- The paper demonstrates that trimodal deep learning with early fusion of histopathology, gene expression, and MRI data achieves the highest composite score (0.854) for glioma survival prediction.
- The study shows that unimodal gene expression outperforms both histopathology and MRI alone, while MRI contributes meaningfully only when combined with RNA and FFPE data.
- The research underscores the impact of fusion strategies, recommending early fusion to mitigate overfitting and effectively integrate diverse modality signals despite limited MRI sample size.
Trimodal Deep Learning for Glioma Survival Prediction: Integration of Histopathology, Gene Expression, and MRI
Introduction
The integration of multimodal data for prognostic modeling in glioma presents a complex challenge due to the inherently heterogeneous character of brain tumors. While prior frameworks have effectively fused histopathology (WSI) and transcriptomics (RNA-seq) for survival prediction, the benefit of incorporating volumetric MRI remains unquantified and methodologically ambiguous. This study addresses this gap by constructing and benchmarking deep learning pipelines across histopathology, gene expression, and FLAIR MRI in the TCGA-GBMLGG/BraTS2021 intersection, utilizing comprehensive unimodal, bimodal, and trimodal evaluations under early, late, and joint fusion paradigms.
Dataset and Preprocessing
The analysis leverages a TCGA-GBMLGG derived cohort (n=664, excluding pediatric/metadata-deficient cases), with the intersection for MRI-labeled data restricted to 162 patients. The detailed preprocessing ensures cross-modality comparability:
- Gene Expression: Feature space of 12,778 genes post z-score normalization and batch correction (ComBat-Seq).
- Histopathology: Extraction of up to 4,000 FFPE WSI patches per slide, normalized for color and stain variability.
- MRI (FLAIR volumes): Volumes resampled to 1283 voxel grids, brain-masked, and augmented for geometric and intensity variations.
The MRI data bottleneck constrains all MRI-inclusive benchmarks to only 19 test cases, creating an acute challenge regarding statistical power and variance in performance estimation.
Neural Architectures and Fusion Strategies
Unimodal Backbones
- Histopathology: A ResNet-50 backbone with patch averaging for per-patient prediction, fine-tuned only at the final block for regularization.
- Gene Expression: An MLP architecture projecting high-dimensional gene signatures to 2048-d latent features.
- MRI: A 3D ResNet-18 trained from scratch (due to sample size and lack of pretraining transferability), with a projection head for fusion compatibility.
Fusion Mechanisms
Three deep multimodal fusion strategies are systematically compared:
- Early Fusion: Feature-level concatenation followed by a fusion MLP.
- Late Fusion: Score-level integration via a cross-validated Cox proportional hazards model.
- Joint Fusion: End-to-end multimodal optimization propagating Cox loss through all encoders.
Additionally, several attention-based fusion mechanisms (bilinear, cross-attention, gated) are evaluated on the FFPE+RNA configuration to assess expressive capacity gains.
All models are trained using Cox partial likelihood loss, with performance reported as Composite Score (CS)—the average of Concordance Index (CI) and $1-$Integrated Brier Score (IBS), with bootstrap-resampled 95% CIs for the CI component. The performance landscape is summarized below.
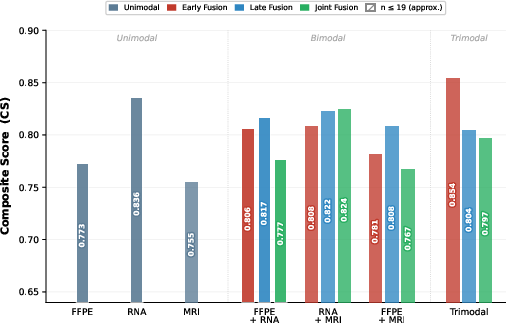
Figure 1: Composite Score (CS) across all modality combinations and fusion approaches, highlighting trimodal early fusion as the top configuration.
Unimodal and Bimodal Results
- Gene Expression sets the performance ceiling among unimodal models (CS=0.836), exceeding both histopathology ($0.773$) and MRI ($0.755$). Notably, the MRI model, despite the absence of transfer learning and low sample size, demonstrates nontrivial discriminative power.
- Bimodal Fusion (FFPE+RNA) achieves CS=0.806 (early)/$0.817$ (late), replicating literature precedent. Importantly, adding MRI to RNA in bimodal settings does not yield a monotonic increase; instead, CS drops from $0.836$ (RNA) to $0.808$ (RNA+MRI), highlighting limitations of simple additive integration in this low-sample regime.
Trimodal Integration
- Trimodal Early Fusion (FFPE+RNA+MRI) yields the top $1-$0 score at $1-$1 on the MRI-intersected subset, which persists as a minor controlled uplift ($1-$2) over FFPE+RNA when both models are restricted to identical patients. Despite the improvement, statistical significance ($1-$3, permutation test) is not achieved, and bootstrap CIs are wide ([0.400, 1.000]) reflecting high variance and limited test set size.
- Late and Joint Fusion are less competitive, with trimodal late fusion dropping to $1-$4 and joint fusion to $1-$5.
Attention-Based Fusion
- Bilinear attention marginally outperforms vanilla MLP ($1-$6 vs. $1-$7) in FFPE+RNA, yet all attention-based configurations exhibit substantial train-test gaps, symptomatic of overfitting at such cohort sizes.
Analysis of MRI’s Marginal Contribution
MRI alone confers prognostic ability comparable to histopathology, but its added value is context-dependent:
- In bimodal settings, MRI’s utility is negligible or negative unless anchored by RNA’s strong signal, and in the case of FFPE+MRI, per-fold discrimination is indistinguishable from random.
- In the trimodal setting, integrated with both gene and histologic context, MRI provides a measurable, if not statistically robust, additive effect.
- The neutral/negative effect in bimodal fusion and positive uplift in trimodal fusion suggests MRI contributes orthogonal information predominantly when sufficient multimodal context exists, supporting a synergy hypothesis over simple signal addition.
Fusion Paradigms and Overfitting Dynamics
Early fusion remains optimal for both bimodal and trimodal integration. The inferiority of joint fusion is especially notable for larger training sets where overfitting risk is amplified by end-to-end fine-tuning of large image encoders. Late fusion, while robust to missing data, suffers from dilution of modality-specific signals, especially in the trimodal setting where it assigns equal weights, thus diminishing RNA’s dominant contribution.
Limitations
The conclusions are tightly bound by the principal limitation: the restricted MRI-matched cohort resulting in small train/test splits (e.g., $1-$8 test for all MRI-inclusive runs), precluding robust IBS estimation, statistical significance, and generalizability. The study's focus on FLAIR alone further limits the exploration of the multi-parametric MRI space, and the absence of external validation (no cohort with WSI/RNA-seq/MRI synergy exists) constrains translational potential.
Clinical and Theoretical Implications
This study affirms the technical feasibility of trimodal fusion for survival prediction in glioma, a scenario increasingly relevant for data-rich, precision oncology centers. The results suggest that—contrary to additive intuitions—imaging only becomes useful in configurations where both spatial and molecular features are present. For real-world deployment, further advances will require expanded, harmonized datasets, robust external validation, and careful consideration of model complexity versus overfitting, especially when integrating high-dimensional medical imaging data.
Conclusion
MRI adds weak but detectable prognostic value when integrated with both histopathologic and transcriptomic data via early fusion, but does not consistently enhance bimodal models. The magnitude and statistical robustness of this effect are severely limited by small sample sizes in the MRI-matched stratum. Methodologically, early fusion remains the preferred integration paradigm at this scale, and attention-based fusion provides only marginal gains. Future research should prioritize assembling larger, multimodal, harmonized cohorts, leveraging multi-sequence MRI, and rigorous out-of-distribution evaluation to realize the translational impact of truly holistic, multimodal neural prognosticators in neuro-oncology.
