ConforNets: Latents-Based Conformational Control in OpenFold3
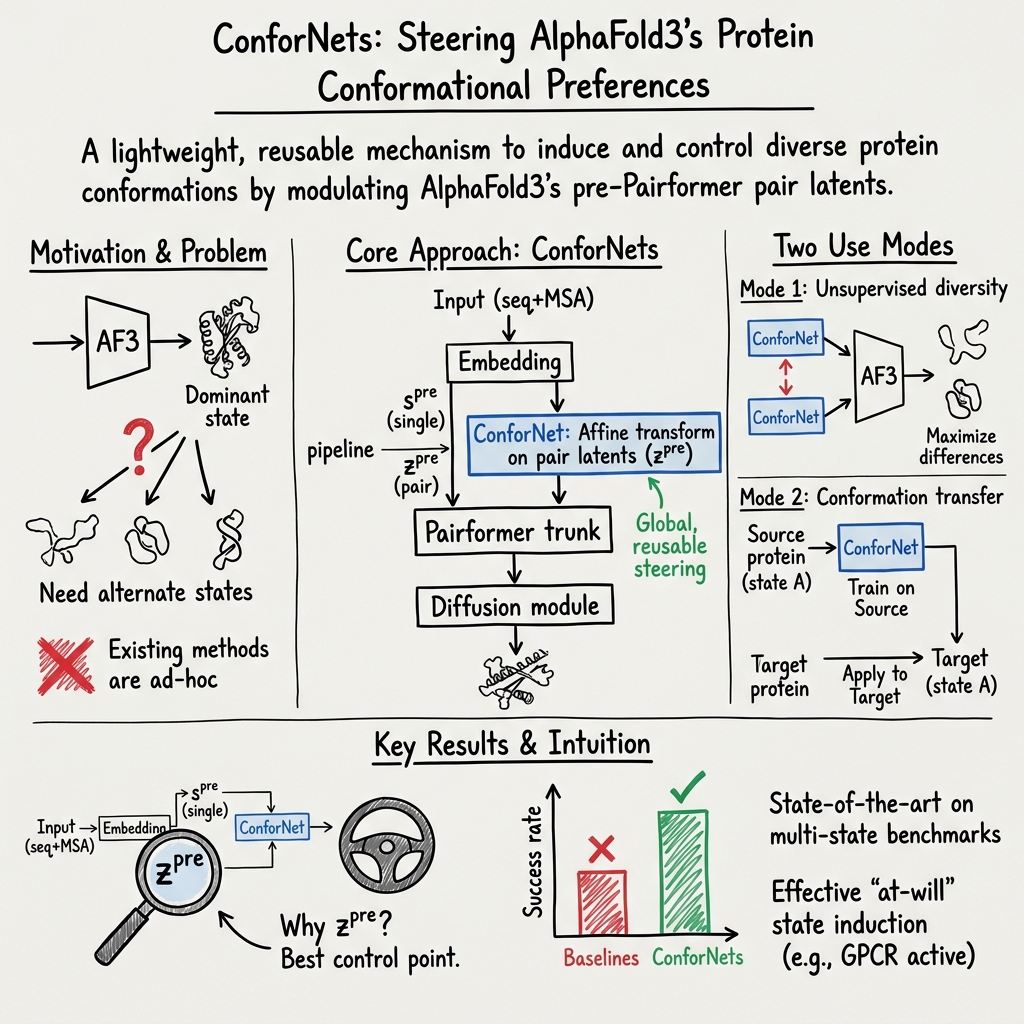
Abstract: Models from the AlphaFold (AF) family reliably predict one dominant conformation for most well-ordered proteins but struggle to capture biologically relevant alternate states. Several efforts have focused on eliciting greater conformational variability through ad hoc inference-time perturbations of AF models or their inputs. Despite their progress, these approaches remain inefficient and fail to consistently recover major conformational modes. Here, we investigate both the optimal location and manner-of-operation for perturbing latent representations in the AF3 architecture. We distill our findings in ConforNets: channel-wise affine transforms of the pre-Pairformer pair latents. Unlike previous methods, ConforNets globally modulate AF3 representations, making them reusable across proteins. On unsupervised generation of alternate states, ConforNets achieve state-of-the-art success rates on all existing multi-state benchmarks. On the novel supervised task of conformational transfer, ConforNets trained on one source protein can induce a conserved conformational change across a protein family. Collectively, these results introduce a mechanism for conformational control in AF3-based models.
Paper Prompts
Sign up for free to create and run prompts on this paper using GPT-5.
Top Community Prompts
Explain it Like I'm 14
What this paper is about
Proteins don’t hold a single frozen shape. They move between different shapes (called “conformations”) to do their jobs, like doors opening and closing. Tools like AlphaFold can predict a protein’s main shape very well, but they often miss other important shapes.
This paper introduces a simple add‑on, called ConforNets, that acts like a set of small “control knobs” inside an AlphaFold3‑style model (OpenFold3). These knobs let scientists steer the model toward different, useful shapes—either to explore multiple possibilities or to specifically push a protein into a known functional state (like an “active” form).
The main questions the authors asked
- Where inside an AlphaFold3‑like model is the best place to “nudge” things so you can change the protein’s predicted shape without breaking the model?
- Can a tiny, reusable steering tool work across many different proteins, not just one at a time?
- Can we:
- Unsupervised: get the model to produce several different sensible shapes for the same protein, on demand?
- Supervised: teach the model a particular shape change from one protein (a “source”) and then transfer that change to other family members?
How they did it (in simple terms)
Think of the model like a factory that:
- Reads the protein’s “recipe” (its amino acid sequence and related evolutionary info),
- Builds internal “notes” about which parts of the protein might be close together,
- Then turns those notes into 3D structures.
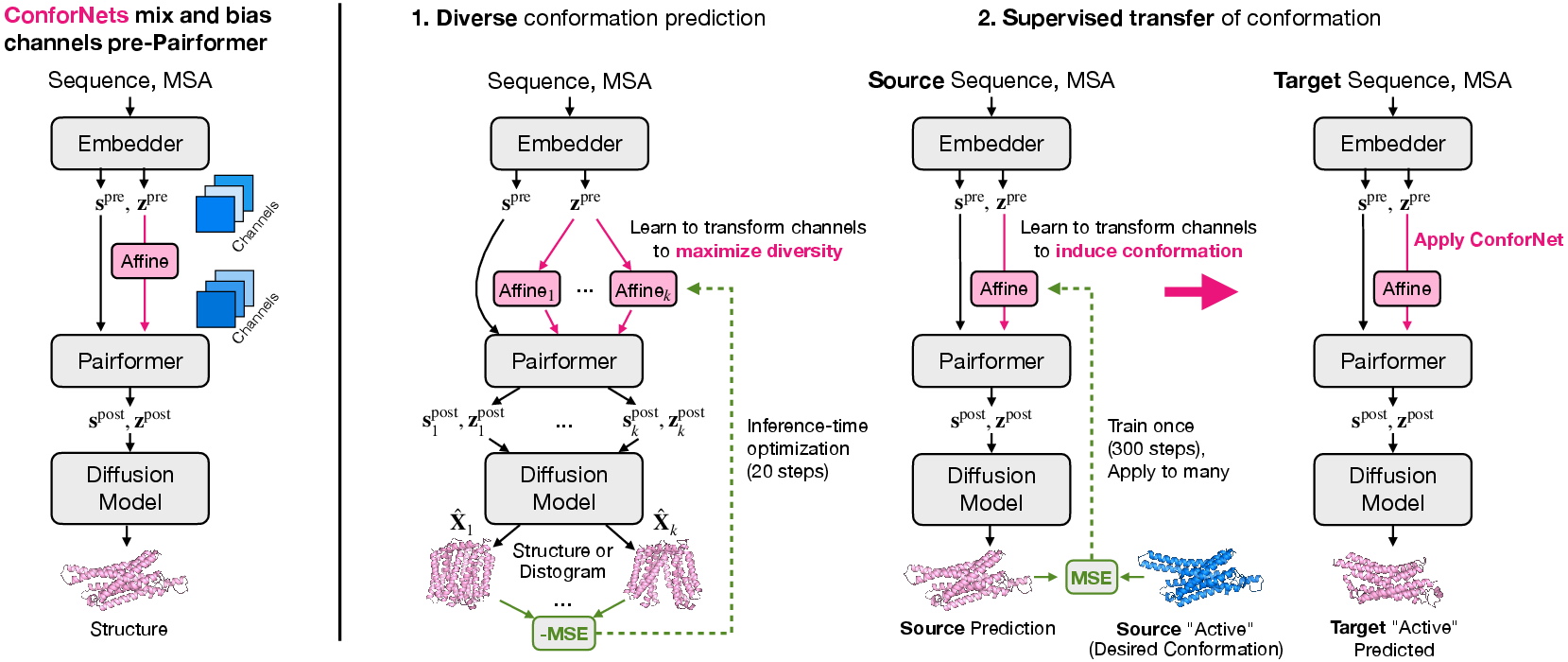
The authors add ConforNets right before a key processing stage called the “Pairformer.” At this point, the model holds a big “distance map”–style set of notes (called pair latents) about which amino acids might contact each other.
- What is a ConforNet?
- It’s a tiny mathematical layer that “remixes” the channels of these notes—like adjusting the sliders on a music equalizer (it scales and mixes channels with a simple formula).
- Because it works on channels (not on individual residues), a trained ConforNet can be reused on different proteins.
- Why here?
- The authors tested several spots and found that nudging the “pair latents” just before the Pairformer gives the most reliable and powerful control. It lets the rest of the model reason as usual and produce physically sensible structures.
- Two ways they use ConforNets:
- Distograms: predicted probability distributions of distances between residues (like a fuzzy map of how far apart pairs of amino acids might be).
- Coordinates: the actual 3D positions of atoms after a quick “denoising” step (like unblurring a photo).
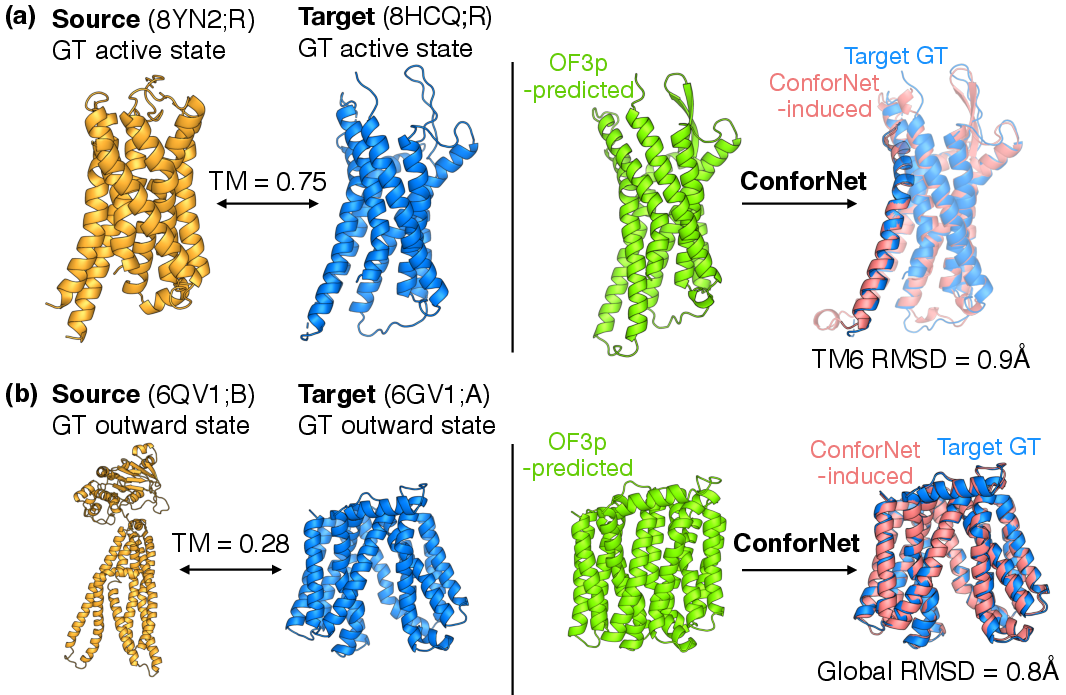
- 2) Supervised transfer: Train one ConforNet on a single protein with a known target shape (for example, the active form of a receptor), so that the model learns to prefer that shape. Then, apply that same ConforNet to other proteins in the same family to push them toward the corresponding shape.
- A note on “diffusion” and “denoising”:
- The structure generator uses a “diffusion” process, which starts with a noisy guess and “denoises” it to a clean 3D structure. ConforNets change the conditioning signal before diffusion starts, so the model still follows its normal, stable routine to make realistic structures.
What they found and why it matters
- Best place to steer: Adjusting the pair latents right before the Pairformer worked best. Tweaks applied later were less stable, and earlier spots didn’t give enough control.
- Unsupervised multiple shapes:
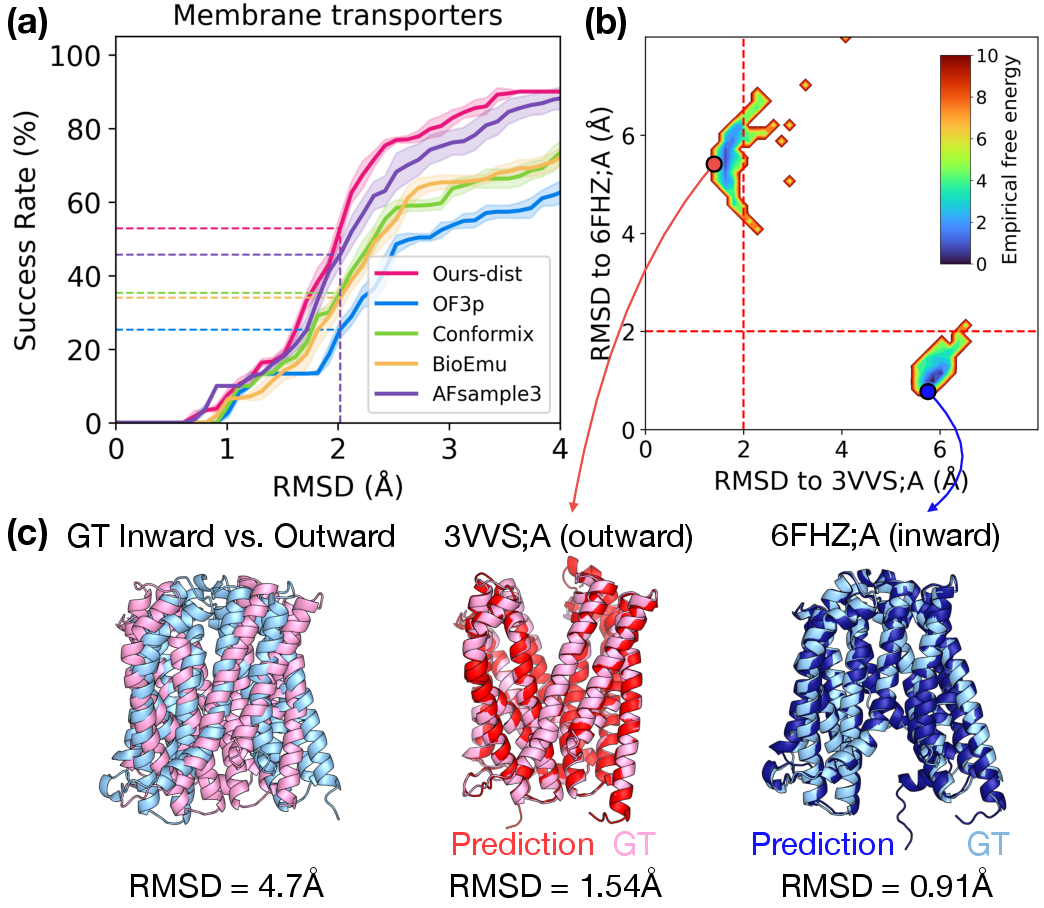
- Across several benchmarks (covering over 100 proteins with two known states each), ConforNets reached the new state‑of‑the‑art at finding alternate shapes. This was especially strong for membrane transporters (proteins that switch between inward‑ and outward‑facing forms).
- Compared to methods that randomly mess with the input data (like the MSA) or guide the last diffusion step, ConforNets’ intentional, upstream control produced more diverse and more plausible results.
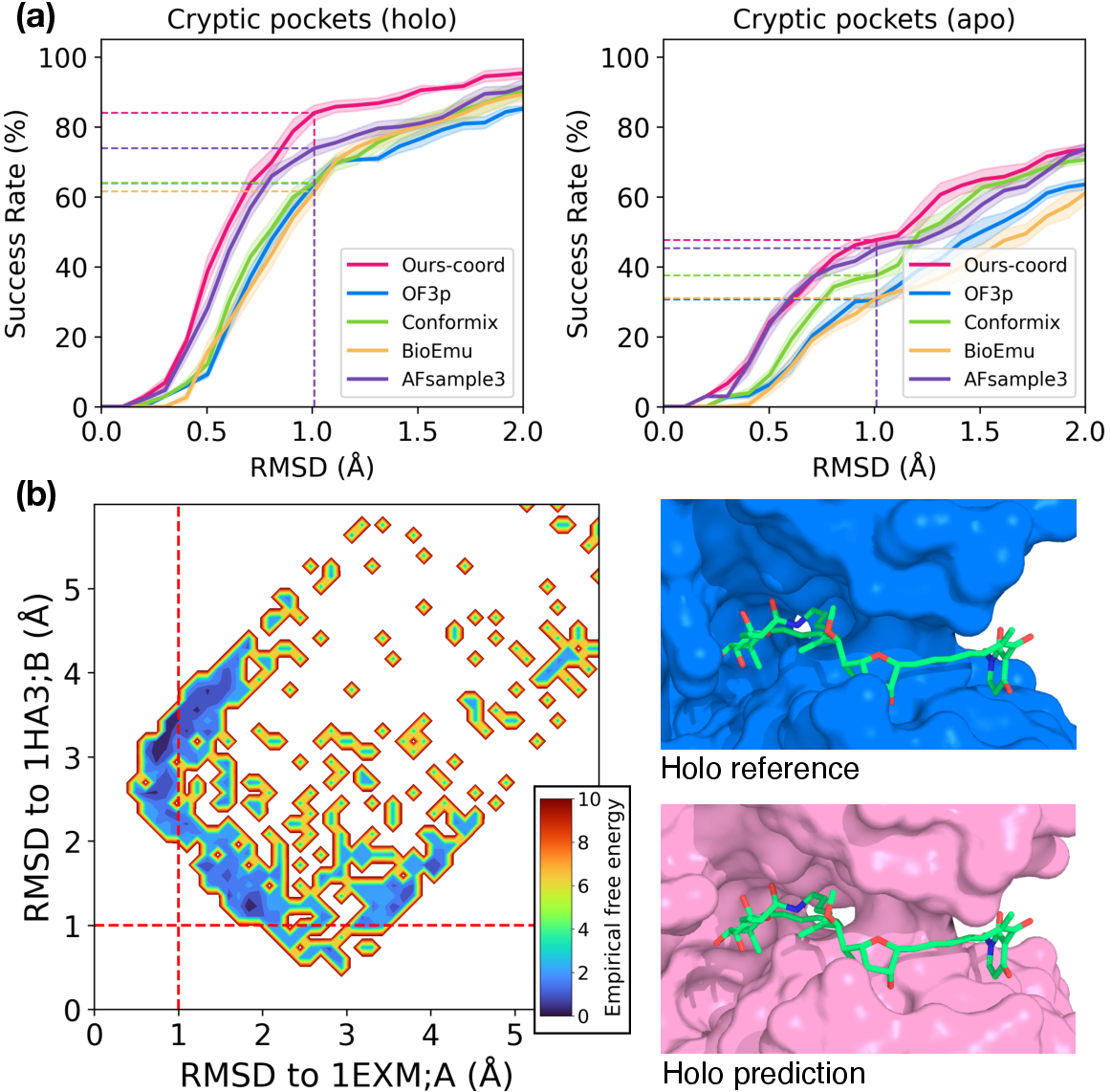
- Local changes when needed:
- Even though ConforNets make a global tweak, they can trigger very localized changes, like opening a hidden “cryptic pocket” (a small, drug‑binding cavity that appears only in certain shapes). That’s valuable for drug design.
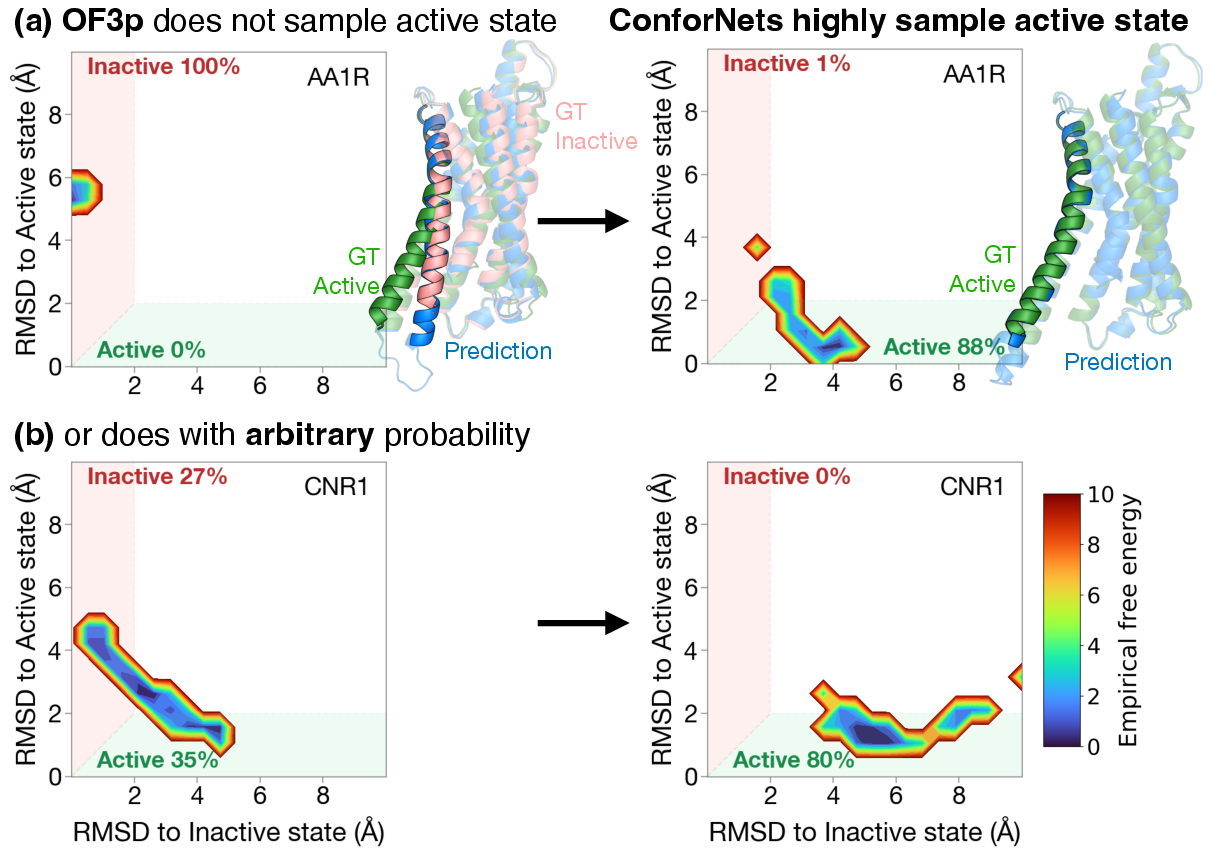
- Supervised conformation transfer:
- They trained a ConforNet on a single example protein to encode a known shape change, then applied it to other family members:
- GPCRs (important cell receptors): boosting success at getting the “active” state from about 24% to about 79% in a small sample test.
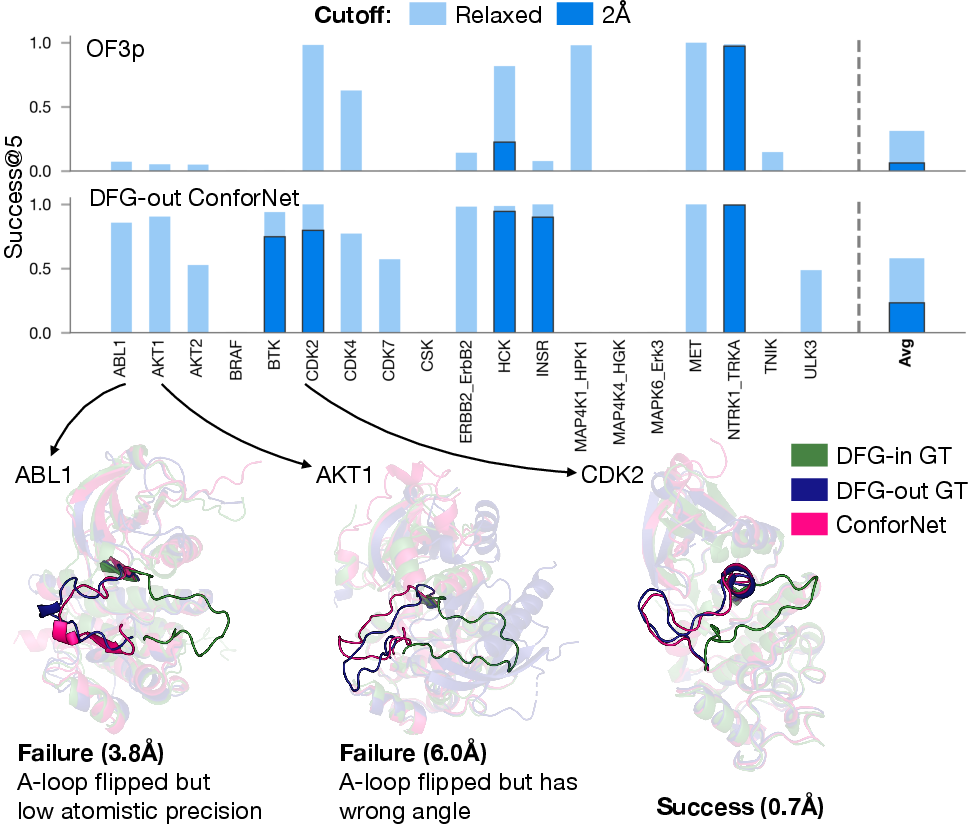
- Kinases (key signaling proteins): increasing the tough “DFG‑out” inactive state from about 6% to about 23%.
- Transporters: increasing the “outward‑open” state from about 16% to about 57%.
- This means you can often push a whole protein family toward a specific state using a ConforNet trained on just one protein.
- Efficiency and reuse:
- ConforNets are tiny and fast to use after training. Training them is relatively quick and the same trained ConforNet can be reused across proteins, giving you practical “knobs” for conformation control.
- Limits to keep in mind:
- The method does not tell you how common a shape is in nature (it’s not “energy‑calibrated”). It finds plausible shapes, not their exact probabilities.
- Some very flexible regions (like certain loops in kinases) remain challenging to place precisely.
Why this is important
- Controlled exploration: Instead of hoping random tricks will find alternate shapes, scientists get a practical steering tool to produce diverse, meaningful conformations.
- Better inputs for downstream tasks: Having the “active” form of a receptor or an “open” conformation of a transporter can make drug docking, mutational analysis, and simulation setup much more effective.
- Reusable “shape bias”: Training once and applying across a family saves time and gives consistent control—something that hasn’t been easy with earlier methods.
Big picture and future impact
ConforNets turn powerful structure predictors into more flexible tools that can explore and target protein shape changes on demand. This could help:
- Drug discovery, by revealing hidden pockets and active states,
- Mechanistic studies, by testing how proteins might switch between key conformations,
- Simulation workflows, by providing better starting shapes for molecular dynamics.
Next steps could include combining ConforNets with experimental data (like cryo‑EM or NMR constraints), calibrating the “energy feel” of the results, and pushing control down to finer details (like specific side‑chain arrangements) while keeping structures physically realistic.
Knowledge Gaps
Knowledge gaps, limitations, and open questions
The following list enumerates concrete gaps and open questions left unresolved by the paper, to guide follow-up research.
- Calibration of conformational distributions: ConforNets produce uncalibrated ensembles; no method is provided to estimate state probabilities or free energies, or to reweight samples to a Boltzmann-consistent distribution.
- Mechanistic interpretability of latent control: The paper shows pre-Pairformer pair latents are effective, but does not map which channels or operations (e.g., specific Pairformer updates) correspond to particular structural motions or contact changes.
- Side-chain and stereochemical quality: Evaluation focuses on backbone RMSD and pocket RMSD; the work does not report side-chain accuracy, clashes, bond/angle violations, Ramachandran outliers, or pLDDT/PAE confidence changes under perturbation.
- Prospective experimental validation: All results are retrospective benchmarks; no prospective tests (e.g., ligand binding assays, mutational scans, crosslink validation, cryo-EM map fitting) verify functional relevance of induced states.
- Dependence on MSA quality: ConforNets assume availability of deep MSAs and use per-step subsampling; robustness for shallow/noisy MSAs, single-sequence inputs, or taxonomically biased MSAs remains unquantified.
- Applicability beyond monomers: The study centers on monomeric cases; it does not test multimeric complexes, transient PPIs, or conformational changes driven by binding partners (e.g., GPCR–G protein complexes).
- Ligand, cofactor, and environment conditioning: Cryptic-pocket and transporter cases are evaluated without explicit ligand/membrane conditioning; it is unknown how ConforNets interact with small-molecule, metal, glycan, or membrane context in AF3/OF3p.
- Generalization breadth of transfer: Supervised transfer is demonstrated on three families (GPCRs, kinases, transporters); transfer to other families (e.g., ion channels, allosteric enzymes, antibodies), multi-domain proteins, and different topologies remains untested.
- Source selection for transfer: The “family centroid” heuristic is used but not systematically compared to alternatives (e.g., multiple sources, subfamily-specific sources, similarity-weighted ensembles, meta-learning across sources).
- Multi-state and continuum control: The method is trained with k=2 to target two-state benchmarks; its ability to induce >2 states, interpolate along reaction coordinates, or sample intermediate conformations is unstudied.
- Region-aware objectives for flexible loops: Kinase results highlight difficulty in precise loop rearrangements; region-targeted losses, torsion-based or angle-aware objectives, and local regularizers are not explored.
- Stability across seeds and hyperparameters: Sensitivity to k, learning rates, number of optimization steps, and objective choice (distogram vs coordinates) is incompletely characterized; automatic step selection (vs. step sweeps) is not addressed.
- Recycling strategy: ConforNets are applied only on the final Pairformer pass; whether earlier or multi-pass application (or per-recycle adaptations) improves control or stability is unknown.
- Mini-rollout optimization gap: One-step (or few-step) diffusion is used for fast optimization, yet can mismatch full-rollout behavior; principled surrogate losses or differentiable proxies that better correlate with full diffusion are not investigated.
- Expressivity of the transform: A global channel-wise affine transform may be underpowered; the benefits/risks of richer parameterizations (e.g., low-rank updates, per-head/per-block transforms, nonlinear MLPs, conditional gates) are not assessed.
- Spatial specificity and off-target changes: The extent to which ConforNets induce unwanted changes outside target regions (e.g., TM6 in GPCRs) is not quantified with regional metrics (e.g., per-residue RMSD/PAE).
- Confidence and uncertainty reporting: The method does not provide calibrated confidence for induced states (e.g., pLDDT shifts, pTM/PAE, ensemble agreement), which is critical for downstream decision-making.
- Benchmark metrics and thresholds: Success uses RMSD cutoffs that may not capture functional hallmarks (e.g., GPCR TM6 displacement angle, kinase DFG scoring, transporter gating metrics); state-specific criteria are not systematically evaluated.
- Data leakage assessments: OOD60 is OOD for BioEmu but not necessarily for AF3/OF3p; the paper does not audit training cutoffs or potential overlap between OF3p pretraining data and benchmark structures.
- Scaling with sequence length and complexity: Although the transform is L-independent, pair latents scale as O(L2); runtime/memory behavior, and performance on very large proteins or multi-domain assemblies remain uncharacterized.
- Synergy with other perturbations: Integration with MSA column/row strategies, dropout, or diffusion guidance (e.g., ConforMix) is not explored; combined methods may yield better coverage or control.
- Cross-family (universal) control priors: While claimed reusable across proteins, ConforNets are only demonstrated within families; whether family-agnostic ConforNets can encode generic motions (e.g., open↔closed) remains open.
- Transfer across large structural divergence: The paper reports some success at low TM-score, but systematic limits of transfer vs. structural/sequence divergence are not quantified (e.g., failure regimes, scaling laws).
- Template interactions: Template baselines use a single centroid template; whether multiple templates, template ensembles, or hybrid template+ConforNet strategies improve results is not investigated.
- Pathway and kinetics: The approach targets endpoints; it does not model transition paths or kinetics, nor does it test if generated intermediates (if any) are physically plausible.
- Energy landscape surrogates: The “empirical energy landscape” is a density over RMSDs and explicitly not calibrated; methods to build or learn better energy proxies or to couple with physics-based reweighting are not proposed.
- Robustness across MSA subsampling: ConforNets are trained with different MSA subsamples each step; the stability of a trained ConforNet when applied with different MSAs (or updated databases) is not quantified.
- Downstream task impact: Improvements in docking, virtual screening, or MD initialization are posited but not empirically evaluated (e.g., docking enrichment, MD convergence vs. baselines).
- Application to nucleic acid–binding proteins and hybrid complexes: Transfer/control for protein–DNA/RNA complexes or RNPs is not tested, despite AF3’s broader modality ambitions.
- Reproducibility and implementation details: Precise hardware/time profiles, randomness control, and open-source checkpoints for trained ConforNets (beyond OF3p weights) are not provided for full reproducibility.
These gaps suggest concrete avenues: develop calibrated scoring/reweighting, add region-aware objectives, explore richer transforms and multi-pass application, broaden family coverage and multimers/ligands, integrate physics or experimental constraints, and establish prospective, functionally grounded validation.
Practical Applications
Immediate Applications
Below are concrete, deployable use cases that can be implemented with the paper’s current methods (OpenFold3-preview + ConforNets), together with sectors, potential tools/workflows, and key assumptions/dependencies.
- Healthcare/Pharma — State-specific ensemble docking and virtual screening
- What: Generate active/inactive/cryptic-pocket conformers (e.g., GPCR active, kinase DFG-out, transporter outward-open) to enable ensemble docking, hit finding, and ranking for orthosteric and allosteric modulators.
- Tools/workflow: OF3p + ConforNets to produce state-specific ensembles → pocket detection (Fpocket, SiteMap) → ligand preparation (RDKit) → docking (Smina, Glide, GOLD) → rescoring/FEP (Schrödinger FEP+, OpenMM).
- Assumptions/dependencies: Sufficient MSA depth; the desired state is encoded in AF3/OF3p latents; structures are not energetically calibrated (post-docking minimization/relaxation needed); cryptic pockets may require additional filtering for physicochemical plausibility.
- Structural Biology — Model fitting and hypothesis generation for cryo-EM/crystallography
- What: Rapidly propose alternative conformers that fit ambiguous density or represent minor states; triage which state-specific constructs/mutants to pursue experimentally.
- Tools/workflow: Generate conformers with ConforNets → rigid/local fit into maps (ChimeraX, Phenix, ISOLDE) → refine and validate.
- Assumptions/dependencies: Map resolution and quality; ConforNet conformers may need restraints/refinement; no explicit ligand/ion cofactors unless modeled downstream.
- Computational Biophysics — Seeding diverse MD simulations
- What: Provide physically plausible, diverse starting structures to accelerate barrier crossing and improve coverage of conformational space.
- Tools/workflow: ConforNet ensembles → restraint/minimization → MD (GROMACS, OpenMM) → enhanced sampling (PLUMED, metadynamics) → MSM/kinetic analysis.
- Assumptions/dependencies: ConforNets are uncalibrated; structures should be relaxed; force-field choices and solvent conditions matter.
- Drug Discovery — Cryptic pocket discovery and pharmacophore modeling
- What: Reveal apo/holo cryptic pockets to build state-specific pharmacophores and prioritize fragment screening.
- Tools/workflow: Generate holo-like conformers → pocket definition (VolSite, DoGSite) → pharmacophore extraction (LigandScout) → screen libraries (OpenEye, Pharmit).
- Assumptions/dependencies: Pocket stability/usefulness must be verified; absence of ligand may bias pocket geometry; follow-up MD or experimental validation recommended.
- Target Assessment — “Reachability” triage of therapeutic states
- What: Use success@N criteria to assess whether a difficult functional state (e.g., DFG-out) is reachable for a target; prioritize targets with favorable state accessibility for downstream programs.
- Tools/workflow: Batch ConforNet sampling → compute RMSD-to-state → rank targets by success rate and proximity.
- Assumptions/dependencies: Benchmarks suggest variable difficulty (e.g., kinase loops harder); relies on having an exemplar/source state for “transfer.”
- Protein Engineering — Rationale for mutations to stabilize desired states
- What: Compare ConforNet-induced and baseline conformers to identify residues to mutate (e.g., disulfide locks, helix capping, cavity-filling) to stabilize target states for assays or crystallization.
- Tools/workflow: Structural comparison (PyMOL, ChimeraX) → mutational hotspot analysis → in silico ΔΔG (Rosetta cartesian_ddg, FoldX) → experimental design.
- Assumptions/dependencies: ConforNet conformers guide hypotheses rather than guarantee stability; mutations must be validated experimentally.
- Academic Research — Mechanistic hypothesis generation and teaching
- What: Visualize large-scale motions (hinges, gate opening/closing), fold switching, and family-conserved rearrangements to generate testable hypotheses and instructional material.
- Tools/workflow: ConforNet sampling → trajectory-like visualizations → mapping to functional motifs.
- Assumptions/dependencies: Interpret with caution in absence of energetic calibration and experimental corroboration.
- Software/Platform Engineering — Conformational control plugin for AF3/OF3p pipelines
- What: Package ConforNets as reusable, family/state-specific adapters (e.g., “GPCR-active”, “Kinase-DFG-out”) that can be applied to new sequences within a family.
- Tools/workflow: Python SDK/CLI for ConforNet inference; integration with ColabFold-like interfaces; batch job orchestration on GPUs.
- Assumptions/dependencies: Availability of OF3p weights and compute; governance for distributing family-specific ConforNet parameters.
- Quality Control of Structure Predictions — Exploring ambiguity and alternates
- What: Systematically sample alternate states to flag conformational ambiguity and possible fold switching, informing caution in downstream use.
- Tools/workflow: ConforNet ensembles → clustering → confidence/violation checks → reporting alternates.
- Assumptions/dependencies: Interpretation requires domain expertise; alternate states may include artifacts.
- Education/Training — Interactive demonstrations of conformational control
- What: Course labs and tutorials showing how latent perturbations induce functional states across families (GPCRs, kinases, transporters).
- Tools/workflow: Notebooks and web apps tying OF3p + ConforNets to basic analysis and visualization.
- Assumptions/dependencies: GPU access for live demos or precomputed examples.
Long-Term Applications
The following use cases require further methodological development, validation, calibration, or scaling before broad deployment.
- Healthcare/Pharma — Calibrated, state-weighted ensembles for free-energy workflows
- What: Combine ConforNet-driven diversity with MD/reweighting to approximate Boltzmann-weighted state populations; feed into FEP and kinetics predictions.
- Tools/workflow: ConforNet ensembles → short MD + enhanced sampling → reweight to ensembles → free-energy pipelines.
- Assumptions/dependencies: Requires robust calibration against experiment/MD; careful bias correction; extensive benchmarking.
- Protein–Ligand Complex Prediction — State- and ligand-conditioned co-prediction
- What: Extend conformational control to jointly predict ligand-bound structures (e.g., learning ligand-guided latent transforms or integrating diffusion guidance with ligand constraints).
- Tools/workflow: ConforNets + diffusion guidance conditioned on ligand/fragment; iterative refinement with docking and restraint-based fitting.
- Assumptions/dependencies: New training and conditioning mechanisms; datasets with paired state/ligand complexes; managing ligand chemistry diversity.
- Family-wide ConforNet libraries and state ontologies
- What: Curate and distribute standardized ConforNets for major protein families (GPCR active, kinase DFG-out variants, transporter conformational cycle states), with metadata and usage guidelines.
- Tools/workflow: Repository of vetted transforms; programmatic API; provenance and benchmark scores per family/state.
- Assumptions/dependencies: Sustained curation; versioning and governance; resolving IP/licensing around base models and weights.
- Integrating experimental constraints (NMR, HDX-MS, crosslinks, cryo-ET)
- What: Train or guide ConforNets using experimental observables to induce conformers consistent with measured restraints, enabling structure refinement and mechanistic inference.
- Tools/workflow: Losses or priors from NOEs/RDCs/HDX-MS/CXL; cryo-ET density restraints; hybrid modeling.
- Assumptions/dependencies: Data availability/quality; robust multi-modal loss design; validation on prospective datasets.
- Multimeric complexes and assemblies — State control across interfaces
- What: Generalize conformational control to oligomers and transient complexes (e.g., GPCR–G protein, kinases with regulatory partners, transporters with periplasmic binding proteins).
- Tools/workflow: OF3p multimer modes + ConforNets at appropriate latent sites; interface-aware objectives.
- Assumptions/dependencies: Architectural extensions and retraining may be required; larger compute; benchmarking datasets for complexes.
- Condition-dependent conformational control (pH, ions, PTMs)
- What: Make conformational control conditional on biochemical context (protonation state, phosphorylation, ion binding), enabling realistic state induction under specific conditions.
- Tools/workflow: Condition labels/embeddings; protonation/PTM-aware pipelines; ion placement protocols.
- Assumptions/dependencies: Requires labeled datasets and conditioning mechanisms; accurate modeling of chemical context.
- Proteome-scale mapping of conformational landscapes
- What: High-throughput screening to annotate state reachability per protein/family; support target selection and systems-level analyses of conformational regulation.
- Tools/workflow: Cloud-native ConforNet inference at scale; automated QC; public database with state coverage metrics.
- Assumptions/dependencies: Substantial compute; standardized metrics; community curation and adoption.
- Allosteric drug design and selectivity engineering
- What: Use state transfer and cryptic pocket induction to systematically reveal allosteric sites and design selective ligands (e.g., kinase family selectivity via state-specific features).
- Tools/workflow: ConforNets → pocket comparison across family → selectivity filters → medicinal chemistry loops.
- Assumptions/dependencies: Requires robust cross-family transfer; careful assessment of pocket conservation and druggability.
- Synthetic Biology/Protein Engineering — Designing switchable proteins
- What: Leverage controllable conformational states to design switchable enzymes/sensors (e.g., conformational toggles coupled to signals or ligands).
- Tools/workflow: ConforNet-guided state mapping → mutational design to stabilize toggles (Rosetta/ProteinMPNN) → experimental screening.
- Assumptions/dependencies: Complex coupling between conformational equilibria and function; extensive experimental cycles.
- Closed-loop discovery with lab automation
- What: Integrate ConforNet proposals with robotic screening for ligands/mutants that stabilize desired states; iteratively update models/ConforNets.
- Tools/workflow: Active learning over conformers and designs; automated biophysics assays; ML-in-the-loop optimization.
- Assumptions/dependencies: Significant infrastructure; robust feedback signals; data/knowledge management.
- Standards and policy for computational evidence in regulatory contexts
- What: Develop best practices for using state-specific structural predictions in regulatory submissions (e.g., mechanism-of-action rationale, off-target risk via alternate states).
- Tools/workflow: Community guidelines; audit trails; uncertainty reporting; benchmark-based validation reports.
- Assumptions/dependencies: Regulatory engagement; consensus on validation thresholds; transparency of methods and datasets.
Cross-cutting assumptions and dependencies
- Methodological scope: ConforNets modulate AF3/OF3p latents and are not energy-calibrated; generated ensembles require downstream physics-based filtering/validation.
- Data prerequisites: MSA depth/diversity matters; transfer is strongest within structural families; a suitable source protein/state is needed for supervised transfer.
- Generalization: Large flexible loops (e.g., kinase A-loops) remain challenging; performance varies across families and targets.
- Compute: GPU access is required; training ConforNets is lightweight relative to model retraining but non-zero; scaling to proteome or multimeric assemblies increases costs.
- Toolchain integration: Best results arise when ConforNet conformers feed into established modeling, docking, and MD refinement pipelines.
- Validation: Experimental confirmation (biophysics, structures, functional assays) remains essential before high-stakes decisions (e.g., clinical programs).
Glossary
- Activation loop (A-loop): A flexible kinase segment whose conformation regulates activity. Example: "Kinases transition between active (DFG-in) and inactive (DFG-out) states by locally rearranging their activation loop and DFG motif."
- AF3 (AlphaFold3): The third-generation AlphaFold architecture for structure prediction, here referring to its open-source reproduction OpenFold3-preview (OF3p). Example: "we investigate both the optimal location and manner-of-operation for perturbing latent representations in the AF3 architecture."
- Affine transform: A linear transformation with optional translation applied to features; here used channel-wise on latents. Example: "ConforNets: channel-wise affine transforms of the pre-Pairformer pair latents."
- Apo state: A protein conformation without bound ligand, often hiding “cryptic” pockets. Example: "apo vs.\ holo pockets"
- Boltzmann-weighted ensemble: A set of conformations sampled in proportion to their thermodynamic probabilities. Example: "producing calibrated, Boltzmann-weighted ensembles."
- Conformational heterogeneity: The existence of multiple biologically relevant protein conformations. Example: "struggle to capture the conformational heterogeneity and context-dependent changes"
- Conformational transfer: Inducing a learned conformational state from a source protein in other family members. Example: "On the novel supervised task of conformational transfer"
- Conditioning signal: The inputs provided to the diffusion model that guide generation, as opposed to altering the model’s score function. Example: "ConforNets alter the conditioning signal, not the score function"
- Cryptic pocket: A ligand-binding site that is not apparent in the apo structure and becomes accessible upon conformational change. Example: "Cryptic pockets (N=34): apo vs.\ holo pockets;"
- Denoiser: The network component in diffusion models that predicts the clean structure from noisy inputs. Example: "The denoiser predicts a clean structure from a noisy input at a given ,"
- DFG-in/DFG-out: Two kinase states defined by the orientation of the Asp-Phe-Gly motif, associated with active/inactive conformations. Example: "DFG-in/out transition of kinases"
- Diffusion guidance: Steering a diffusion process during inference to bias samples toward desired properties. Example: "ConforMix uses diffusion guidance to bias AF3 towards states with fixed target RMSDs"
- Distogram: A probabilistic representation (logits/probabilities) over pairwise inter-residue distance bins. Example: "From $\mathbf{z}^{\mathrm{post}$, OF3p also predicts distogram logits"
- Distogram entropy: A measure of uncertainty in the distance distribution; maximizing it can promote diversity. Example: "we maximize the entropy of the predicted distogram"
- Elucidated Diffusion Model (EDM): A continuous-time diffusion parameterization used for generative modeling. Example: "OF3p adopts the Elucidated Diffusion Model (EDM) parameterization"
- Euler–Maruyama-like solver: A numerical scheme for simulating stochastic differential equations during diffusion sampling. Example: "an Euler-Maruyama-like solver with 200 discretizations"
- Evoformer: The AF2/AF-derived module that processes sequence/MSA features to produce structure-relevant embeddings. Example: "conditioned on fixed AF2 Evoformer embeddings"
- Flow matching: A generative approach that learns continuous flows mapping noise to data, used here for protein backbones. Example: "as a sequence-conditioned flow-matching model"
- Free energy landscape: The thermodynamic surface governing conformational preferences and transitions of proteins. Example: "learning the free energy landscapes of proteins"
- GPCR (G protein–coupled receptor): A membrane protein family undergoing conserved activation-associated conformational changes. Example: "GPCR activation couples agonist binding to G-protein recruitment by rearranging transmembrane helices (TM) 5--7"
- Hinge motion: A large-scale domain movement around a flexible “hinge” region. Example: "Domain motions (N=21): large-scale hinge motions."
- Holo state: A ligand-bound protein conformation exposing or reshaping binding pockets. Example: "apo vs.\ holo pockets"
- Multiple sequence alignment (MSA): An alignment of homologous sequences used by AF models to infer structural couplings. Example: "subsamples input multiple sequence alignments (MSAs)"
- Out-of-distribution (OOD): Data that differs from the training distribution; here a benchmark of newer structures. Example: "out-of-distribution for BioEmu (but not necessarily for other methods)"
- Pairformer: The AF3 trunk that refines single and pair representations of residues with specialized attention operations. Example: "The Pairformer then refines the two representations via triangular updates and self-attention"
- Pair latents (pair representation): Learned pairwise residue–residue features used by AF3 to reason about contacts and geometry. Example: "pre-Pairformer pair latents"
- Recycling (Pairformer passes): Iteratively reprocessing representations through the trunk to refine predictions. Example: "OF3p refines single and pair representations through multiple recycles of Pairformer passes"
- Rigid alignment: Superposition of structures via rotation/translation before comparing coordinates. Example: "After rigid alignment, we maximize deviation in coordinate space:"
- SE(3)-equivariant: Neural network property ensuring outputs transform consistently under 3D rotations/translations. Example: "an SE(3)-equivariant diffusion model"
- Triangular updates: Specialized attention operations in AF models for pairwise information flow among residues. Example: "via triangular updates and self-attention"
- US-align: A structural alignment tool used to compare protein conformations. Example: "using US-align"
- Variance-exploding diffusion: A diffusion process where noise variance increases over time, used in EDM. Example: "continuous-time variance-exploding diffusion."
- pLDDT: AlphaFold’s per-residue confidence metric used to assess prediction reliability. Example: "to maximize the model's predicted confidence (pLDDT)"
Collections
Sign up for free to add this paper to one or more collections.