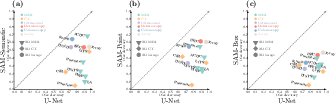- The paper finds that SAM's zero-shot segmentation consistently yields lower Dice scores than specialized medical models.
- It utilizes a comprehensive analysis over 12 datasets to assess how factors like dimensionality, contrast, and target size affect segmentation accuracy.
- Results indicate SAM performs better on higher contrast 2D images, while struggling with complex 3D and low contrast scenarios, highlighting areas for improvement.
The Segment-Anything Model (SAM) in Medical Image Segmentation
Introduction
The Segment Anything Model (SAM) marks a significant advancement in computer vision, characterized by its ability to generalize across diverse image segmentation tasks without requiring task-specific re-training or fine-tuning. While SAM demonstrates impressive zero-shot performance in natural image segmentation, its accuracy and applicability to medical imaging tasks—characterized by complexity and diversity—remains uncertain. This essay discusses the exploratory study evaluating SAM's efficacy in medical image segmentation across multiple datasets, its comparative performance relative to established medical-image-specific algorithms, and the influencing factors impacting its segmentation accuracy.
Methods
The study employs SAM's zero-shot capability to segment medical images without any retraining, assessing its performance across 12 public datasets comprising 7,451 subjects and spanning a variety of organs, pathologies, dimensions, and imaging modalities. The accuracy of SAM is measured using the Dice coefficient, comparing its performance with five state-of-the-art algorithms tailored for medical image segmentation tasks like U-Net, U-Net++, Attention U-Net, Trans U-Net, and UCTransNet. Diverse other factors including segmentation difficulty, image dimension, modality, target region size, and contrast were analyzed for their potential influence on SAM's accuracy.
Evaluation of SAM's Accuracy
The results indicate SAM's performance lagging behind specialized medical-image segmentation algorithms. In all datasets, SAM exhibited lower Dice scores by margins of 0.1 to 0.7, underscoring the challenges posed by medical images differing from natural images in terms of complexity, dimensionality, and target characteristics.
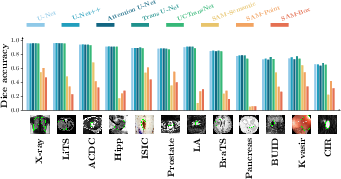
Figure 1: Accuracy of SAM in 12 medical image segmentation datasets. (a): Dice overlaps of 5 medical-image-specific algorithms and 3 variations of SAM. (b): scatter plots of SAM's Dice (y axis) with U-Net's Dice (x axis) across datasets, for SAM-Semantic (left), SAM-Point (middle), and SAM-Box (right).
Furthermore, SAM's lower accuracy is most pronounced in complex images—those with low contrast, smaller target regions, and 3D dimensions, corroborated by its comparatively better performance on 2D images with higher U-Net Dice values.
The degree to which different factors impact SAM's segmentation accuracy was analyzed using both single-factor and multi-factor analyses. It was observed that SAM's accuracy was significantly influenced by image dimensionality, segmentation ability scores, and target region characteristics.
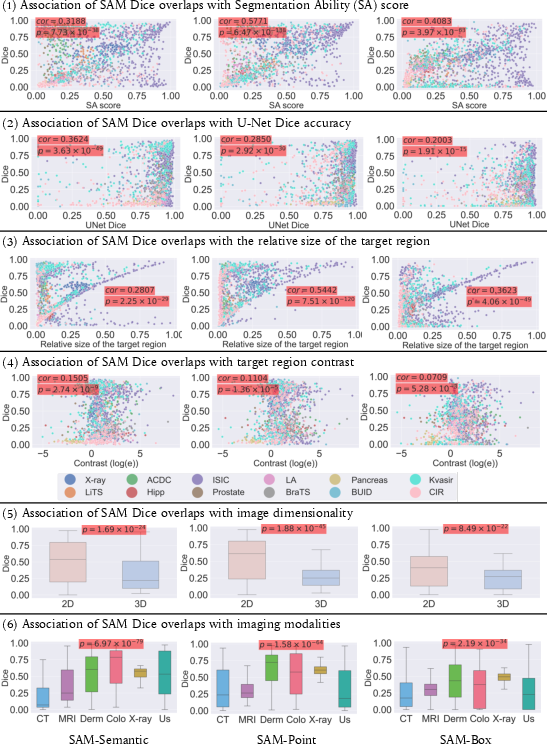
Figure 2: Single-factor analysis associating 6 potential factors with SAM's accuracies in 12 medical image datasets. In the top four rows, every dot is a subject, and the color of the dot denotes the dataset this subject comes from. p-values are provided and highlighted with a red background if a factor has a significant association with the SAM Dice overlap. In the 6th row, Derm--Dermology; Colo--Colonoscopy; US--Ultrasound.
SAM variants demonstrated different sensitivities to these factors: SAM-Semantic showed an association with medical image segmentation difficulty, whereas SAM-Point and SAM-Box were more impacted by target region size, dimensionality, and contrast.
Discussion
The analysis confirms that while SAM represents a step towards unified segmentation across different contexts, its performance in medical imaging is sub-optimal compared to specialized models. The distinctive characteristics of medical images, such as higher dimensionality, lower contrast, and smaller target regions, pose unique challenges that SAM currently does not surmount effectively.
SAM's underperformance highlights the necessity for strategic adaptations or alternative frameworks when applying zero-shot segmentation models to medical images. Promising avenues for future research include the development of medical-image-specific benchmark models or adapting existing models by incorporating medical image datasets into their training regime.
Conclusion
In sum, this study provides a critical evaluation of SAM's capabilities in medical image segmentation and sets the groundwork for future explorations. Enhancements to SAM or the development of specialized models tailored to medical imaging needs are imperative for achieving comparable accuracy to current state-of-the-art algorithms. Such efforts must consider the unique characteristics inherent in medical images—including 3D imaging, small and low-contrast target regions—that challenge current methodologies.