- The paper introduces a weakly-supervised U-Net model using scribble-based annotations to segment prostate cancer aggressiveness with performance near fully supervised methods.
- Experiments using only 6.35% annotated voxels achieved lesion-wise Cohen’s kappa scores close to baseline values, highlighting efficient segmentation.
- The method reduces annotation effort, integrates heterogeneous datasets, and offers promising clinical applications in rapid MRI-based cancer screening.
Learning to Segment Prostate Cancer by Aggressiveness from Scribbles in Bi-parametric MRI
Introduction to Weakly-supervised Segmentation in Medical Imaging
The task of segmenting prostate cancer (PCa) aggressiveness in bi-parametric MRI, using scribble annotations, presents a significant challenge in medical imaging. While deep learning models for detection and segmentation typically require extensive fully annotated data, such data is often challenging to obtain due to the expertise and time required for annotation. The paper addresses this dilemma by employing a weakly-supervised learning approach, leveraging only partial labels—scribbles—to perform segmentation. This method is notably beneficial in clinical settings where rapid annotation is necessary. By focusing on cancer aggressiveness characterized by the Gleason score (GS), the study explores how biopsy sample data can facilitate assessing lesion extent without exhaustive annotation, augmenting datasets like ProstateX-2, which lacks comprehensive lesion contours.
Data Collection and Preparation
The study utilizes two key datasets: a proprietary dataset containing precise axial T2 weighted (T2w) and apparent diffusion coefficient (ADC) MR images from 219 patients, and the ProstateX-2 challenge dataset with data from 99 patients. The proprietary dataset is meticulously annotated with pixel-level lesion details and GS values, while ProstateX-2 provides only the lesion centroid and GS. To tackle the segmentation problem, circular scribble annotations are automatically generated with constrained radii across slices, enabling rapid annotation without exhaustive detail.
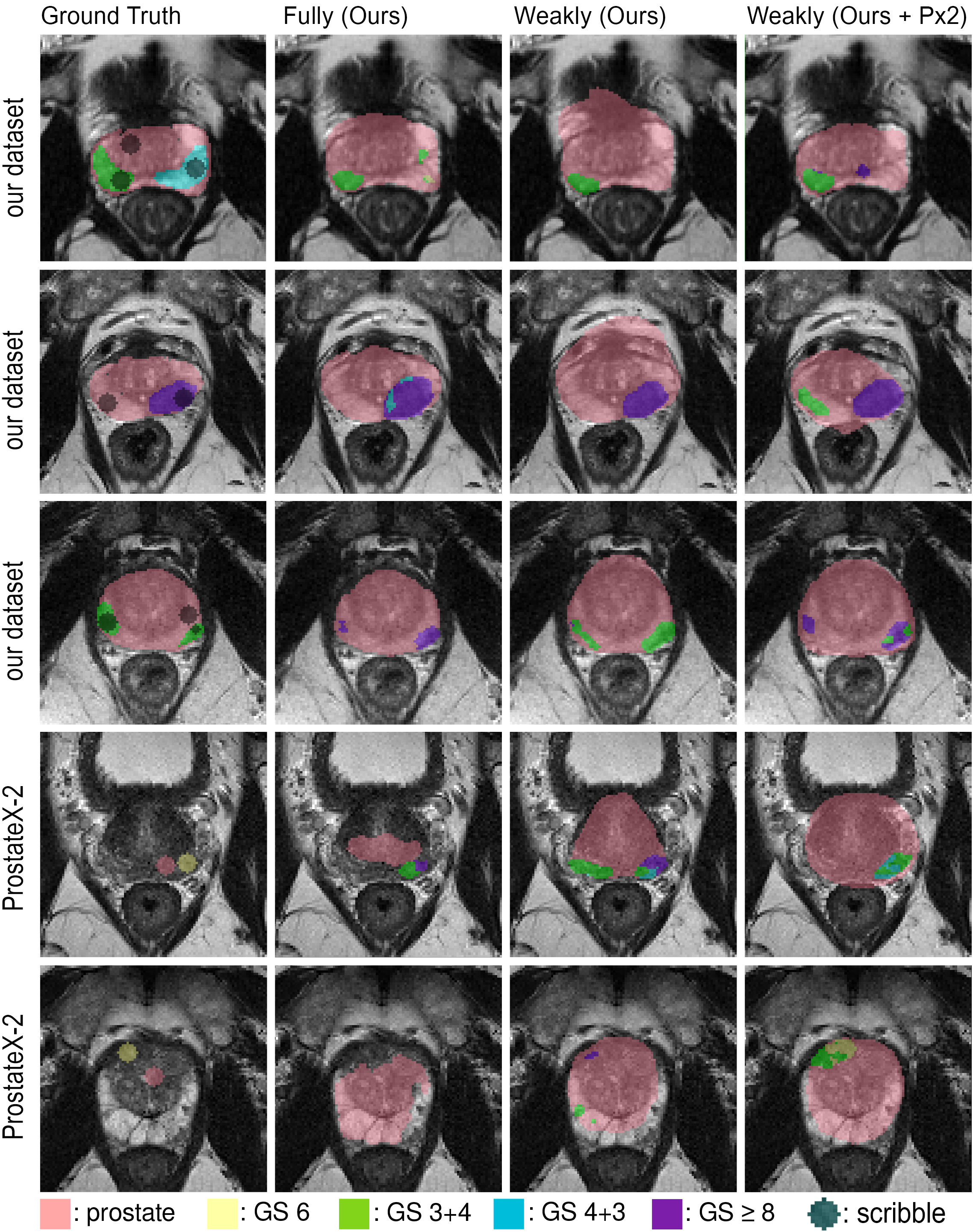
Figure 1: Input distribution and preparation for lesion segmentation via scribbles.
Weak Multi-class Segmentation Model
To address the challenges of weak supervision, the paper adopts a U-Net-based architecture with a custom constrained loss function informed by prior work from Kervadec et al. This loss integrates partial cross-entropy focused on annotated pixels and a constraint penalizing segmentations deviating from defined size bounds. The architecture favorably adapts to multi-class outputs, using Gleason scores to label segmentation classes. The implementation benefits from data augmentation and a systematic training regimen that mitigates overfitting and leverages 3D connectivity to derive final lesion maps.
Experimental Results
Experiments reveal the effectiveness of the proposed weak U-Net model. Using only 6.35% annotated voxels, the model closely approaches the performance of a fully supervised baseline in segmenting lesions based on aggressiveness. The reported lesion-wise Cohen's kappa scores, particularly 0.29±0.07 compared to 0.32±0.05, illustrate the model's potential. Moreover, on the ProstateX-2 dataset, the weak U-Net achieves a leading kappa score of 0.276±0.037, outperforming previous reported values for segmentation tasks.
(Figure 2)
Figure 2: Comparison of segmentation maps obtained using different models on validation datasets.
Practical Implications and Future Work
The findings suggest substantial implications for clinical practice, notably in reducing the time required for training segmentation models without sacrificing significant performance. Furthermore, the use of weak annotations supports integrating diverse datasets, potentially addressing domain shift issues prominent in medical imaging. Looking forward, this methodology encourages exploring hybrid models combining fully and weakly annotated data and refining lesion shape constraints for segmentation tasks to improve precision further.
Conclusion
The paper successfully demonstrates that through strategic use of weak supervision and modified loss functions, it is possible to segment prostate cancer aggressiveness with high efficiency, comparable to fully supervised models. The method showcases crucial advancements in using MRI for cancer screening, emphasizing the value of integrating less exhaustive annotative efforts for robust clinical applications. With continuous exploration and refinement, such techniques hold promise for broader adoption in medical imaging tasks, particularly where annotation is resource-intensive.
