- The paper introduces MonoMSK, a framework combining inverse dynamics with a differentiable forward dynamics ODE solver to achieve realistic 3D musculoskeletal motion estimation.
- It integrates transformer-based modules with anatomically accurate musculoskeletal models, yielding significant error reductions on benchmarks like BML-MoVi and BEDLAM.
- The study demonstrates practical applications in clinical analysis, sports, and robotics by ensuring physical plausibility and biomechanical consistency in motion estimation.
Physically Grounded 3D Human Motion Dynamics from Monocular Video: A Technical Analysis of MonoMSK
Introduction
"MonoMSK: Monocular 3D Musculoskeletal Dynamics Estimation" (2511.19326) establishes an integrated framework for estimating both kinematics and kinetics of full-body human motion from monocular video input. Leveraging a hybrid of transformer-based neural modules and differentiable physics simulation, MonoMSK aims to address inherent limitations in current monocular pose estimation methods, specifically the inability to produce biomechanically realistic and physically plausible motion reconstructions.
Prevailing monocular frameworks typically employ simplified skeletal models (e.g., SMPL) and disregard the causality underpinning biomechanical dynamics. The novelty of MonoMSK is its fusion of data-driven inverse kinematics/dynamics prediction with a Forward Dynamics ODE solver over an anatomically precise musculoskeletal model, producing interpretable motion and physically plausible force/torque estimates.
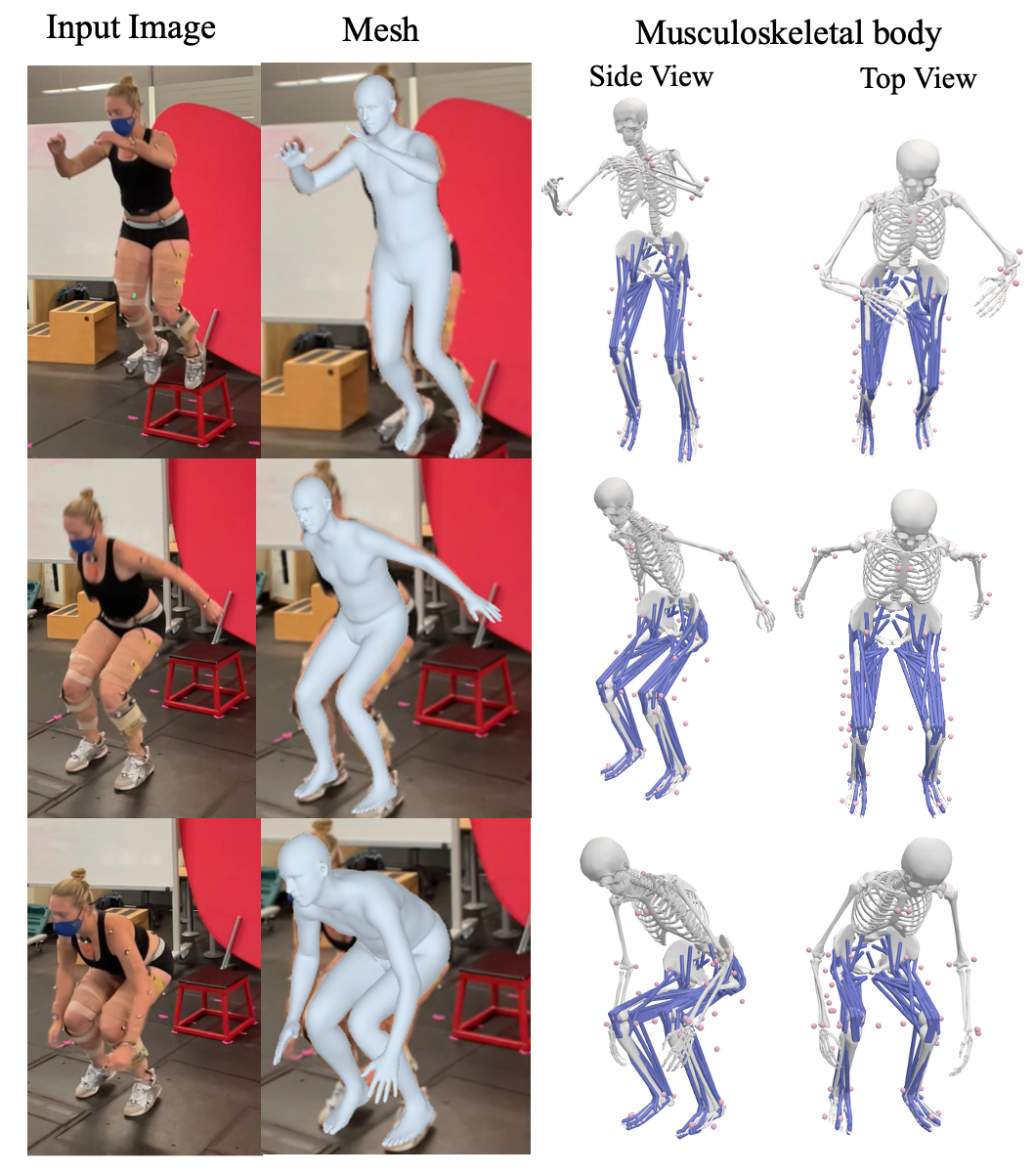
Figure 1: MonoMSK couples transformer-based inverse dynamics with a differentiable Forward Dynamics ODE solver, yielding biomechanically consistent motion reconstructions from monocular videos.
Methodological Framework
Pipeline Overview
MonoMSK incorporates five sequential stages:
- Human Mesh Recovery (HMR): Pretrained models generate 3D meshes and virtual markers.
- Inverse Kinematics Transformer (IKT): Converts marker positions into anatomical musculoskeletal joint states q.
- Inverse Dynamics Transformer (IDT): Infers latent dynamic quantities—internal joint torques τ and external ground reaction forces λ.
- Forward Kinematics (FK) Layer: Converts simulated joint states back to marker positions for evaluation.
- Forward Dynamics (FD) ODE Solver: Simulates continuous motion evolution under predicted forces and torques for physical verification.
The ODE-based FD solver integrates the physics of the musculoskeletal model, enforcing plausibility and ensuring closed-loop consistency between inferred kinetics and kinematics.
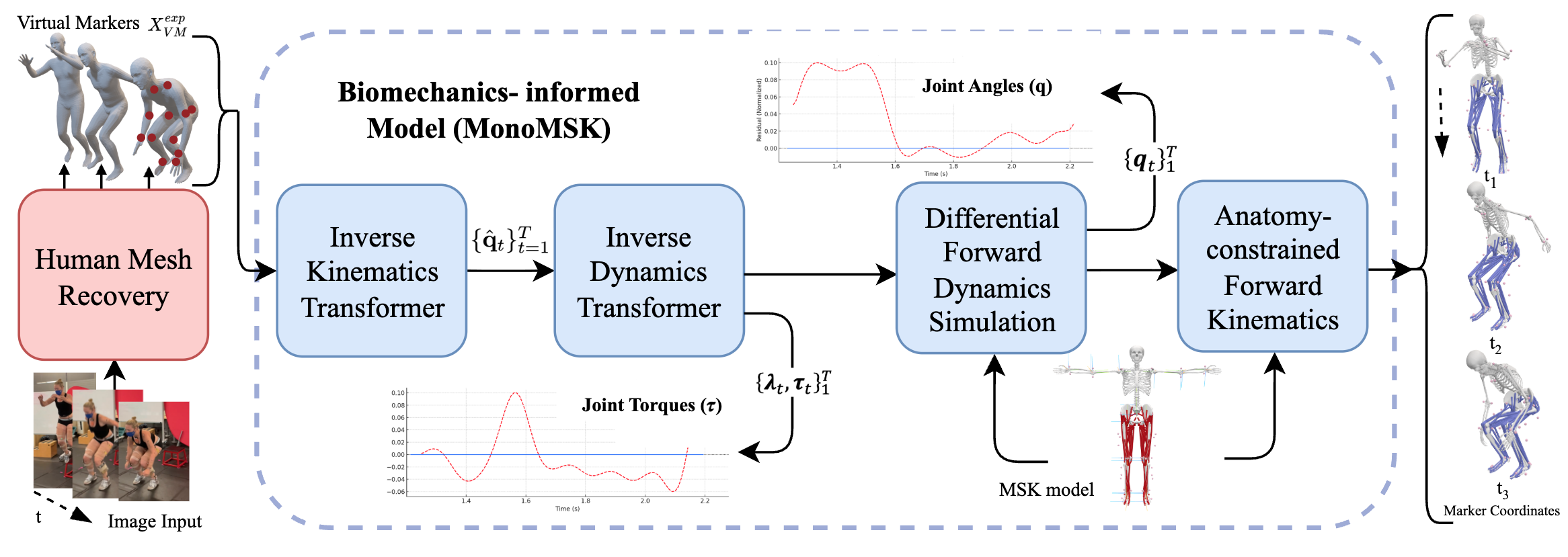
Figure 2: Schematic of MonoMSK pipeline with integration of HMR, inverse transformers, ODE dynamics, and anatomical FK.
Anatomically Accurate Musculoskeletal Modeling
The core of MonoMSK is its detailed musculoskeletal (MSK) model, reflecting true joint locations, muscle insertions, segment masses, and physiological DoFs. The MSK model enables subject-specific anatomical adaptation and explicit modeling of muscle dynamics, joint torques, and ground interactions.
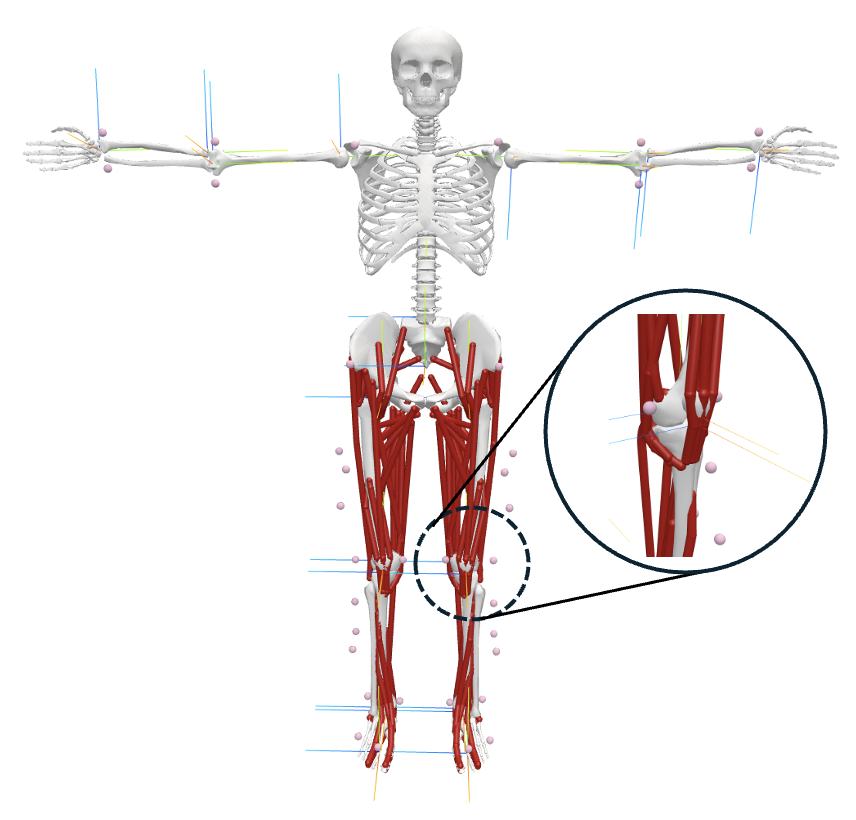
Figure 3: The MSK body model, capturing precise musculature, joint geometry, and virtual markers for biomechanical tracking.
Forward simulation in MonoMSK leverages Newton-Euler ODEs and Hunt–Crossley contact models, capturing nuanced force transmission, ground reaction dynamics, and joint torques based on muscle activation profiles. This enables robust simulation of motion phases such as heel-strike and toe-off, which are poorly represented in traditional parametric models.
Training Strategy
Ground-truth for supervised learning is sourced from physics-based optimal control simulations (OpenSim-Moco), yielding precise kinematics and kinetics reference trajectories. Training losses consist of:
- Kinetics Losses: Mean squared error on predicted versus reference joint torques (Lτ) and ground reaction forces (Lλ).
- Consistency Losses: Inverse-forward loop regularization enforcing agreement between ODE-integrated predictions and original kinematic observations, through joint rotation (Lq) and anatomical marker position (LJ) errors.
This combination ensures the formation of an interpretable latent space aligned with physical causality and anatomical feasibility.
Experimental Results
Quantitative Evaluation
MonoMSK is benchmarked against leading monocular and biomechanics-based approaches on BML-MoVi, BEDLAM, and OpenCap datasets. Key metrics include mean per-bony-landmark position error (MPBLPE), joint-angle MAE, acceleration and velocity errors, and direct MAE of predicted forces and torques.
MonoMSK achieves substantial improvements:
- BML-MoVi: Joint-angle MAE reduced by 32.0% and MPBLPE by 5.4% over the BioPose baseline; acceleration/velocity errors reduced by 30–39%.
- BEDLAM: 18.1% reduction in MAEangle, 32.6% in acceleration, and 23.8% in velocity error.
- OpenCap: 11% joint-angle, 19.9% acceleration, and 22.2% velocity reductions.
MonoMSK delivers the first precise monocular kinetics estimates, outperforming direct force/torque optimization approaches.
Ablation Studies
- HMR Backbone Quality: Upstream mesh recovery fidelity (e.g., MQ-HMR vs. CameraHMR) directly impacts force/torque estimation accuracy and plausible motion trajectories.
- Training Objective Configuration: Joint optimization of external and internal force losses, plus consistency regularization, is critical for SOTA biomechanical and kinetic fidelity.
- Temporal Prediction Strategy: Single Frame Out autoregressive setups outperform multi-frame forecasts, yielding lower drift and superior stepwise physical integration.
Theoretical and Practical Implications
MonoMSK's integration of differentiable physics into transformer motion models advances biomechanically faithful human motion understanding. The framework supports fine-grained force/torque analysis from camera videos, enabling noninvasive studies in clinical motion analysis, sports performance, robotics, rehabilitation, and digital human twins. The demonstrated generalization across synthetic and real-world datasets and robust cross-backbone performance indicate practical portability.
From a theoretical standpoint, MonoMSK highlights the importance of embedding physical priors and causality constraints in deep neural architectures for interpretable and physically realistic pose estimation. The physics-regulated bidirectional loop can inform broader research in physically consistent generative modeling, simulation-based inference, and learning dynamics from sparse observations.
Future Directions
Potential avenues include extending MonoMSK to multi-person tracking, incorporating richer muscle activity estimation, real-time deployment for wearable-free motion analysis, and adaptively learning subject-specific biomechanical parameters. Further, the general forward-inverse consistency paradigm may be applied to other domains demanding interpretability and causal reasoning in physical simulation.
Conclusion
MonoMSK offers a unified system for biomechanically accurate 3D human motion estimation, bridging the gap between visual perception and physical reasoning from monocular video. By combining deep learning with differentiable physics over anatomical skeletons, MonoMSK advances the field of human motion dynamics, enabling noninvasive, interpretable reconstruction of both kinematics and kinetics, and establishes new standards for physical plausibility and biomechanical analysis in computer vision.


