- The paper introduces the GT4SD toolkit that consolidates diverse generative models into a unified Python-based API for material and drug discovery.
- It demonstrates a molecular optimization pipeline that applies both conditional and unconditional generative methods to enhance candidate property profiles.
- The framework emphasizes reproducibility and democratization by integrating model lifecycle management, standardized interfaces, and accessible CLI/web tools.
Motivation and Context
The protracted pace and escalating costs of hypothesis generation in fields such as material and drug discovery are pressing challenges, with vast chemical spaces (>1033 molecules) precluding exhaustive, human-driven search. Data-driven generative models, including VAEs, GANs, GFlowNets, and diffusion models, have recently been adopted for efficient navigation of these spaces to design molecules or materials that conform to specific target properties. Despite rapid methodological advances, the application of generative models in scientific discovery is hampered by the lack of unified, accessible software environments, standardized interfaces, and model versioning, all critical for democratizing the field and supporting an open-science paradigm.
The GT4SD Library: Design Principles and Capabilities
The Generative Toolkit for Scientific Discovery (GT4SD) addresses these critical gaps with an extensible Python-based library that consolidates state-of-the-art (SOTA) generative pipelines for material design, encapsulating training, inference, and model lifecycle management within a harmonized API. GT4SD's infrastructure is modular, facilitating seamless integration of newly developed algorithms and supporting interoperation with wider scientific machine learning libraries, including PyTorch, Hugging Face Transformers, Diffusers, and targeted packages such as GuacaMol and MOSES.

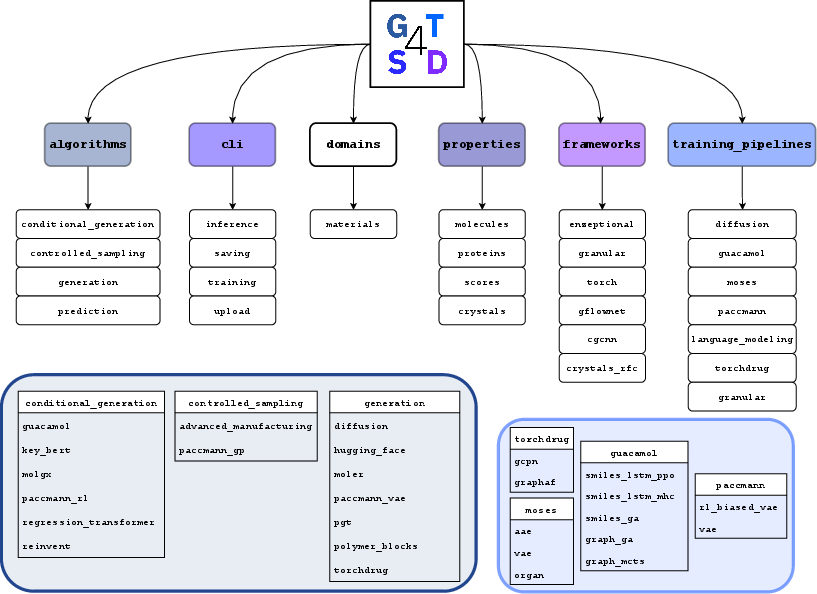
Figure 1: Schematic overview of GT4SD’s architecture, illustrating its standardized interfaces for generative model instantiation, CLI tools, and integration with material-science-focused algorithm and property prediction frameworks.
The library features:
- Unification of Algorithms and Interfaces: A single application registry for generative models and property predictors obviates the need for users to interact with disparate codebases or ad hoc wrappers.
- Pipeline Abstraction: Both inference and training are exposed via high-level, standardized classes, with further fine-grained control available through configurable data classes for each model family.
- CLI and Web Integration: All library features are accessible via CLI endpoints, and inference over pre-trained models is further democratized through browser-based web applications.
- Model Versioning and Distribution: Utilities for local or hub-based model versioning facilitate collaborative, reproducible science.
Algorithmic Scope and Interoperability
GT4SD encompasses comprehensive support for both unconditional and conditional generative models trained on molecular graphs (e.g., MoLeR, TorchDrug, GraphAF) and sequence-based representations (via SMILES, SELFIES, etc.), interfaced through MOSES or GuacaMol wrappers. Algorithms range from standard VAEs and AAE/ORGAN variants to GFlowNets and diffusion models, with support for both random sampling and property/structure-conditioned candidate generation.
The toolkit also bridges to downstream property prediction using integrated regression and classification models for small molecules, proteins, and crystalline materials, closing the loop for active design and optimization cycles. Advanced model release and deployment are structured towards instant integration into custom discovery workflows.
Case Study: Molecular Optimization Pipeline
An applied molecular discovery scenario exemplifies GT4SD’s capabilities. The workflow involves starting with a literature-derived hit compound (gentrl-ddr1, a DDR1 kinase inhibitor identified by deep RL [zhavoronkov2019deep]), and executing a constrained search for analogs with enhanced estimated water solubility (ESOL)—a significant real-world optimization objective for drug development.
The process entails:
- Sampling candidate molecules from the local chemical space surrounding gentrl-ddr1, using both unconditional generative models and conditional methods (MoLeR, REINVENT, Regression Transformer).
- Imposing structure similarity constraints (e.g., Tanimoto similarity) and/or direct conditioning on ESOL values.
- Quantitatively, conditional generative approaches delivered candidates respecting the similarity constraint (Tanimoto >0.5) with a statistically significant improvement in ESOL, exceeding 1M/L in several cases.
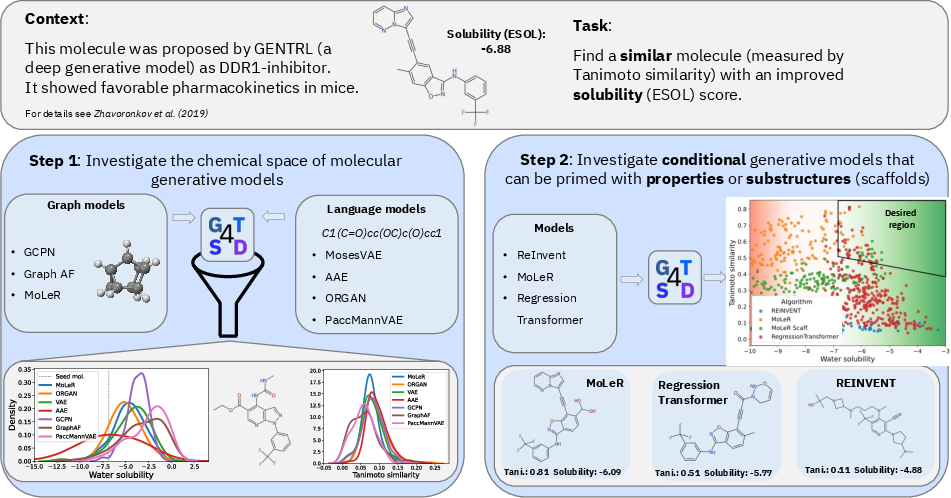
Figure 2: Illustration of GT4SD-driven iterative candidate generation, showing workflow from hit selection, through both unconditional and property-constrained generative pipelines, to output set evaluation.
This pipeline streamlines the translation from hit identification to lead optimization and prototyping, bypassing substantial human and computational cost.
Architecture and Model Lifecycle Management
The API ensures model serving, training, and lifecycle events (persistence, sharing, deployment) strictly follow a standardized protocol. Inference pipelines handle local caching, cloud-synced model storage, and algorithm instantiation regardless of back-end ML library. Training pipelines are designed for arbitrariness in hyperparameterization and data configuration. CLI endpoints and associated commands (for running, configuring, saving, and uploading models) automate discovery workflow management in a reproducible manner, essential for collaborative open science.
Practical and Theoretical Implications
By amalgamating disparate generative pipelines and predictive analytics in a unified, extensible interface, GT4SD enables researchers at institutions of all sizes to access, evaluate, and extend SOTA algorithms for scientific design and discovery. The implications are multifold:
- Democratization: Substantially lowers technical barriers, enabling non-experts and students to deploy advanced generative models with minimal code.
- Standardization and Reproducibility: Common interfaces and model versioning protocols foster consistency and facilitate direct comparative evaluation and benchmarking.
- Acceleration of Discovery: Streamlined integration and orchestration reduce human effort and time-to-insight for hit-to-lead material or molecule optimization workflows.
- Open Science Synergy: Integrated release, sharing, and fine-tuning ecosystems promote rapid dissemination and collaborative extension of new generative architectures.
Future Directions
Ongoing and anticipated developments include expanding application domains to encompass climate, earth sciences, and sustainability; integrating new classes of generative models; and strengthening model evaluation by extending current benchmarks (e.g., GuacaMol, Moses) with additional bias and property metrics. The vision includes a community-driven model hub akin to Hugging Face for generative science workflows, enabling users to fine-tune, upload, and leverage models tailored to their domain-specific datasets.
Conclusion
GT4SD represents a robust, harmonized framework facilitating end-to-end generative modeling workflows for material design, aggregating SOTA algorithms, streamlined APIs, and model sharing protocols. This infrastructure is poised to be instrumental in rendering advanced generative techniques accessible and interoperable for a broad spectrum of researchers, thus catalyzing innovation and progress in material and drug discovery through open, collaborative computational science.
(2207.03928)


