- The paper introduces MolDA, a diffusion-driven framework that replaces autoregressive generation with iterative denoising to ensure global molecular validity.
- It combines a hybrid graph encoder with a Q-Former cross-modal projector, achieving competitive benchmarks such as an AUROC of 0.846 on the SIDER side-effect dataset.
- It employs multi-stage training with domain-adaptive tokenization and MolPO optimization to enforce chemical constraints and address modality imbalance.
MolDA: Molecular Understanding and Generation via Large Language Diffusion Model
Motivation and Context
Molecular LLMs (MolLMs) have led to significant advances in computational chemistry and drug discovery. Yet, mainstream approaches predominantly leverage autoregressive (AR) generation, typically representing molecules as 1D strings (e.g., SMILES, SELFIES) and relying on left-to-right decoding. This methodology, while effective for language tasks, is fundamentally sub-optimal for molecular applications due to its inability to enforce non-local, global chemical constraints such as ring closure and proper atom valence. AR models readily accumulate errors in early decoding steps, leading to globally invalid or incoherent molecular structures.
Recent multimodal frameworks have attempted to alleviate sequential representation limitations by integrating graph neural network (GNN) encoders, aligning molecular graph features with LLM token spaces via cross-modal projectors. However, these architectures still retain AR backbones, inheriting their inductive biases and structural limitations. Discrete diffusion LLMs (DLM) have recently emerged as an effective alternative for text by reframing generation as iterative denoising of fully masked sequences, enabling global context modeling and token revision. Their extension into holistic molecular understanding is, however, underexplored.
Model Architecture
MolDA proposes a fundamentally non-AR paradigm for molecular language tasks by constructing a fully multimodal system grounded in the discrete diffusion generation process. The architecture is structured as follows:
- Hybrid Graph Encoder: A parallel architecture combining GINE for neighborhood interactions and TokenGT transformers for global topology processing, producing rich local and long-range graph embeddings.
- Q-Former Cross-Modal Projector: A query-based cross-modal alignment block that compresses hybrid graph embeddings into language-model-aligned token features using learnable queries via multi-head cross-attention, efficiently bridging molecular graphs and LLM token spaces.
- Diffusion-Driven LLM Backbone: The core generation mechanism is LLaDA-8B-Instruct, a DLM configured for molecular instruction finetuning, operating through iterative denoising (discrete masked diffusion) instead of AR next-token prediction. This allows modeling of joint sequence probabilities, providing bidirectional context and correction capabilities at each generation step.
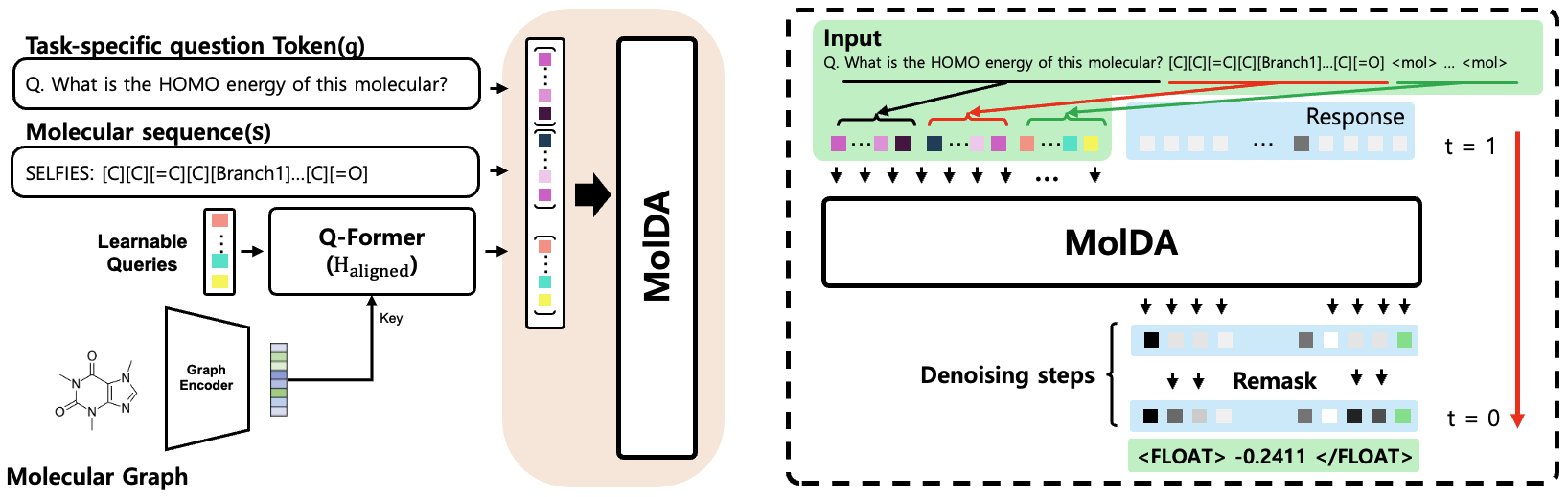
Figure 1: MolDA architecture overview; a hybrid graph encoder for local/global features, a Q-Former for cross-modal projection, and an LLM backbone performing iterative denoising for robust generation.
Training and Optimization
MolDA employs a robust, multi-stage training strategy:
- Domain-Adaptive Tokenization: Extends the tokenizer with nearly 3,000 SELFIES-specific tokens, initialized to preserve embedding statistics, directly addressing vocabulary inadequacies found in pre-trained LLMs for the chemistry domain.
- Hybrid Encoder Pretraining: The graph encoder is initially trained using functional group prediction (binary cross-entropy objective for chemical substructures) and SELFIES reconstruction (via AR GPT-2 decoder), ensuring the extraction of both local and global chemical semantics.
- Textual DLM Initialization: The LLaDA backbone is subjected to supervised finetuning (SFT) using mask-based denoising objectives on instruction datasets, without graph inputs, injecting domain-relevant language priors.
- Cross-Modal Alignment: The Q-Former is then trained (with backbone and GNN frozen) to project graph embeddings into token space, using alignment losses derived from the masked diffusion objective.
- Molecular Structure Preference Optimization (MolPO): A central innovation, MolDA mathematically adapts MolPO for the diffusion context by defining reward margins in terms of masked diffusion log-likelihoods, computed between embeddings derived from the reference and perturbed graphs. This scheme actively penalizes reliance on textual information alone and enforces meaningful graph feature utilization, addressing the classic "modality imbalance" in multimodal molecular learning.
Denoising and Generation Dynamics
During inference, MolDA utilizes iterative masked denoising with task-adaptive sampling:
- Captioning/Reasoning: Block diffusion with low-confidence remasking for natural language tasks, where tokens with the highest uncertainty are selectively revised, facilitating strong bidirectional modeling.
- Molecule Generation: Full-sequence pure diffusion, essential when molecular validity depends on simultaneous satisfaction of multiple non-local atomic constraints, such as in the realization of aromaticity or ring closure.
This process ensures global structural consistency and validity, providing an advantage over unidirectional AR approaches for molecular structure-critical tasks.
Empirical Results
MolDA's performance is evaluated across major molecular understanding, property prediction, and reaction prediction benchmarks:
- Molecular Understanding: While MolDA’s generation and captioning metrics on ChEBI-20 lag behind top AR baselines, it demonstrates competitive or leading performance in property-centric tasks. Notably, MolDA achieves the highest AUROC (0.846) on the SIDER side-effect benchmark, indicating robust efficacy in structure-sensitive predictions, and is highly competitive on the HIV and HOMO datasets.
- Reaction Prediction: MolDA attains the second-highest Exact Match and MACCS scores across forward synthesis, retrosynthesis, and reagent prediction on Mol-Instructions—dramatically outperforming generalist AR LLMs in tasks where global molecular transformations are critical.
- Denoising Steps: An ablation demonstrates that increasing the number of denoising steps (T) improves exactness and MACCS up to T=64, after which returns diminish; generation efficacy thus benefits from controlled iterative revision rather than shallow denoising.
These results collectively demonstrate the value of discrete diffusion in complex chemical tasks—particularly when modeling structurally constrained transformations where AR models’ locality and error accumulation are problematic.
Theoretical and Practical Implications
The modeling choices in MolDA have direct implications:
- Theoretical: The migration from AR to diffusion-based generation in molecular language modeling redefines inductive biases, favoring models that enforce global consistency over those that optimize for local fluency. Preference optimization in a diffusion backbone may inspire similar practices in other multimodal domains suffering from modality imbalance.
- Practical: MolDA enables more reliable molecular generation and interpretation for tasks where chemical validity is non-negotiable, such as inverse design, retrosynthesis, and reaction prediction. The design is inherently scalable to additional modalities or instruction sets.
- Limitations: Despite advances, diffusion-based generation (particularly with DLMs) still trails AR approaches in tasks demanding maximum fluency in natural language, and its molecule generation on synthetic benchmarks remains to be improved.
Future Directions
Potential avenues of exploration include:
- Architectural Refinements: Enhanced hybridization strategies for graph encoders and more expressive cross-modal projector mechanisms.
- Task-Specific Sampling: Adaptive denoising policies driven by explicit chemical constraints or property predictors.
- Scalability and Efficiency: Compression of denoising steps via learned step-size adaptivity, improving computational efficiency.
Furthermore, broader integration of domain-specific constraints—e.g., quantum chemistry, bioactivity priors—could further harness the flexibility of diffusion backbones in molecular LLMs.
Conclusion
MolDA establishes a principled framework for moving beyond the autoregressive paradigm in molecular language modeling by harnessing discrete masked diffusion. Through bidirectional generation, advanced preference optimization, and tailored cross-modal alignment, MolDA provides competitive property prediction and robust reaction modeling, underscoring the viability of DLMs for global structure-enforcing molecular AI. This direction is expected to inform subsequent generations of multimodal, graph-aware, and structure-coherent LLMs for chemical sciences.
