- The paper introduces a novel expert-in-the-loop approach using LLM-assisted competency question extraction to formalize and validate experimental data semantics.
- It presents a modular ontology framework with core and technique modules that ensure scalable, robust integration of heterogeneous experimental data.
- The evaluation, featuring SPARQL queries and SHACL negative tests, demonstrates its effectiveness in mitigating semantic drift and data inconsistencies.
The AnIML Ontology: Semantic Interoperability for Experimental Data in Scientific Labs
Motivation and Problem Statement
The proliferation of heterogeneous scientific data sources and distributed experimental systems has rendered semantic interoperability a critical barrier in computational science. The Analytical Information Markup Language (AnIML) emerged as a de facto XML-based standard for encoding analytical chemistry and biology data. However, the flexibility of the XML schema has proven problematic: divergent practices across stakeholders induce inconsistencies, which fundamentally undermine reliable data integration and automated reasoning. Existing approaches to ontology extraction from XML for such domains fail to formalize the semantics of reusable core entities and context-dependent data linking.
Ontology Engineering Methodology
To overcome the ambiguity inherent in AnIML's schema and enable robust, cross-system interoperability, the AnIML Ontology leverages an expert-in-the-loop engineering pipeline. The process synergistically combines LLM-assisted requirements elicitation with collaborative ontology development:
- LLM-Assisted Competency Question (CQ) Extraction: The schema and associated documentation are processed using Gemini 2.5 Pro, generating candidate CQs that reflect intended data modeling requirements.
- Expert Validation and Iteration: Domain experts validate, refine, and accept/reject CQs, with rejected items triggering prompt-based guided reformulation cycles.
- Ontology Design Patterns (ODPs): The ontology heavily reuses and extends established ODPs (e.g., Set, Sequence, Situation), enabling modularity and compatibility with foundational ontologies such as BFO and DOLCE.
- Graphical Modeling: Graffoo notation supports interdisciplinary consensus in module definition and relationship scoping.
Through this approach, over 100 expert-validated CQs were formalized, ensuring precise coverage of key experimental data abstractions.
Ontology Structure and Content
The AnIML Ontology (v1.1.0, OWL 2) is partitioned into two main modules—core and technique—and introduces the AnIML Reference Pattern for explicit linked data modeling.
Core Module
The core encapsulates structural aspects of experimental records:
- aml:Document aggregates experimental workflows, sample sets, provenance/audit trail, and signatures.
- Sample Management: Employs the Set ODP for representing complex, hierarchical sample/container arrangements, and the Situation pattern to handle physical and logical groupings.
- Data Containers: The aml:Category pattern uniformly annotates experimental parameters (aml:Parameter) and multimodal data series (aml:Series), supporting consistent extensibility.
- Experiment Workflow: The Sequence ODP structures aml:Experiment and its ordered aml:ExperimentStep constituents, capturing methods, responsible agents, infrastructure, and results.
- Provenance and Compliance: A versioned audit trail (aml:AuditTrail) rigorously tracks atomic changes, document states, and agent attributions, mapped explicitly within the class hierarchy.
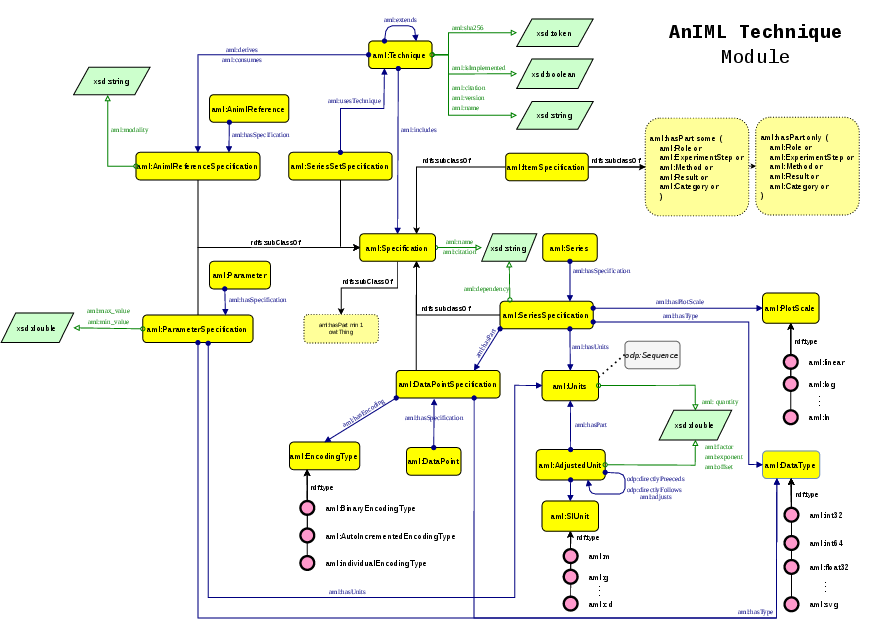
Figure 1: The AnIML technique module, modeling blueprint definitions, constraint imposition, and metadata for methods in the ontology.
Technique Module
Technique-specific definitions (i.e., ATDDs in AnIML) are handled as formal blueprints:
- aml:Technique details the analytical method, versioning, modular hierarchical inheritance, and extensibility via aml:extends.
- Specification Classes: Fine-grained control of data types, units, array structures, and visualization constraints is provided by a suite of Specification subclasses (aml:ParameterSpecification, aml:SeriesSpecification, aml:DataPointSpecification).
- Encoding Semantics: Explicit modeling of data encoding, binary representations, and dependencies supports instrument-agnostic interpretation and visualization.
AnIML Reference Pattern
XML’s ID/IDREF mechanism for indirect references is insufficient for semantic reasoning. The ontology introduces the AnIML Reference Pattern as an ODP to explicitly model references as first-class entities (aml:AnimlReference):
- Instances link with aml:pointsTo, carrying context via aml:hasFunction (role semantics) and aml:startValue/aml:endValue (array slicing semantics).
- Reference sets (aml:AnimlReferenceSet) are strictly scoped to a subject container, ensuring contextually-valid linking.
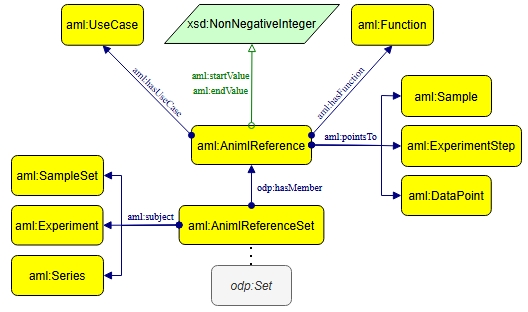
Figure 2: The AnIML reference pattern, elevating ID/IDREF-style links into explicit, context-rich, first-class semantic entities.
Ontology Alignment and Interoperability
Cross-standard interoperability is operationalized via mappings to the Allotrope Foundation Ontologies (AFO), leveraging state-of-the-art ontology alignment systems (LogMap, BERTMap, DeepOnto, OntoAligner):
- Scalable Coverage: Instead of exhaustive class-level equivalence, the focus is on pragmatic alignments underpinning experimental data integration.
- Curation and Relaxation: Automatically discovered equivalences are curated with manual SKOS redefinitions for semantically looser correspondences.
- Numerical Evaluation: Out of 176 candidate alignments, 121 were certified as strict equivalences and 20 as SKOS matches, producing a curated set of 47 unique crosswalks.
Evaluation and Validation Strategy
Evaluation rigorously checks both representational adequacy and unintended model exclusion.
- Structural/Logical Verification: OOPS! is applied for anomaly detection; model refinements guarantee axiomatic consistency.
- Knowledge Graph Instantiation: Industrial AnIML files (anonymized) are transformed to test instance data fidelity; successful AFO-aligned population demonstrates practical integrability.
- CQ Formulation (Positive): 40 CQs are translated into SPARQL queries confirming the ontology’s ability to answer core domain requirements.
- Adversarial Negative CQs: A notable methodological advance is the generation of 20 SHACL-enforced negative CQs, targeting empirical anti-pattern categories (e.g., role/type incompatibility, scope misalignment, cycle violations). Each negative requirement includes designed test datasets, mechanistically validating that the ontology excludes undesired states—an approach rarely systematized in ontology engineering for experimental science.
Implications and Future Directions
The AnIML Ontology delivers a formal, extensible, and modular framework for transforming large, heterogeneous scientific data silos into knowledge graphs enabling semantic interoperability. The design promotes direct mapping from legacy AnIML XML, supporting both rapid adoption in industrial environments and rigorous constraint validation. The explicit modeling of references and provenance is essential in automated scientific workflows and compliance-oriented institutional contexts.
On the theoretical side, the application of LLMs as requirement elicitors within iterative expert-validation cycles presents a template for ontology engineering in data-rich, documentation-heavy domains. The adversarial CQ protocol sets a new standard for validation—directly targeting real sources of semantic drift and unintended modeling.
Future developments will prioritize richer integration with established ontologies for processes, agents, units, and provenance, extend crosswalks to OBO/BioPortal standards, and expand validation as new use cases and stakeholder requirements emerge. This work exemplifies how semantic technologies—when paired with expert/AI-in-the-loop engineering and rigorous model testing—can support next-generation autonomous scientific laboratories and data-driven discovery.
Conclusion
The AnIML Ontology sets a technical foundation for scalable, robust semantic interoperability of experimental data in modern scientific laboratories. It systematically addresses the shortcomings of XML-based syntactic standards, operationalizes advanced ontology alignment and validation strategies, and directly serves the interoperability needs of both industrial and academic research environments.



