- The paper presents an ontology-driven approach that leverages UMLS and a modified Tversky Index to enhance radiology report retrieval, yielding higher AUROC than embedding baselines.
- The method employs a four-stage pipeline including entity extraction with RadGraph-XL and CUI linking via SapBERT to ensure clinically meaningful alignment.
- The framework improves rare disease detection and labeling accuracy by integrating structured medical knowledge, offering interpretable and efficient retrieval for clinical AI.
Ontology-Based Concept Distillation for Radiology Report Retrieval and Labeling
Introduction
This paper presents an ontology-driven framework for radiology report retrieval and labeling, specifically targeting long-tail medical imaging tasks such as rare disease detection in chest X-rays. The authors critique the prevailing reliance on high-dimensional text embeddings (e.g., CLIP, CXR-BERT) for report similarity, highlighting their lack of interpretability, computational inefficiency, and poor alignment with the structured nature of medical knowledge. In response, the proposed method leverages the Unified Medical Language System (UMLS) ontology to extract and compare clinically grounded concepts, yielding a transparent, interpretable, and semantically meaningful approach to report retrieval and labeling.
Methodology
The framework comprises four principal modules: entity extraction, similarity search, retrieval experiment, and labeling procedure.
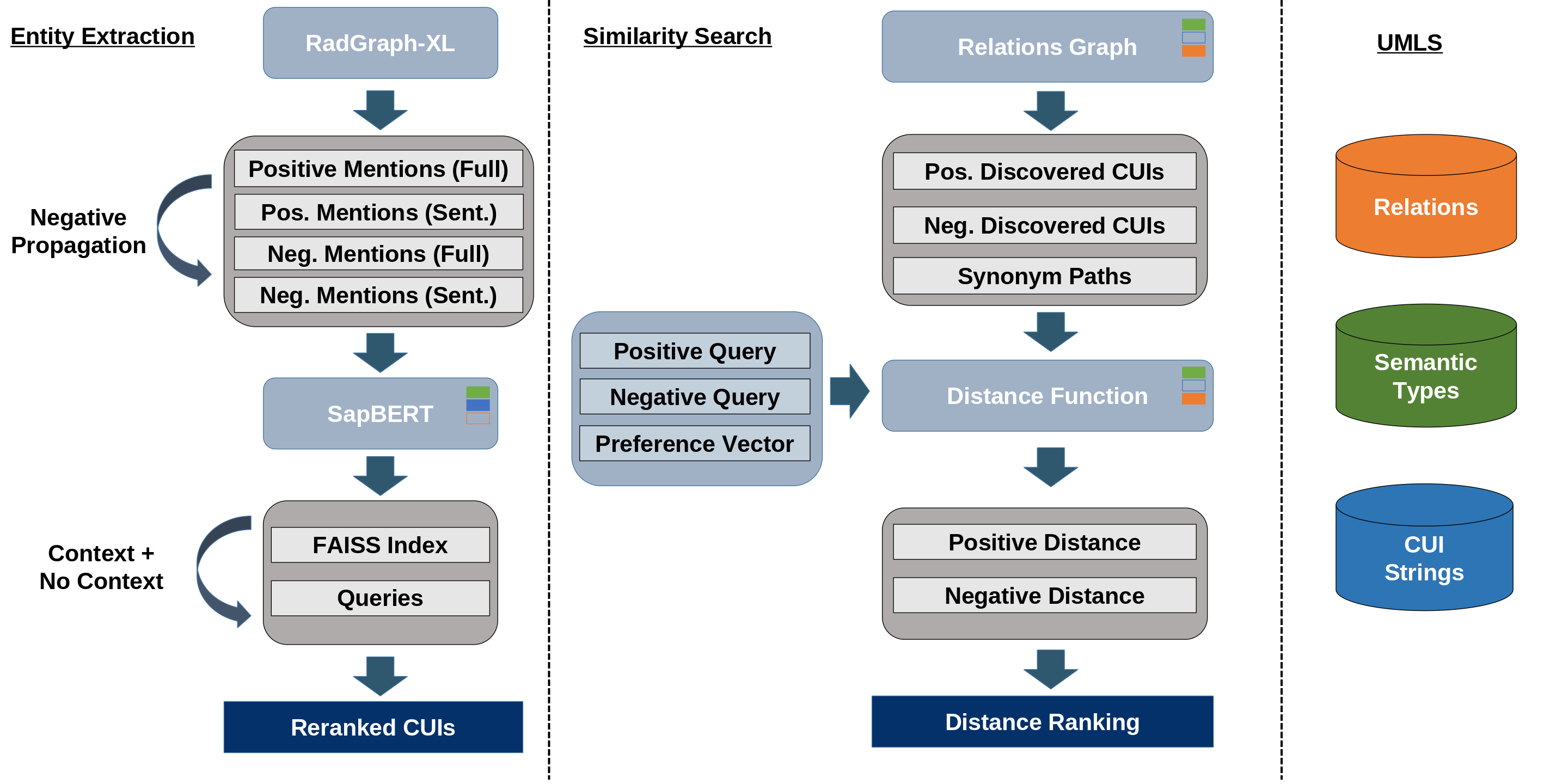
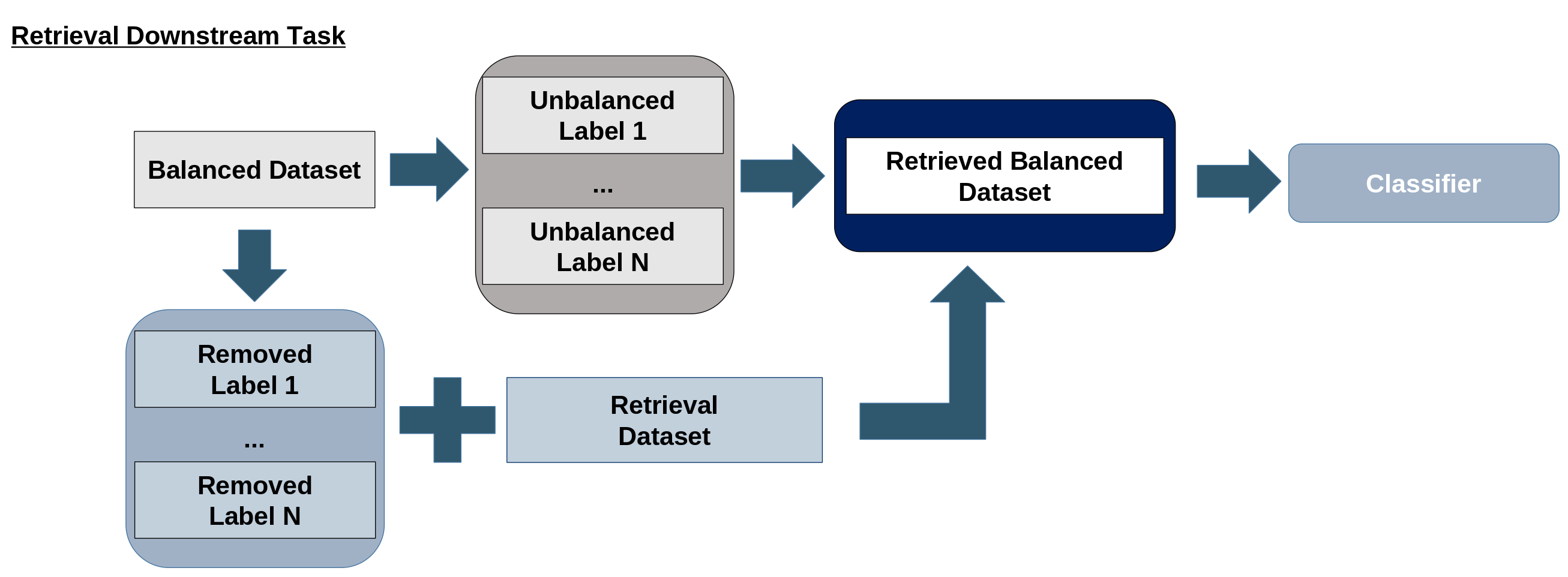
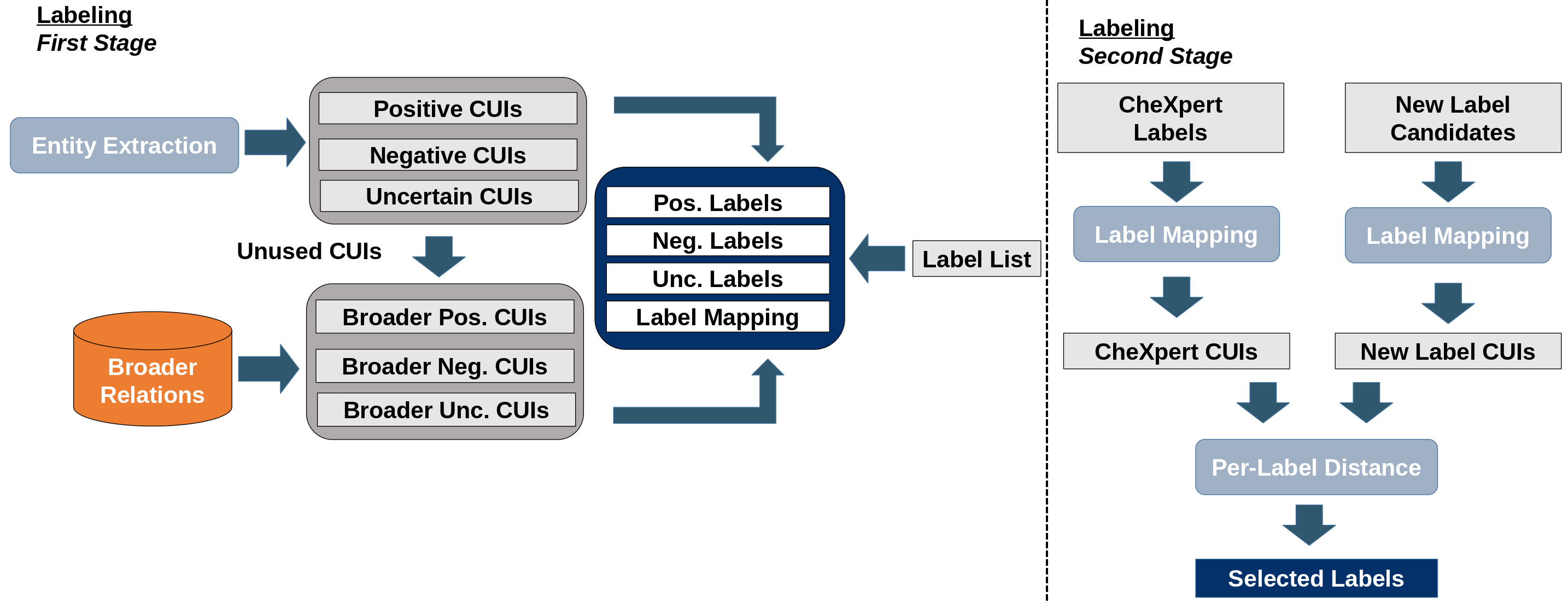
Figure 1: The method's four modules: entity extraction, similarity search, retrieval experiment, and labeling procedure.
Medical entities are extracted from free-text radiology reports using RadGraph-XL, which identifies anatomical and observational entities along with their assertions (presence, absence, uncertainty). The pipeline propagates negations to anatomies lacking corresponding positive observations, addressing limitations in RadGraph-XL's assertion logic. Both full-report and per-sentence contexts are provided to RadGraph-XL to maximize extraction accuracy, with rule-based assertion handling for edge cases.
Entities are then linked to UMLS Concept Unique Identifiers (CUIs) using SapBERT, fine-tuned on SNOMEDCT-US vocabulary to ensure terminological consistency. SapBERT embeddings are computed for both isolated entities and their contexts, and top-128 cosine similarity search is performed to select the most semantically appropriate CUI for each entity.
Knowledge Graph Construction
A cyclic, undirected graph is constructed from UMLS relations emphasizing hierarchy and semantic closeness. This graph supports efficient candidate report retrieval via BFS and enables synonym discovery by traversing synonym relations. Semantic distances are not directly used due to computational overhead and lack of observed performance gains.
Preference-Based CUI Set Similarity
The core similarity metric is a modified, weighted Tversky Index, which generalizes set similarity measures (Jaccard, Dice-Sørensen) and incorporates asymmetric similarity, synonymy, negation, and hierarchical relationships. The symmetric reformulation dynamically selects the prototype set, and a contradiction term penalizes conflicting assertions. A task-adaptive preference vector filters and ranks semantic types, allowing the similarity measure to be tailored to specific downstream tasks (e.g., disease localization vs. disease detection).
Downstream Retrieval and Labeling
For evaluation, a DenseNet-121 classifier is trained on datasets retrieved using the proposed similarity metric, with performance compared to embedding-based and other ontology-based baselines. The labeling procedure uses CUI set representations to generate ontology-backed disease labels for MIMIC-CXR, improving label accuracy and downstream classification performance. The final label for each report is selected based on maximal containment similarity between report and label CUI sets.
Experimental Results
The proposed ontology-based retrieval methods (Jaccard and Prototypical Retrieval) consistently outperform embedding-based baselines (CXR-BERT, CLIP, Bio-ClinicalBERT) in radiograph classification tasks on MIMIC-CXR, particularly in long-tail settings. The mean AUROC for ontology-based methods is higher across most disease classes, with notable improvements for Atelectasis, Cardiomegaly, and Pleural Effusion. Path-based ontology retrieval yields inferior results, affirming the superiority of set-based approaches.
Labeling experiments demonstrate that the ontology-backed labels generated by the pipeline correct mislabelings present in CheXpert and outperform modern LLM-based labeling approaches for certain disease classes. The asymmetric similarity measure further enhances labeling accuracy compared to symmetric baselines.
Discussion
The results substantiate the claim that embedding-based similarity is not optimal for long-tail retrieval in clinical domains, where interpretability and domain knowledge integration are critical. The ontology-based approach provides transparent, clinically meaningful similarity assessments and enables the generation of high-quality, ontology-backed labels for large-scale datasets. The modular pipeline facilitates adaptation to diverse retrieval and labeling tasks by adjusting the preference vector and semantic type filters.
Limitations include the exclusion of certain ontology-based retrieval algorithms and IC metrics due to scalability constraints. Entity extraction and linking remain bottlenecks, as even state-of-the-art models like RadGraph-XL exhibit assertion errors in simple cases. The framework's reliance on SNOMEDCT-US vocabulary may limit generalizability to other medical ontologies.
Implications and Future Directions
Practically, this work enables more reliable and interpretable retrieval-augmented learning in clinical AI systems, with direct applicability to rare disease detection, dataset curation, and automated labeling. Theoretically, it advances the integration of structured medical knowledge into retrieval and classification pipelines, challenging the dominance of opaque embedding-based methods.
Future research should focus on extending the pipeline to other imaging modalities and ontologies, improving entity extraction and linking accuracy, and exploring the impact of ontology-backed labels on advanced multimodal learning tasks. The generated MIMIC-CXR labels constitute a valuable resource for benchmarking and training in medical imaging research.
Conclusion
This paper introduces a robust ontology-based framework for radiology report retrieval and labeling, demonstrating superior performance and interpretability compared to embedding-based baselines in long-tail classification tasks. The integration of UMLS concepts, task-adaptive similarity metrics, and ontology-backed labeling establishes a new paradigm for clinically meaningful retrieval in medical AI, with significant implications for both research and real-world deployment.


