- The paper introduces BioMol-LLM-Bench, a framework that evaluates 13 LLMs across 26 tasks spanning four levels of molecular complexity.
- The paper finds that while high-parameter models excel in classification tasks, they often struggle with regression and output validity on complex molecular tasks.
- The paper shows that tool integration significantly enhances performance, underscoring the need for hybrid intelligence in precise biomolecular modeling.
Cross-Scale Evaluation of LLMs for Bio-Molecular Modeling
Introduction and Motivation
Bio-molecular modeling across multiple scales presents substantial challenges, demanding accurate reasoning about small molecules, proteins, and their interactions. The proliferation of LLMs, including both generalist and domain-specialized architectures, has driven rapid adoption of these models for tasks such as molecular property prediction, protein annotation, and biomolecular interaction inference. However, a systematic, fine-grained evaluation of LLMs covering the entire landscape of molecular complexity has been lacking. "The limits of bio-molecular modeling with LLMs: a cross-scale evaluation" (2604.03361) introduces BioMol-LLM-Bench, a benchmarking framework that integrates 26 tasks at 4 hierarchical levels of molecular complexity, and enables rigorous, tool-augmented assessment of 13 diverse LLMs. The work addresses critical gaps in evaluation methodology, architectural analysis, and mechanistic insights for LLMs in molecular science.
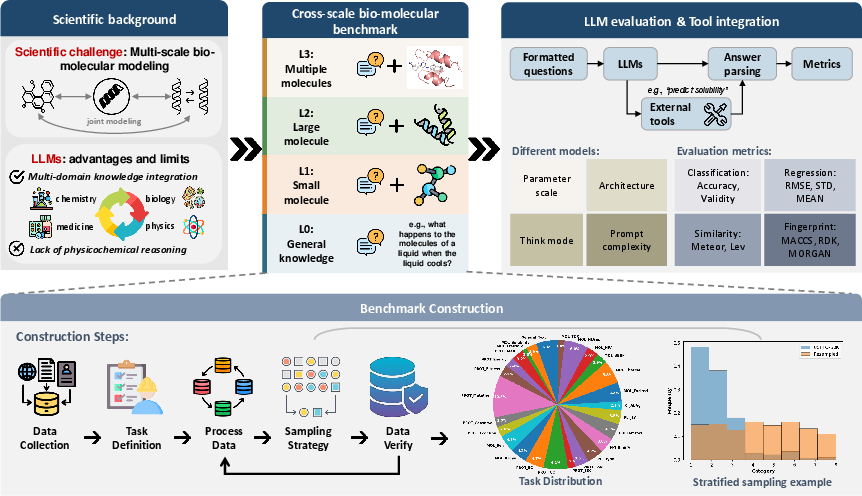
Figure 1: Overview of BioMol-LLM-Bench, showing the cross-scale evaluation pipeline and tool integration for benchmarking 13 LLMs across 26 tasks.
Benchmark Design and Methodology
BioMol-LLM-Bench is constructed to provide comprehensive coverage across bio-molecular text understanding (L0), small molecule property and reaction tasks (L1), protein property and function prediction (L2), and multi-entity molecular interaction problems (L3). Data is curated through deduplication, strict structural validation, and LLM-based filtering for bio-molecular relevance. A notable feature is the integration of bio-chemistry computational tools within the benchmark pipeline, enabling not only direct evaluation of LLM outputs, but also assessment of the agentic and tool-using capabilities of the models. The evaluation protocol extends beyond accuracy, explicitly quantifying output validity by assessing consistency with domain-required answer formats.
Analysis of 13 LLMs on 8 bio-molecular classification tasks reveals that high-parameter models (e.g., DeepSeek-v3.1, Qwen3-235b-a22b-2507) are consistently among the top performers in terms of accuracy. However, model performance is not uniform; for instance, models fine-tuned for specific domains, such as NatureLM-8x7B, may fail to generalize across tasks and even attain zero accuracy on tasks outside their training distribution. Output validity remains largely high, but there exist cases, notably with GPT-oss-20B and Phi-4-14B, where high-accuracy predictions do not always conform to required formats, highlighting an orthogonality between memorization-based correctness and task-constrained generation.
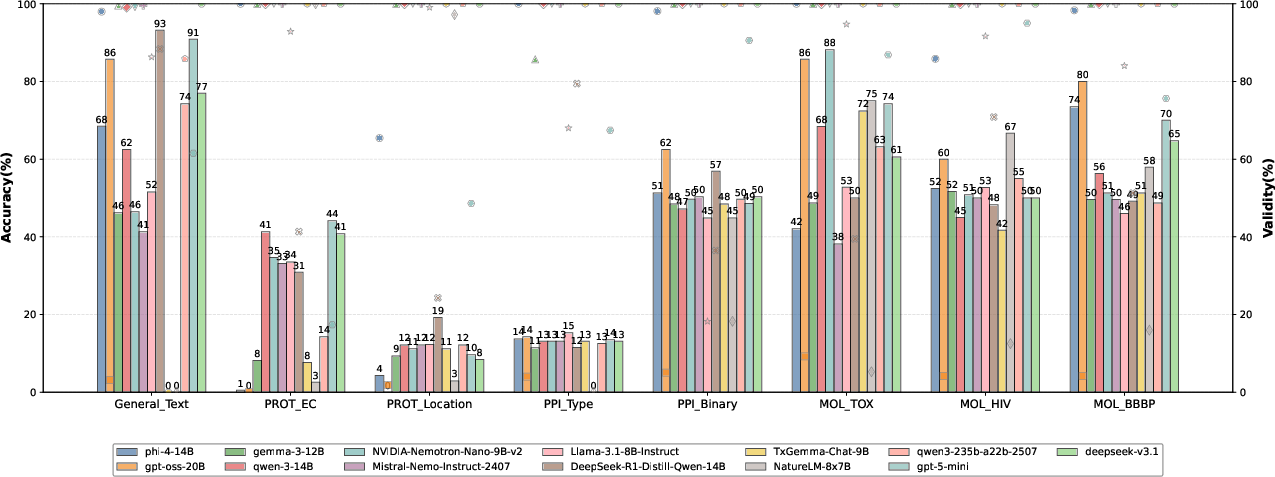
Figure 2: LLM classification task accuracy and output validity indicate that scale correlates with accuracy, but validity violations are non-negligible for some models.
Regression Task Limitations
Regression tasks, particularly those involving molecular thermodynamic or protein-level mutation and stability effects, expose the pronounced limitations of both small and large LLMs. Models such as NVIDIA-Nemotron-Nano-9B-v2 achieve lower numerical errors and variance on average, yet even the best models fail to consistently solve inherently complex or long-range tasks. There is no monotonic improvement with increased model size, especially for L1-level tasks—implicating fundamental limits in purely language-based estimation of physical quantities.
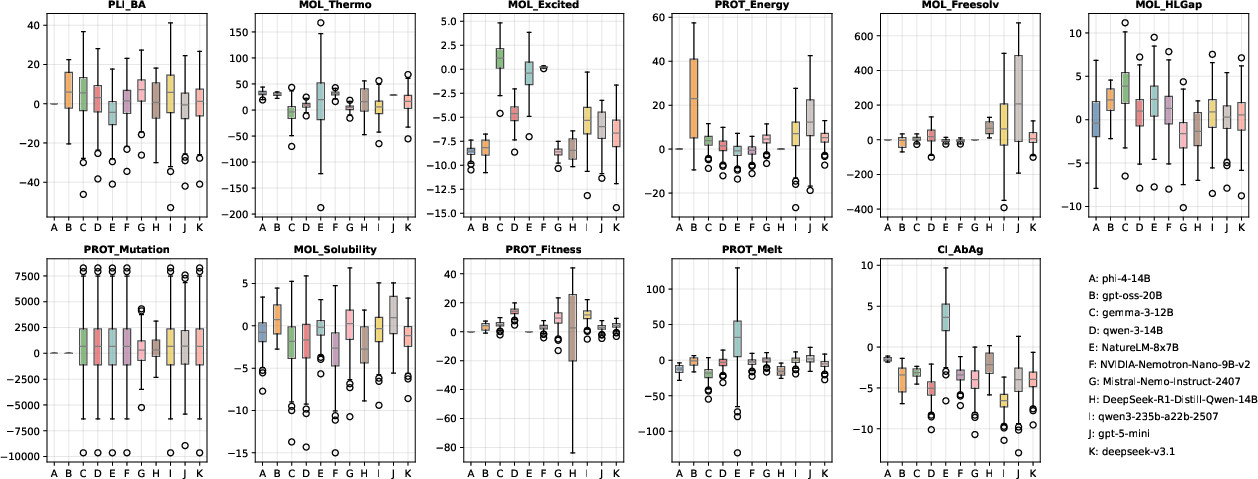
Figure 3: Distribution of numerical regression errors across models and tasks; challenging targets (especially MOL_Thermo and PROT_Mutation) reveal model inadequacy.
Generative Bio-molecular Task Evaluation
In generative tasks, such as forward and retrosynthetic prediction or protein functional annotation, most models deliver only moderate performance, with some (e.g., TxGemma-Chat-9B) showing improved fidelity due to domain-specific fine-tuning, as reflected in molecular fingerprint similarity metrics and string similarity scores. Nevertheless, generative failures are frequent, with several models unable to produce valid structures or outputs at all for certain tasks. CoT-based fine-tuning on these tasks does not result in universal gains; for example, DeepSeek-R1-Distill-Qwen-14B was shown to have degraded generation compared to its non-CoT baseline.
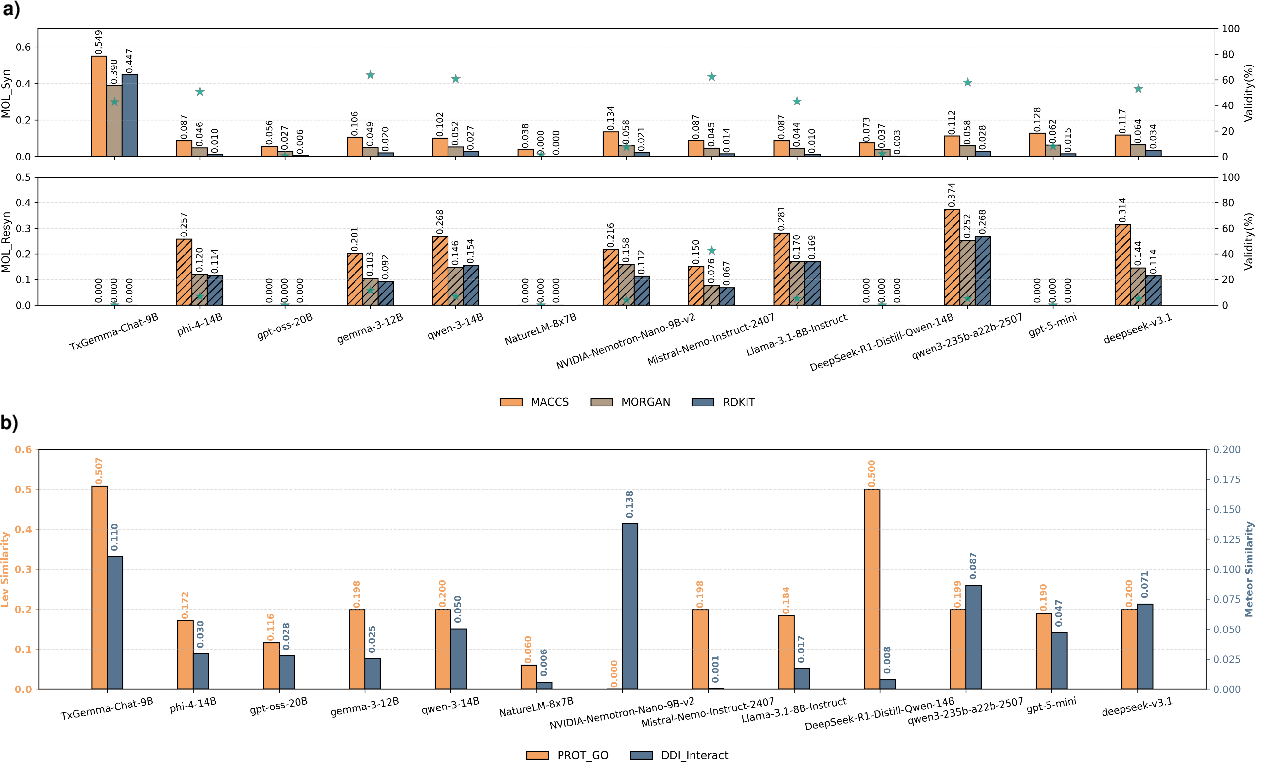
Figure 4: LLM generative performance assessed by molecular similarity and sequence metrics; domain-tuned models lead on certain tasks, but broad generative reliability remains elusive.
The inclusion of external domain tools for prediction provides marked improvements in both regression and classification performance. Disabling tool use results in extreme errors, particularly for regression tasks, demonstrating that unconstrained language modeling remains insufficient for precise, mechanistic scientific tasks. Tool integration yields enhanced consistency and accuracy, matching or exceeding that of much larger models without tools. Moreover, continual pre-training with bio-molecular corpora yields only marginal improvements, hinting at the inherent value of external computation for accurate quantitative prediction.
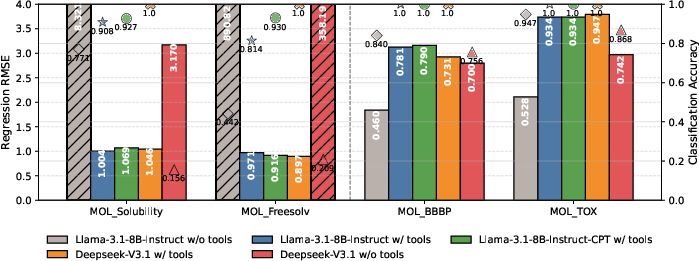
Figure 5: Tool-assisted LLM workflows demonstrate significant improvements in regression accuracy and robust output validity, especially with structured domain information.
Prompt Engineering, Thinking Modes, and Model Sensitivity
Prompt structure, domain knowledge enrichment, and the application of explicit "think" modes have non-trivial influences on model performance, with NVIDIA-Nemotron-Nano-9B-v2 exhibiting particular sensitivity. Detailed, knowledge-informative prompts generally increase accuracy. However, effects of think mode and prompt phrasing are often model- and task-dependent, underscoring the instability and brittleness of LLMs when extrapolating beyond their training regime.
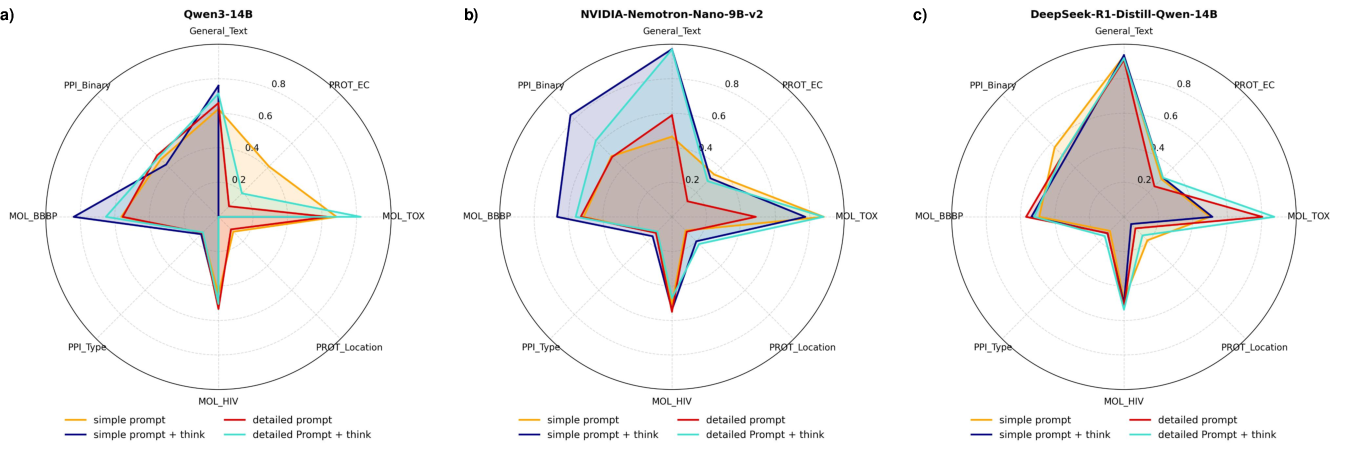
Figure 6: Impact of prompt engineering and think modes across tasks; models benefit from more structured domain guidance but exhibit inconsistent responsiveness.
Mechanistic Reasoning and Interpretability in LLM Outputs
An in-depth examination of the chain-of-thought (CoT) outputs reveals that, despite outwardly coherent scientific explanations, LLMs frequently generate rationales with logical and chemical inconsistencies. The ability to arrive at correct molecular property predictions via spurious or plausible-sounding but incorrect mechanistic reasoning is prevalent, supporting the claim that LLM scientific problem-solving remains largely pattern-driven rather than principled or mechanistically grounded.
Implications, Contradictory Findings, and Theoretical Insights
Several strong and sometimes contradictory findings emerge from the comprehensive evaluation:
- Hybrid Mamba-attention architectures (e.g., NVIDIA-Nemotron-Nano-9B-v2) demonstrate superior performance on tasks requiring the processing of long molecular or sequence data, outperforming models with many times more parameters.
- Chain-of-thought fine-tuning does not improve—indeed may degrade—model performance on specialized biochemistry tasks.
- Supervised fine-tuning drives specialization but causes a marked loss of generalization outside the training distribution.
- All models, regardless of size or training, demonstrate across-the-board failure on regression tasks involving complex, long-sequence dependency resolution, confirming that language-based abstraction alone remains insufficient in these scientific regimes.
- Tool integration is consistently beneficial, enabling otherwise impossible performance on quantitative tasks, and suggests that compositional intelligence combining autonomous LLMs with external computation may be essential.
Conclusion
The cross-scale benchmarking presented in "The limits of bio-molecular modeling with LLMs: a cross-scale evaluation" (2604.03361) establishes a new empirical standard for evaluating LLMs in molecular science. The work demonstrates that architectural innovations such as hybrid Mamba-attention networks are crucial for sequence-heavy biomolecular domains, while further highlighting the domains in which LLMs fundamentally struggle: long-range, mechanistic, and quantitative tasks without tooling. Theoretical implications point toward the necessity of hybrid intelligence paradigms, where LLMs act as orchestrators coordinating expert tools rather than direct replacements for symbolic and computational software. Future progress will likely depend on deeper integration of symbolic reasoning, explicit physics models, and dynamic tool invocation, rather than brute-force language pretraining or scaling alone.





