Genomic Next-Token Predictors are In-Context Learners
Abstract: In-context learning (ICL) -- the capacity of a model to infer and apply abstract patterns from examples provided within its input -- has been extensively studied in LLMs trained for next-token prediction on human text. In fact, prior work often attributes this emergent behavior to distinctive statistical properties in human language. This raises a fundamental question: can ICL arise organically in other sequence domains purely through large-scale predictive training? To explore this, we turn to genomic sequences, an alternative symbolic domain rich in statistical structure. Specifically, we study the Evo2 genomic model, trained predominantly on next-nucleotide (A/T/C/G) prediction, at a scale comparable to mid-sized LLMs. We develop a controlled experimental framework comprising symbolic reasoning tasks instantiated in both linguistic and genomic forms, enabling direct comparison of ICL across genomic and linguistic models. Our results show that genomic models, like their linguistic counterparts, exhibit log-linear gains in pattern induction as the number of in-context demonstrations increases. To the best of our knowledge, this is the first evidence of organically emergent ICL in genomic sequences, supporting the hypothesis that ICL arises as a consequence of large-scale predictive modeling over rich data. These findings extend emergent meta-learning beyond language, pointing toward a unified, modality-agnostic view of in-context learning.
Paper Prompts
Sign up for free to create and run prompts on this paper using GPT-5.
Top Community Prompts
Explain it Like I'm 14
Overview
This paper asks a big question: Can computers learn patterns “on the fly” from examples, not just in human language, but also in DNA? The authors show that a DNA-focused model, trained to predict the next nucleotide (A, T, C, or G), can learn rules from examples given in its input—just like LLMs do with words. This ability is called in-context learning.
What the researchers wanted to find out
Put simply, they explored:
- Does in-context learning only happen because human language has special properties?
- Or can it also appear in other kinds of sequences—like DNA—if models are large and trained to predict the next symbol?
They tested this by comparing:
- A LLM (Qwen3), trained on text.
- A genomic model (Evo2), trained on DNA sequences.
How they tested it (in everyday terms)
Think of the models as puzzle-solvers. They are shown a few examples of input and output pairs that follow some hidden rule. Then they must figure out the rule and apply it to a new example. No model parameters are changed; the model just uses the examples in the prompt to “learn” the rule on the spot.
Here’s how they set up the puzzles:
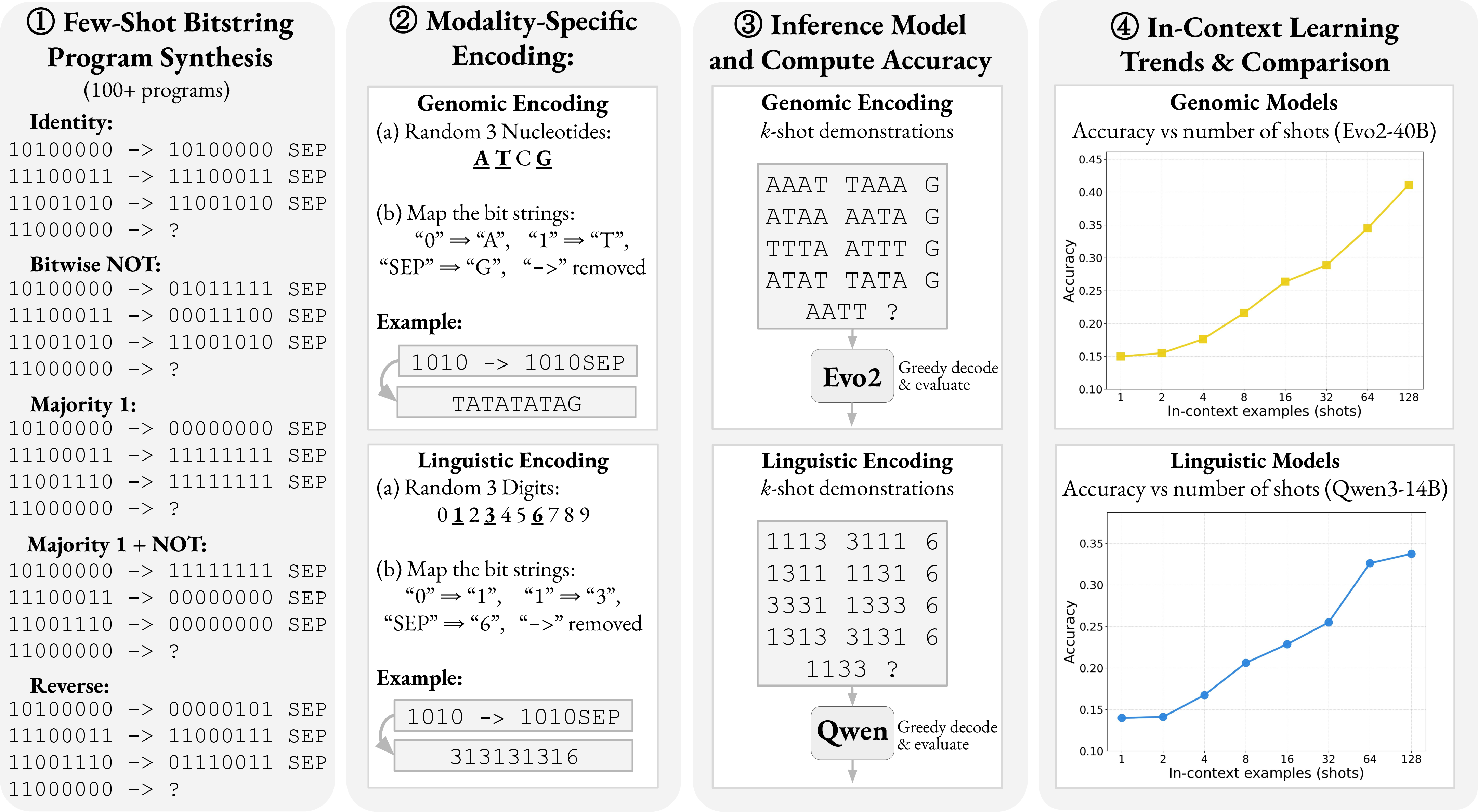
- They used 8-bit binary strings (like 01010101). Binary is simple and fair for both models.
- They made many kinds of rules that transform one bitstring into another (for example: copy it exactly, flip all bits, shift bits left or right, count how many ones, reverse the order).
- To compare across domains, they presented the same puzzles in two forms:
- Linguistic form: bits shown as digits (0 and 1).
- Genomic form: bits shown as DNA letters (A/T/C/G), using two letters to mean “0” and “1,” and others as separators.
- They gave both models different numbers of example pairs in the prompt (from 1 up to 128) and measured how often the answer was exactly correct.
A small sample of puzzle types they used, just to give you a feel:
- Identity: “Just copy the input.”
- NOT: “Flip each bit (0→1, 1→0).”
- Majority: “If more bits are 1 than 0, output all 1s; otherwise all 0s.”
- Reverse: “Reverse the order of the bits.”
- Shift: “Move bits over by one position.”
They also compared against a simple “mode baseline,” which just guesses the most common output seen in the examples, without trying to understand the input-output rule.
Main findings and why they matter
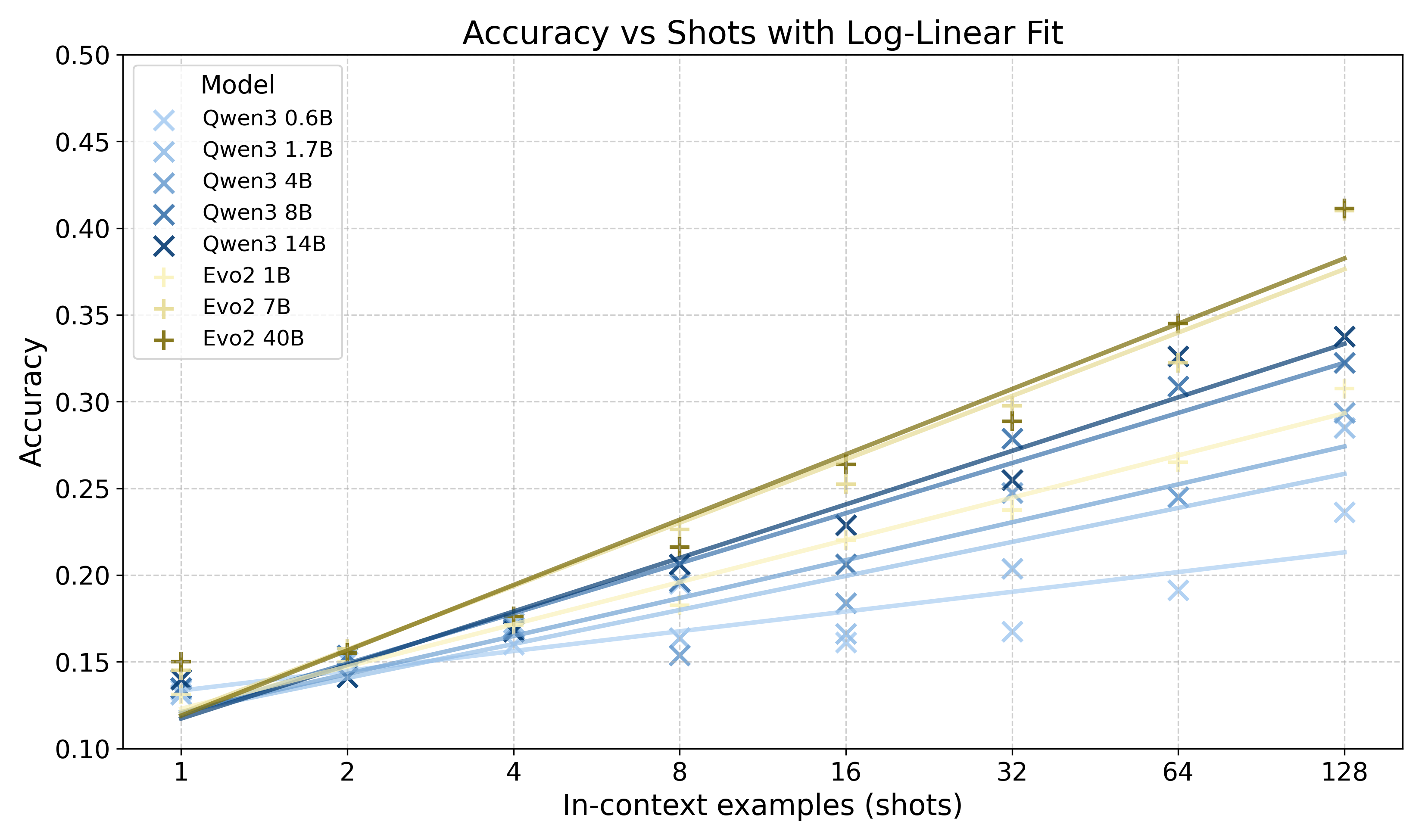
- Both models learn better with more examples. Each time the number of examples increased (like doubling from 8 to 16), accuracy rose in a steady, predictable way. In plain terms: more demonstrations → better rule learning.
- The DNA model (Evo2) showed clear in-context learning. This is strong evidence that in-context learning isn’t only a language thing; it can appear in other sequence types too, if the model is large and trained to predict the next token.
- Compared at similar sizes, the genomic model often performed better than the LLM on these bitstring tasks, especially as the number of examples grew.
- Both models beat the “mode baseline,” meaning they weren’t just guessing the most common output. They actually learned the rule connecting inputs to outputs from the examples.
- Different strengths:
- Evo2 was better at “whole-string” operations, like flipping every bit or copying exactly, and stayed more robust on harder rules.
- The LLM often shined on tasks like simple shifts and counting-based rules (like parity or majority), which rely on global properties of the string.
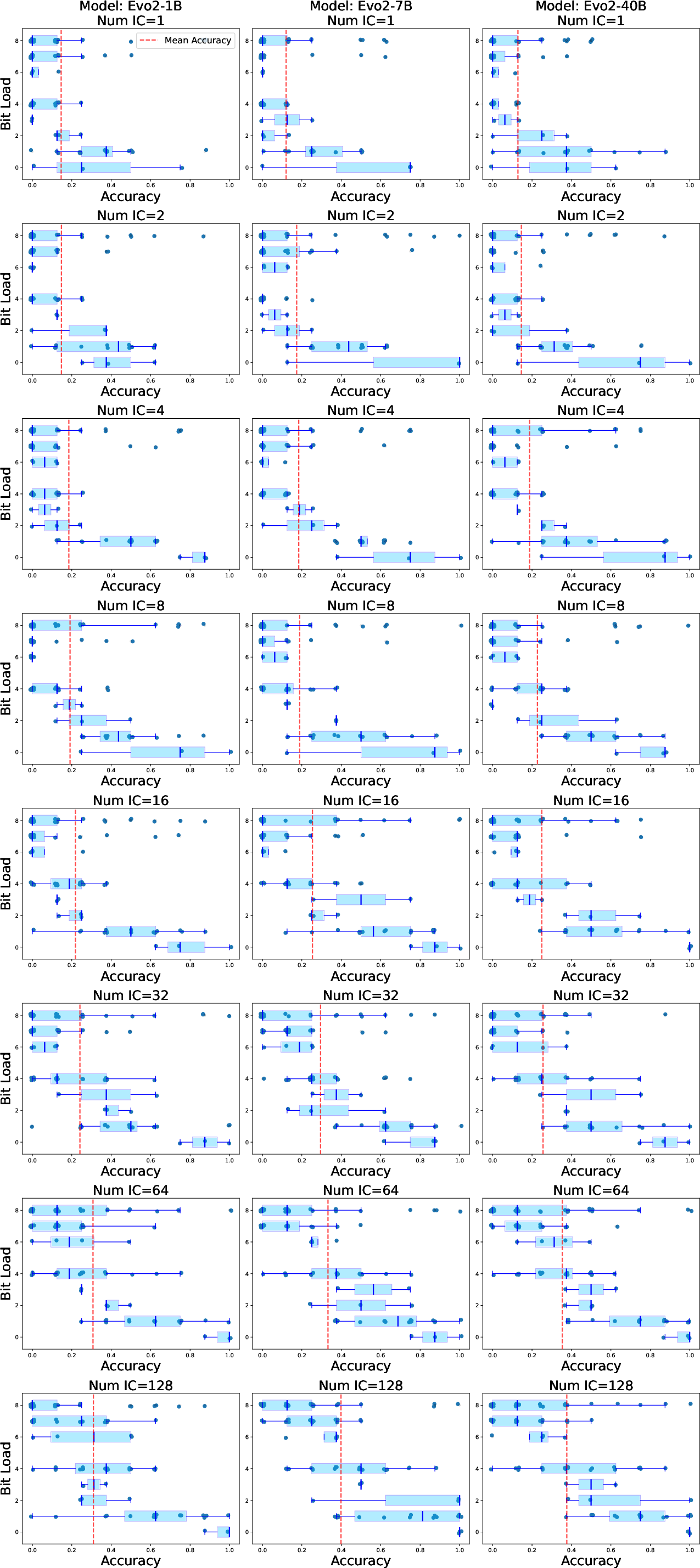
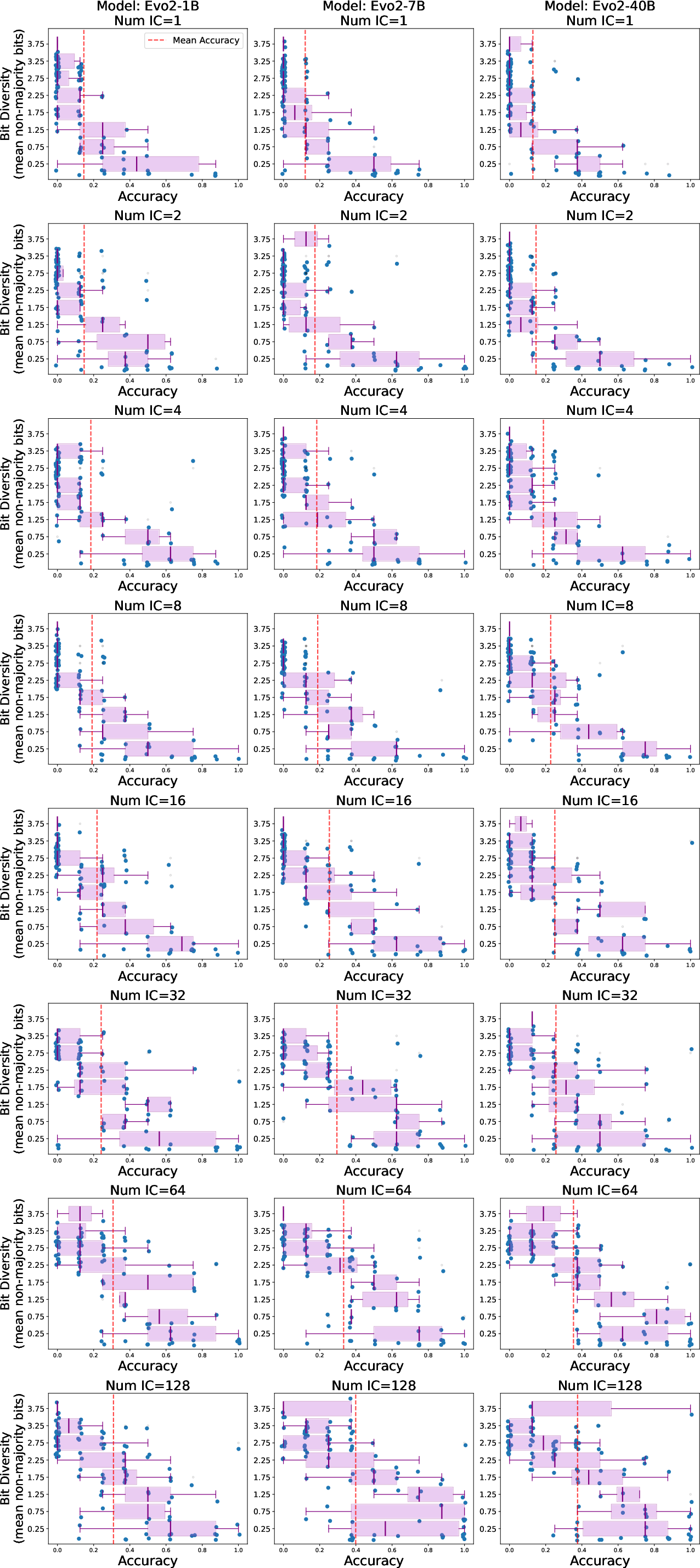
- Task complexity matters. The authors introduced “BitLoad,” a measure of how many input bits affect the output. Higher BitLoad means a harder rule. Evo2’s accuracy dropped more gently as BitLoad increased; the LLM’s accuracy fell more sharply on complex rules. This suggests Evo2 generalizes better as rules require paying attention to more bits.
Why this is important: It’s the first strong evidence that in-context learning can show up in genomic sequences naturally, just from large-scale next-token training—no special extra teaching needed.
What this means going forward
- In-context learning seems to be a general effect of training big models to predict the next symbol on rich, structured data. It’s not something magical about human language alone.
- This points to a “modality-agnostic” view: if you have sequences with patterns (words, DNA, maybe even other signals), and you train a big enough model to predict the next item, it can learn to use examples in its input to solve new tasks on the fly.
- For biology, this could lead to smarter tools that quickly spot patterns in DNA or learn new transformation rules from a few examples, helping with analysis of genetic sequences, designing molecules, or understanding the effects of mutations.
- For AI research, it supports the idea that compression and prediction over large datasets can naturally create flexible, “learn-from-context” behavior, across different kinds of data and architectures.
Knowledge Gaps
Knowledge gaps, limitations, and open questions
Below is a concise, actionable list of what remains missing, uncertain, or unexplored in the paper’s evidence and methodology.
- Causal attribution remains unresolved: the paper infers that ICL is modality-agnostic and driven by large-scale predictive compression, but offers no causal tests disentangling data distribution effects, model architecture, and training objective; controlled ablations (matched architectures, matched datasets, shuffled/perturbed corpora) are needed.
- Architectural confounds are unaddressed: Evo2 is a hybrid AR model with very long context, while Qwen3 is a transformer trained on text; no experiments hold architecture constant across modalities or vary architecture within a modality to pinpoint which components are necessary/sufficient for ICL.
- Compute and training differences are only approximately matched: there is no control for training data size, diversity, curriculum, optimizer, or training regime; stronger matching or controlled from-scratch training would reduce attribution ambiguity.
- Context-length differences could bias results: Evo2-7B/40B have 1M-token contexts whereas Qwen3 has much shorter contexts; the paper does not test whether long-context pretraining per se enhances few-shot ICL on short prompts.
- Prompt-format sensitivity is not systematically explored: the authors acknowledge Qwen3 can solve identity under familiar formatting (arrows/newlines), yet the study fixes an intentionally unfamiliar, compressed format; systematic prompt ablations (delimiters, whitespace, line breaks, markers, ordering) are missing.
- Tokenization and surface-form confounds persist: digits for language vs nucleotides for genomics may confer unequal familiarity and tokenization benefits; cross-encoding controls (e.g., Qwen3 on A/C/G/T encodings; letter-like symbols) and tokenizer-aware analyses are absent.
- No cross-domain, cross-encoding stress tests: LLMs are not evaluated on nucleotide encodings and genomic models cannot process digit encodings; this limits the claim that ICL is surface-agnostic across modalities.
- Limited task scope risks overgeneralization: tasks use 8-bit strings and at most depth-2 compositions; it is unknown whether findings hold for longer bitstrings (e.g., 16/32/64 bits), deeper compositions, or richer function classes (arithmetic, automata, regular/context-free grammars).
- Ambiguity/identifiability of functions is not controlled: random demonstration sets can be insufficient to uniquely identify the latent function; best-case/worst-case/active selection of examples and identifiability analyses are missing.
- Few-shot sampling is statistically underpowered: only m=8 Monte Carlo trials per function are used; accuracy estimates and p-values may be fragile; power analyses and larger m are needed.
- Metric choice may hide partial competence: exact-match is the sole metric; no Hamming distance, per-bit accuracy, or calibrated log-probability analyses to capture near-misses or graded learning.
- Baseline comparisons are weak: only a “mode” baseline is used; no comparisons to simple algorithmic few-shot solvers (nearest neighbor, decision trees on bit features, linear in-context solvers, ridge-regression/attention-as-inference baselines) or to meta-learners.
- No zero-shot or few-shot-to-zero-shot extrapolation tests: whether models can generalize without demonstrations, or interpolate/extrapolate with fewer/more demonstrations than trained, remains untested.
- Mechanistic explanations are lacking: there is no interpretability analysis (e.g., induction heads, attention patterns, probing, activation patching, KV-cache dynamics) to reveal how Evo2 or Qwen3 implement ICL on these tasks.
- The BitLoad complexity measure is limited: it counts bit influence but ignores compositional depth, nonlinearity, invariances, and algorithmic complexity; alternative or complementary complexity measures and controlled task sets are needed.
- Plateau in Evo2 (7B ≈ 40B) is unexplained: the paper does not determine whether saturation arises from optimization limits, data bottlenecks, architecture, or evaluation ceiling effects.
- No robustness tests to noise or adversarial prompts: the impact of label noise, corrupted demonstrations, adversarial ordering, or out-of-support queries is unmeasured.
- No analysis of demonstration order and recency effects: whether example ordering (curriculum), clustering, or recency in the prompt affects ICL accuracy is unknown.
- No examination of calibration or uncertainty: confidence estimates, selective prediction, and abstention policies are not studied; models might be overconfident on wrong functions.
- Temperature/decoding effects are not tested: only greedy decoding is used; it is unknown whether sampling, nucleus/beam search, or reranking changes ICL success.
- Potential contamination is not ruled out: Qwen3 likely saw bitstring/parity-like tasks in pretraining; systematic OOD controls (synthetic symbols, held-out function families, spurious token mappings) are needed to discount memorization.
- Pretraining data properties are untreated experimentally: hypotheses about “compression over rich data” are not tested via corpus ablations (e.g., shuffled genomes preserving unigram/bigram stats, motif-scrambled controls) to separate structure vs frequency effects.
- No comparison of objectives: only autoregressive training is considered; whether masked-LM, permutation-LM, or next-k-token prediction produce similar ICL on these tasks is unknown.
- No study of instruction tuning or RLHF effects: results are for base models only; do post-training methods enhance or hinder ICL on symbolic induction in each modality?
- Generalization to ecologically valid genomic tasks is untested: the paper does not evaluate in-context motif discovery, splice site induction, variant-effect pattern induction, or other real genomic reasoning tasks using demonstrations.
- Cross-modality breadth is narrow: claims of modality-agnostic ICL would be stronger with additional non-linguistic domains (protein amino-acid sequences, music, clickstreams, symbolic math, vision sequences) under the same framework.
- Sample complexity per function is uncharacterized: the minimal number of demonstrations required to reliably identify each transformation and how this varies by BitLoad or composition depth is not reported.
- No learning dynamics analysis within prompts: whether intermediate examples are “used” (e.g., via causal tracing) and whether models update internal hypotheses across the demonstration list is unclear.
- Effects of GC-content and nucleotide frequency biases are untested: the random mapping of bits to nucleotides may interact with Evo2’s learned nucleotide priors; experiments controlling GC-skew and symbol frequency are needed.
- Reproducibility details are thin for statistical claims: seeds, hardware variability, and sensitivity to random remappings are not fully reported; rigorous reproducibility artifacts (configs, seeds, full result tables) would strengthen claims.
- Utility for practitioners is not demonstrated: guidelines for using Evo2’s ICL in real genomic workflows (e.g., how many examples to include, formatting recipes, robustness expectations) are missing.
- Broader theoretical claims are not formalized: while the discussion favors compression-based explanations, no predictive theory or scaling law is proposed to bound ICL emergence across modalities and tasks.
Glossary
- 6ND estimate: A rule-of-thumb for estimating pretraining compute where total FLOPs scale as approximately 6 × tokens × parameters. "Using the standard $6ND$ estimate"
- ARC-AGI: A benchmark of abstract reasoning/program-induction tasks used to probe generalization and compositionality. "and more recently in ARC-AGI"
- analogical reasoning: Solving new problems by mapping relational structure from examples to queries. "few-shot generalization and analogical reasoning"
- autoregressive model: A sequence model that predicts each token conditioned on all previous tokens. "A pretrained autoregressive model performs ICL"
- BitLoad: A complexity measure counting how many input bits influence a function’s output; higher BitLoad implies harder inference. "We introduce BitLoad"
- burstiness: A distributional property where events occur in clusters rather than uniformly over time. "``burstiness''"
- compositionality: The property that complex structures or meanings are built by combining simpler parts according to rules. "such as parallelism or compositionality"
- context length: The maximum number of tokens a model can condition on in a single input sequence. "trained at a context length of 8192 nucleotides"
- exact-match accuracy: An evaluation metric that counts a prediction correct only if it exactly equals the target. "are greedily decoded to compute exact-match accuracy."
- few-shot: Performing a task or inferring a rule from only a handful of in-context examples. "few-shot generalization"
- FLOPs: Floating-point operations; a standard unit for measuring computational training or inference cost. "about FLOPs"
- greedy decoding: A decoding strategy that selects the highest-probability token at each step without lookahead search. "are greedily decoded to compute exact-match accuracy."
- in-context learning (ICL): The ability of a model to infer and apply a task from examples in its input without updating parameters. "In-context learning (ICL)"
- instruction fine-tuning: Supervised tuning on instruction–response pairs to make a base model follow instructions. "soft prompting and instruction fine-tuning"
- intersection-over-union: A set-similarity metric defined as |A ∩ B| / |A ∪ B|, common in comparative evaluations. "yielding an intersection-over-union of 0.54."
- log-linear: Describing a relationship that is linear in the logarithm of a variable (e.g., performance vs. log shots). "exhibit log-linear gains in pattern induction"
- meta-learning: Learning to learn across tasks so a system can rapidly adapt to new tasks from few examples. "emergent forms of meta-learning"
- Meta-ICL: In-context learning behavior trained explicitly with meta-learning objectives over input–output pairs. "We refer to such systems as Meta-ICL"
- mode baseline: A simple baseline that predicts the most frequent output observed in the context examples. "We define a mode baseline"
- Monte Carlo estimate: An estimate computed via random sampling to approximate expectations or performance metrics. "we use a Monte Carlo estimate"
- motifs: Short, recurring patterns in genomic sequences that often have functional roles. "motifs, repeats, dependencies"
- next-nucleotide prediction: An autoregressive objective on DNA sequences to predict the next base (A/T/C/G). "trained solely on `next-nucleotide' prediction."
- parallelism: Recurrent structural regularities in data that can facilitate pattern induction. "such as parallelism or compositionality"
- perplexity: The exponentiated average negative log-likelihood; lower values indicate better next-token prediction. "increase perplexity far more than other common mutations"
- program synthesis: Inferring a transformation or program from example input–output pairs. "program-synthesis tasks"
- quaternary encoding: Representation using four symbols, such as mapping to DNA bases A, T, C, and G. "via quaternary encoding"
- Ravenâs Progressive Matrices: A nonverbal analogical reasoning test involving pattern completion in matrices. "Ravenâs Progressive Matrices"
- soft prompting: Conditioning a model with learned continuous prompt vectors instead of discrete text tokens. "soft prompting and instruction fine-tuning"
- variant effect estimation: Predicting the functional impact of genetic mutations on biological sequences or traits. "variant effect estimation"
- z-test: A statistical hypothesis test using a normal approximation to assess whether an effect is significant. "one-sided z-test on bootstrapped standard errors"
Practical Applications
Immediate Applications
The following applications can be deployed now, leveraging the paper’s methods (cross-domain ICL benchmarking with bitstring program induction), findings (emergent ICL in genomic next-token models), and tools (released code and evaluation protocol).
- Cross-domain ICL evaluation toolkit for non-linguistic models
- Sector: academia, software
- What: Use the released code to benchmark emergent ICL in genomic models (Evo2) and compare with LLMs using parallel symbolic tasks and the BitLoad metric.
- Products/workflows:
- A standardized “ICL audit” harness for sequence models (genomics, proteomics, time series).
- Model report cards that include accuracy-vs-shots curves, mode-baseline comparisons, and BitLoad sensitivity.
- Assumptions/dependencies: Access to base (non-instruction-tuned) models and inference compute; ability to represent domain tasks as symbolic transformations or simple sequence mappings.
- Prompt-based genomic pattern induction for exploratory bioinformatics
- Sector: biotech, healthcare (R&D and translational)
- What: Use few-shot prompts (examples of input-output sequence pairs) to induce local transformations or pattern recognition without fine-tuning (e.g., recognizing CRISPR PAMs, simple motif presence/absence, reverse-complement checks, local sequence rearrangements).
- Products/workflows:
- Jupyter/CLI utilities that accept a small set of labeled sequences and return predictions on new sequences.
- Integration into genomic browsers or QC dashboards to prototype rules from a few examples.
- Assumptions/dependencies: Tasks should have clear local dependencies and structured regularities (Evo2 excels on copying/full-bitstring manipulations); careful prompt formatting and adequate shot counts (log-linear gains with shots).
- Sequence quality control and anomaly detection with few-shot prompts
- Sector: sequencing pipelines, bioinformatics tooling
- What: Provide a few examples of artifacts (adapter contamination, low-complexity repeats, homopolymer stretches) and query new reads for similar anomalies.
- Products/workflows:
- A “few-shot QC” stage in pipelines that flags candidate reads for review.
- Lightweight triage tools for lab technicians to refine rules during assay development.
- Assumptions/dependencies: Reliability depends on how well anomalies map to structured sequence features; requires validation against conventional QC metrics.
- Educational modules on emergent ICL beyond language
- Sector: education (computational biology, machine learning)
- What: Use the paper’s symbolic tasks and code to demonstrate how next-token models learn rules in-context across modalities.
- Products/workflows:
- Interactive notebooks that visualize accuracy gains vs. shot count and BitLoad.
- Classroom exercises comparing genomic and linguistic ICL behaviors.
- Assumptions/dependencies: None beyond standard pedagogical setup; tasks are synthetic and safe.
- Model auditing and capability assessment for biofoundation models
- Sector: policy, AI governance, industry labs
- What: Adopt the mode baseline and BitLoad analysis to assess whether a model’s “pattern induction” exceeds naive pattern matching, and how performance scales with shots/size.
- Products/workflows:
- Capability audits included in release notes for biofoundation models.
- Internal red-teaming protocols to characterize emergent behaviors prior to deployment.
- Assumptions/dependencies: Stakeholder willingness to include non-linguistic ICL metrics in capability reporting; transparent model licensing (e.g., Apache 2.0).
- Practical prompt engineering guidelines for genomic tasks
- Sector: software, biotech R&D
- What: Apply the observed log-linear scaling with shots and model-specific strengths (Evo2: robust to higher BitLoad; strong at full-bitstring transformations) to select demonstration counts and task formulations.
- Products/workflows:
- “Shots vs. difficulty” cheat sheets and prompt templates for sequence tasks.
- Heuristics to favor demonstration-heavy prompting for genomic tasks (given stronger gains from additional examples).
- Assumptions/dependencies: The synthetic scaling trends carry over to real sequence tasks with similar dependency structure; requires iteration to tune prompts.
Long-Term Applications
These applications require further research, domain alignment, validation, scaling, or productization before deployment.
- Patient-specific clinical genomics decision support via in-context prompts
- Sector: healthcare
- What: Adapt variant interpretation to specific cohorts/diseases by conditioning on a few known pathogenic/benign exemplars and associated sequence contexts.
- Products/workflows:
- Decision support modules that refine predictions for rare variants using cohort-specific few-shot examples.
- Clinician-facing tools to prototype sequence rules and generate candidate explanations.
- Assumptions/dependencies: Rigorous clinical validation, regulatory clearance, bias and privacy safeguards, robust mapping from synthetic ICL tasks to medically meaningful sequence features.
- Adaptive sequence design assistants for synthetic biology
- Sector: biotech, therapeutics, bioengineering
- What: Generate or edit DNA/RNA/protein sequences to match desired properties using few-shot demonstrations of functional patterns (e.g., promoter strength motifs, guide constraints).
- Products/workflows:
- Design UIs that accept exemplar sequences and constraints, returning candidate designs with uncertainty estimates.
- Closed-loop lab integration for iterative validation.
- Assumptions/dependencies: Safety and dual-use risk management, integration with wet-lab data, validation across diverse sequence families; potential need for hybrid training (generative + discriminative).
- Standards and governance for emergent capabilities in non-linguistic foundation models
- Sector: policy, standards bodies, industry consortia
- What: Formalize tests for emergent ICL across biological sequence models; require capability disclosure beyond language benchmarks.
- Products/workflows:
- Certification checklists including cross-domain ICL tests, mode-baseline comparisons, and BitLoad profiling.
- Reporting guidelines for open releases and clinical-adjacent tools.
- Assumptions/dependencies: Community consensus on metrics; incentives or mandates to adopt such standards.
- Multimodal biological foundation models with robust ICL (genomics, proteomics, single-cell, epigenomics)
- Sector: academia, industry research
- What: Train next-token predictors across diverse biological modalities to harness modality-agnostic ICL for cross-assay generalization and transfer.
- Products/workflows:
- Unified sequence assistants that can learn task-specific mappings from few examples across data types.
- Cross-modal prompt curricula for lab workflows.
- Assumptions/dependencies: Scaled training data and compute; careful modality alignment; safety and interpretability.
- BitLoad-informed curriculum design and robustness training
- Sector: ML research, software tooling
- What: Use BitLoad (dependency depth) to design curricula that gradually increase task complexity, improving robustness to higher-dependency transformations.
- Products/workflows:
- Pretraining schedules and evaluation suites stratified by BitLoad.
- Automated prompt selection to match target task complexity and available shot budget.
- Assumptions/dependencies: Empirical confirmation that BitLoad correlates with downstream task difficulty in real biological tasks; tooling to estimate task complexity.
- General “sequence assistant” for symbolic domains beyond biology
- Sector: cybersecurity (malware bytecode), DevOps (log analysis), finance (tick-level time series), industrial IoT
- What: Train next-token predictors on non-linguistic sequence data to unlock few-shot pattern induction (e.g., anomaly detection, local transformations, protocol recognition).
- Products/workflows:
- Prompt-driven analyzers that learn rules from small exemplar sets (e.g., identifying packet patterns or event signatures).
- Workflows that exploit log-linear gains with shots to tune demonstration size.
- Assumptions/dependencies: Availability of large, structured sequence datasets; task formulations compatible with symbolic prompting; domain-specific validation.
- Architecture and systems design for non-linguistic ICL
- Sector: ML systems, hardware-software co-design
- What: Explore hybrid architectures (e.g., convolution + attention, long context memory) that support ICL in million-token contexts, as seen in Evo2, for other sequence domains.
- Products/workflows:
- Memory- and context-optimized inference stacks for long-range sequence modeling.
- Benchmarks to compare transformer-only vs. hybrid models on emergent ICL.
- Assumptions/dependencies: Sustained research investment; evidence that architectural choices materially improve non-linguistic ICL beyond data/compute scaling.
- DNA data storage management with adaptive error correction
- Sector: data storage, biotech
- What: Apply emergent ICL to learn few-shot error correction or transformation rules for DNA-based storage systems.
- Products/workflows:
- Adaptive decoders that refine error models with a handful of exemplar corrupted/clean pairs.
- Maintenance tools that learn new artifact patterns on-the-fly.
- Assumptions/dependencies: Mature DNA storage deployments; mapping of storage error patterns to structured sequence transformations; rigorous reliability testing.
Collections
Sign up for free to add this paper to one or more collections.