clinical task transfer without backbone adaptation.
In summary, VoxelFM articulates and validates a foundation modeling blueprint for CT: prioritize robust, scalable visual feature learning under self-supervision, and leverage this for efficient downstream clinical adaptation. The model and code are publicly available, establishing a reproducible foundation for further CT modeling research and clinical translation (2604.04133).







