- The paper demonstrates a multi-stage validation framework that enhances clinical extraction reliability using LLMs with minimal annotation.
- It details pre-extraction prompt calibration, semantic grounding with cosine similarity, and a judge LLM for confirmatory validation.
- Outcome-based predictive analysis confirms that LLM-extracted features can outperform traditional ICD-10 codes in predicting clinical events.
Introduction
The proliferation of LLMs has led to transformative advances in clinical NLP. However, their operational deployment for extracting diagnoses such as substance use disorders (SUDs) from large-scale electronic health records (EHRs) remains circumscribed by limited trustworthiness and the absence of scalable, annotation-efficient validation. The paper "A Multi-Stage Validation Framework for Trustworthy Large-scale Clinical Information Extraction using LLMs" (2604.06028) addresses this core challenge with a comprehensive framework designed to assess and enhance the reliability of LLM-extracted clinical information, leveraging complementary validation signals under weak supervision and minimal manual annotation.
Multi-Stage Validation Framework
The authors propose a modular, human-in-the-loop validation pipeline, encapsulating both pre-extraction and post-extraction procedures.
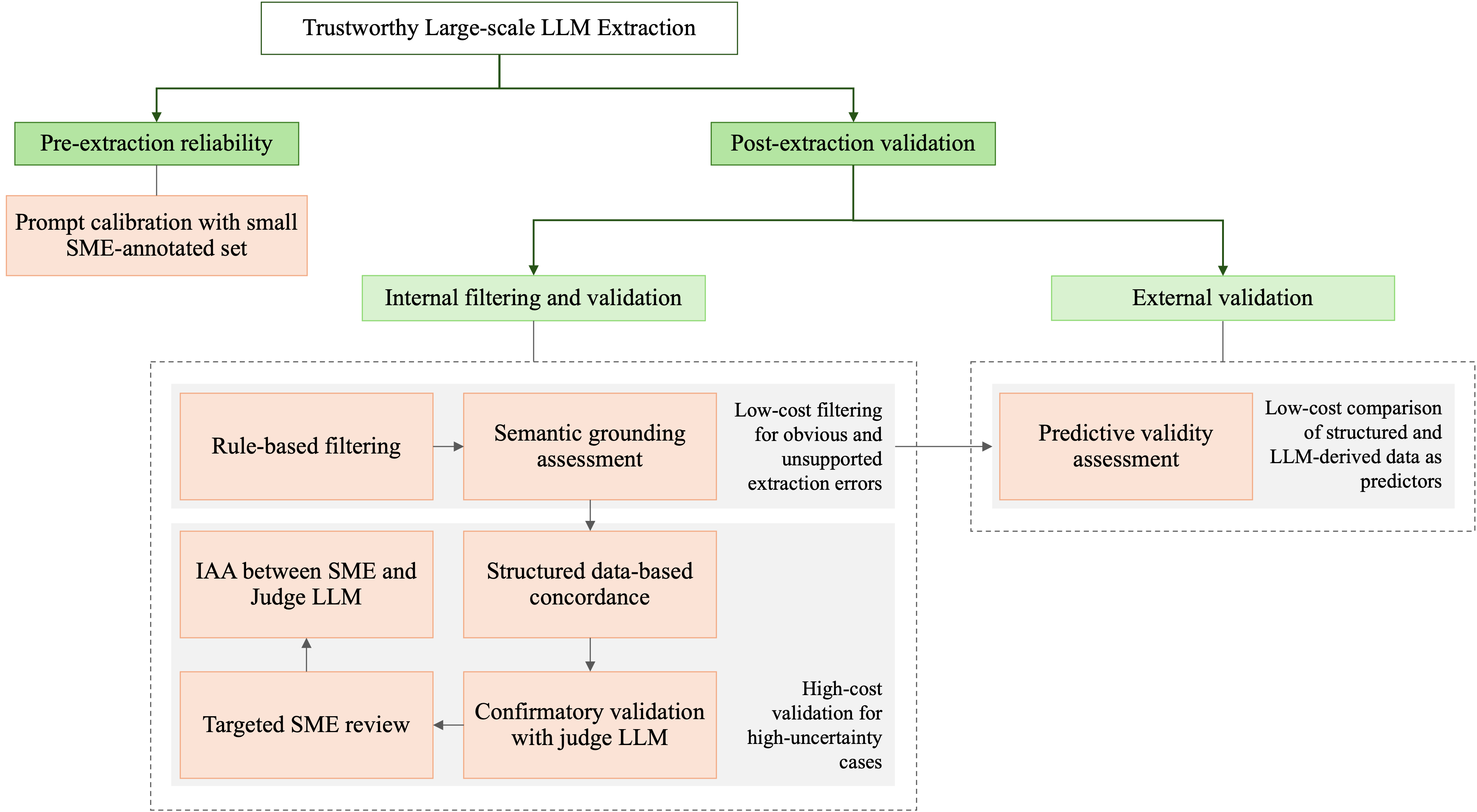
Figure 1: A multi-stage framework for trustworthy large-scale LLM extraction of clinical information.
Pre-extraction reliability centers on prompt calibration, using a limited set of annotated notes to optimize extraction quality and reduce susceptibility to propagation of prompt-induced errors. Post-extraction validation is decomposed into a series of cascading filters and assessments: rule-based plausibility filtering, semantic grounding via embedding similarity, confirmatory evaluation by a larger judge LLM, selective SME review, and outcome-driven predictive validity analysis. This structure ensures that higher-cost human and model resources are allocated only to high-uncertainty cases.
Prompt Engineering and Calibration
Extraction performance is contingent on prompt quality. The framework extensively explores zero-shot, one-shot, and two-shot prompting, both in direct and chain-of-thought (CoT) configurations.
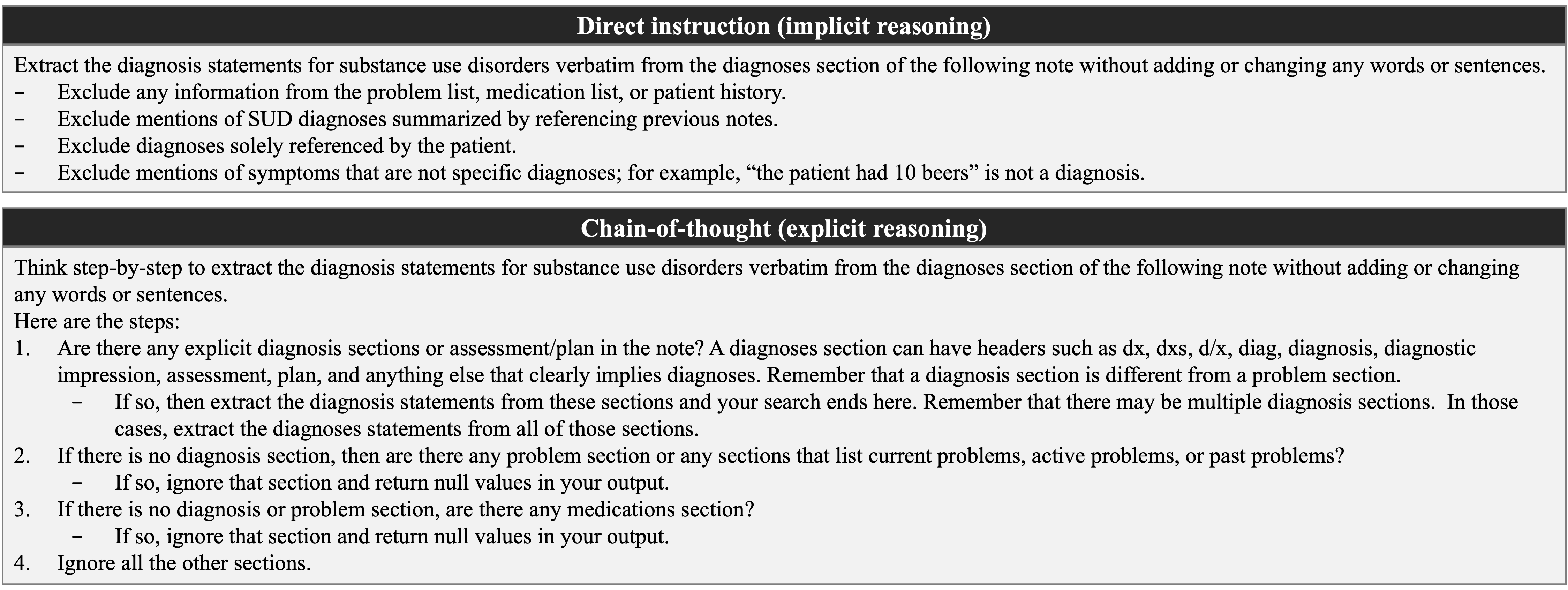

Figure 2: Direct vs chain-of-thought prompting to extract SUD diagnoses information from clinical notes.
Numerical results indicate that zero-shot prompting yields higher F1 scores than few-shot strategies, likely due to the avoidance of distributional bias from in-context examples. Further, chain-of-thought reasoning confers an additional performance gain (F1strict=0.8582, F1relaxed=0.9004), attributed to explicit decomposition of clinical reasoning steps, which is essential for capturing the nuanced language of SUD diagnoses. The inference model is Llama-3.1-8B-Instruct.
Post-Extraction Filtering and Semantic Grounding
Immediately following extraction, lightweight rule-based filtering and semantic grounding are applied. Rule-based checks employ curated inclusion/exclusion patterns to flag unsupported or irrelevant model outputs. Subsequently, semantic grounding compares extraction embeddings to sliding windows over the source note, flagging outputs without clear textual support via cosine similarity metrics. A threshold of 0.65 was empirically selected for grounding. The joint effect of these steps eliminates 14.59% of LLM-positive extractions, demonstrating substantial mitigation of spurious or hallucinated outputs, with high per-category precision.
Confirmatory Validation with Judge LLM and SME Review
For extractions that are positive but unsupported by structured data (i.e., lacking ICD-10 codes), a higher-capacity judge LLM (gpt-oss-20b) independently verifies the extraction using an explicit adjudication prompt.

Figure 3: Prompt for judge LLM.
On a set of 80,006 high-uncertainty notes, the primary LLM achieves F1relaxed=0.7962 and F1strict=0.7735 when benchmarked against judge-adjusted references, with most SUD subtypes exceeding an F1 of 0.78 under relaxed matching. Targeted SME review on random sub-samples validates the judge LLM’s reliability, yielding high inter-annotator agreement (Gwet’s AC1 = 0.91 overall, and 0.80 for high-uncertainty, LLM+/ICD- cases), which are robust levels for clinical tasks with skewed label distributions. These results substantiate the use of a model-based judge as a scalable surrogate to expert review for routine validation while reserving SME time for edge cases or framework calibration.
Predictive Validity as External Validation
A critical component of the framework is outcome-anchored predictive validity analysis. The extracted SUD diagnoses are used as features to predict subsequent SUD-specialty care engagement, compared to predictions made using structured ICD-10 codes and combinations thereof, across multiple patient cohorts.
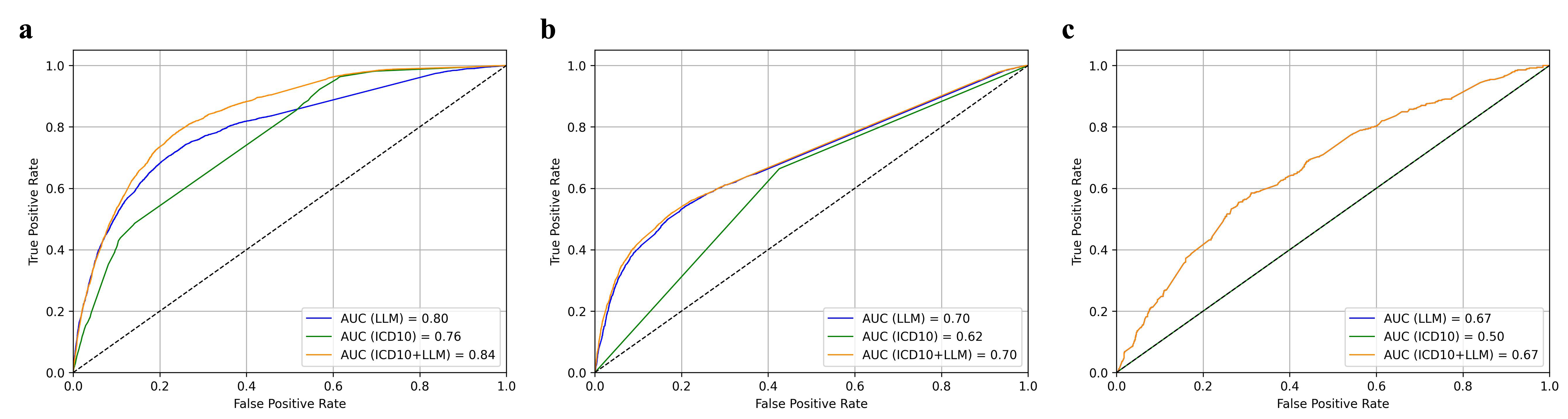
Figure 4: Predictive validity of LLM-extracted SUD diagnoses shown by ROC curves comparing outcome prediction using LLM-extracted, ICD-10–based, and combined features.
The LLM-extracted features robustly outperform structured-data baselines: in the full study population, area under the ROC curve (AUC) values are 0.80 (LLM), 0.76 (ICD-10), and 0.84 (combined). In both concordant and narrative-only cohorts, LLM features maintain or exceed structured data predictive performance. Importantly, even in narrative-only (structured-code negative) cases, LLM-extracted diagnoses retain discriminative power (AUC = 0.67), revealing clinical signal absent from codified EHR data and corroborating the clinical utility of narrative extraction.
Discussion and Implications
The framework integrates and extends prior work on LLM validation, prompt optimization, semantic assessment, LLM-as-a-judge, and outcome-based validation. Each stage targets distinct failure modes: structural implausibility, semantic hallucination, and mismapped or ambiguous diagnoses. The strategic sequencing allows precision to be maximized where clinical risk is greatest (i.e., unsupported label insertions) while preserving recall elsewhere. The judge LLM’s high reliability as an adjudicator is particularly salient, sidestepping the bottleneck of annotation-intensive SME review. Outcome-based validation unambiguously connects extracted clinical signals to future clinical events, offering evidence of construct validity that is otherwise unavailable at scale.
Practically, these findings reinforce the feasibility of deploying general and domain-adapted LLMs for large-scale, trustworthy diagnosis extraction in the presence of weak or incomplete ground truth. This has direct implications for operationalizing LLM-powered clinical decision support and population health analytics, especially in domains where informative content is systematically missing from structured fields. The approach is model-agnostic, requiring only minor adaptation for task or institution-specific documentation practices.
Future Directions
The proposed multi-stage pipeline is extensible to other clinical entities, coding systems, and even broader information extraction tasks subject to similar weak-supervision realities. Further research could formally characterize learned uncertainty measures at each stage, automate adaptation to new documentation styles, or integrate federated SME feedback for continuous improvement. Additionally, real-time framework adaptation in evolving EHR ecosystems and robust quantification of the trade-offs between computational efficiency and human oversight merit further study.
Conclusion
A robust, weakly supervised, multi-stage framework for large-scale, trustworthy extraction of clinical information using LLMs is feasible and effective. Human-in-the-loop strategies focused on calibration, automated semantic assessment, adjudication, and predictive validity can systematically manage risk, minimize annotation burden, and maximize the clinical relevance of extracted information. These results provide a foundation for deploying LLMs as reliable clinical information extractors in high-stakes environments, where annotation- or ground-truth limitations have previously limited progress.



