- The paper introduces a Bayesian finite mixture model that unifies clustering of mixed-type data with robust variable selection and handling of censored biomarkers.
- The methodology employs multiple covariance structures and likelihood-based imputation with spike-and-slab priors, achieving high clustering accuracy (up to 0.98 ARI) in simulation studies.
- Applications to sepsis phenotyping and acute lung injury demonstrate its ability to uncover clinically relevant subgroups and treatment heterogeneity.
Bayesian Finite Mixture Modeling for Mixed-Type Data Clustering and Variable Selection with Censored Biomarkers
Introduction
The paper "A Bayesian Finite Mixture Model Approach for Mixed-type Data Clustering and Variable Selection with Censored Biomarkers" (2603.29316) presents a Bayesian finite mixture model (BFMM) framework tailored for clustering heterogeneous biomedical datasets comprising both continuous and categorical variables, often complicated by cluster-specific dependencies and censored biomarker measurements. The model directly addresses the inadequacies of prevalent clustering strategies—in particular, their failure to accommodate cluster-specific covariance, censoring mechanisms, and variable selection in a unified fashion.
The BFMM models heterogeneous data through a finite mixture of parametric distributions with distinct treatment of continuous and categorical variables. For continuous features, each cluster is endowed with its own multivariate normal component, with a choice among three covariance structures—EEI (diagonal, equal), EEE (general, equal), and VVV (general, cluster-specific)—supporting varying levels of within-cluster dependency. Categorical variables are modeled through cluster-specific multinomial (categorical) probabilities.
Censoring in continuous variables (such as those caused by assay lower or upper detection limits in clinical biomarkers) is integrated via a fully likelihood-based imputation scheme inside the Gibbs sampler, with censored values sampled from their appropriate truncated conditional posteriors at every iteration.
Feature selection is performed through the incorporation of spike-and-slab priors: a normal mixture for continuous variables and a Dirichlet mixture for categorical probabilities, controlled by latent binary inclusion indicators with cluster and variable specificity. The variance ratio between the spike and the slab is adaptively calibrated using empirical cluster means, ensuring separation between informative and noise variables.
Posterior Inference and Label Switching
BFMM parameters are inferred via an MCMC Gibbs sampler with direct updates for all parameters given conjugate priors. Posterior inference includes relabeling for the mixture components via the Stephens' KL divergence minimization procedure, addressing the label-switching phenomenon and producing stable posterior summaries.
Model selection for the number of clusters (G) and the covariance structure utilizes BIC and ICL criteria, balancing penalized likelihood and cluster assignment uncertainty.
Simulation Studies
Comprehensive simulation studies investigate cluster recovery accuracy (ARI), convergence, and variable selection under diverse data-generating schemes, including various correlation structures (mapped to EEI, EEE, VVV) and degrees of censoring in key variables.
(Figure 1)
Figure 1: Violin plots summarizing adjusted rand index (ARI) by different clustering methods in each simulated scenario. Categorical variables in simulated datasets were treated as continuous when applying the mclust and bootstrap K-means clustering methods.
Results demonstrate that BFMM, with the appropriate choice of covariance structure, delivers superior ARI across all simulated conditions, including high levels of censoring, outperforming both marginal independence latent class models and distance-based approaches. Notably, for strong within-cluster correlations or high censoring, only BFMM[VVV] achieves robust, high ARI, while standard methods exhibit high failure rates or instability. Variable selection is sharp: dominant features consistently receive high posterior inclusion, while noise variables are down-weighted regardless of censoring or heteroscedasticity.
Application: Sepsis Phenotyping with EHR Data (SENECA)
BFMM[VVV] was applied to the SENECA sepsis cohort comprising 20,189 patients and 28 variables (continuous and categorical). Model selection via BIC and ICL supports 4 clusters with VVV structure.
(Figure 2)
Figure 2: BIC and ICL evaluation for SENECA data by BFMM given G=1,…,9 clusters. Information criteria evaluation used Markov chains with 10000 Gibbs sampling iterations and 5000 burn-in.
Clusters correspond to clinically meaningful phenotypes, differentiated principally by markers of organ dysfunction (troponin, AST, lactate, GCS, SBP, bicarbonate), each carrying high variable importance weights (≥0.85). Categorical demographic variables were correctly identified as non-influential (importance <0.3). Phenotypic assignment corresponds to distinct risk gradients in ICU admission and multi-hospital mortality.
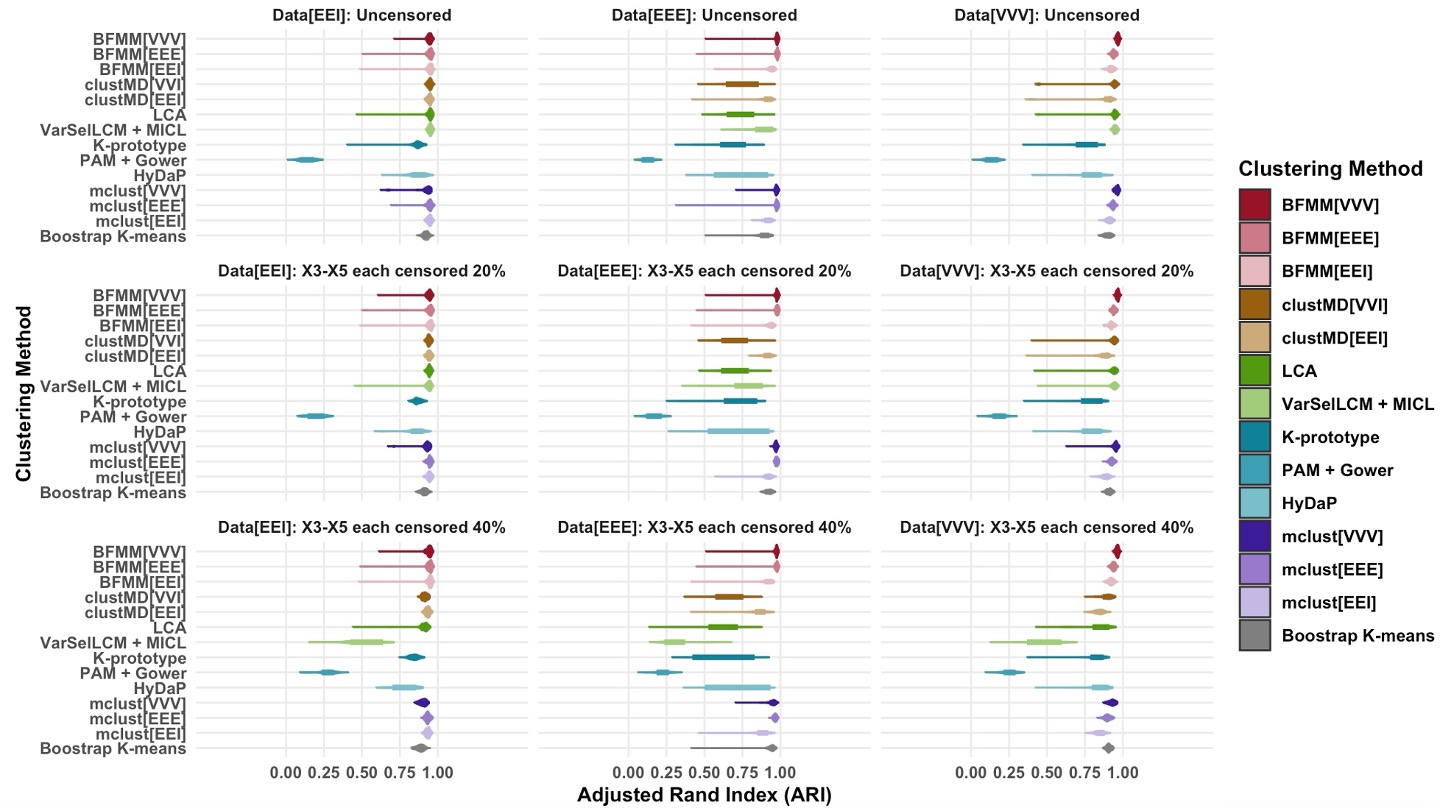
Figure 3: Distributions of the six clinical endpoints by BFMM[VVV] clustering results of SENECA dataset. The annotated p-values are based on Chi-squared test comparing the proportions among the four clusters.
Application: Acute Lung Injury Subgroups in Clinical Trial Data (EDEN)
In the EDEN trial (N=889, 37 variables, multiple left-censored biomarkers), only BFMM[EEI] with G=3 clusters is supported; full-covariance models demonstrate overfitting given sample size limitations.
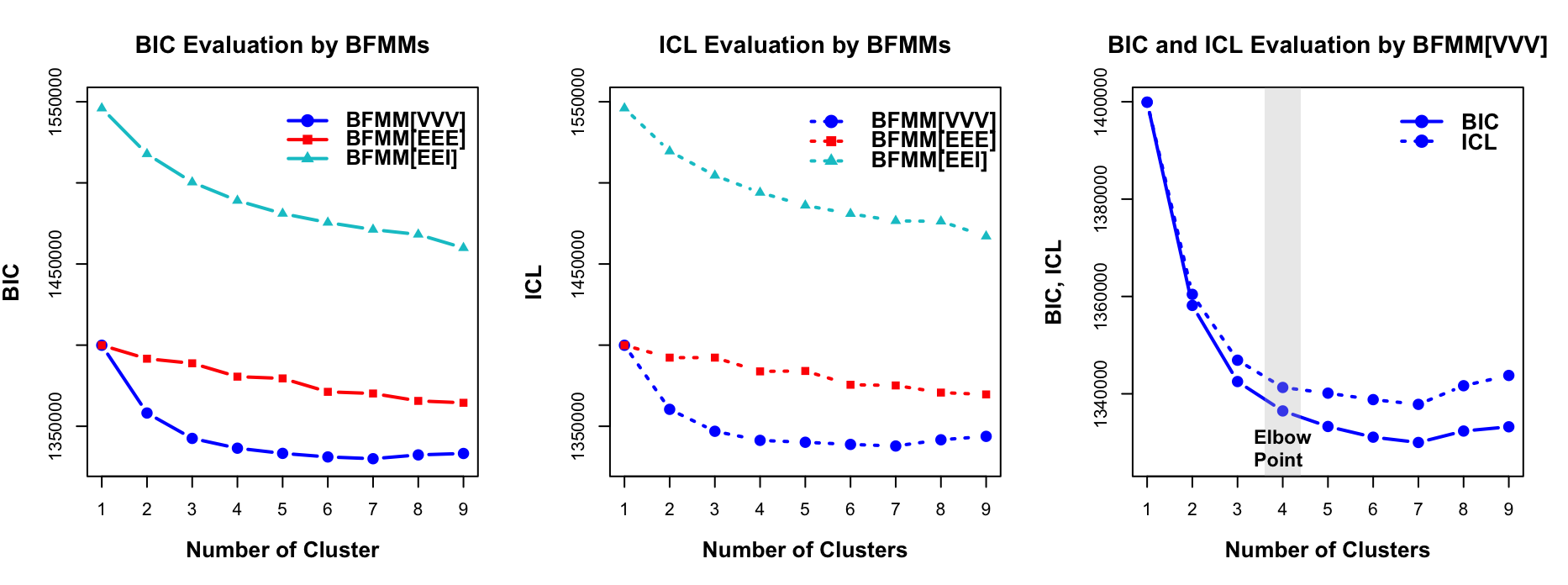
Figure 4: BIC and ICL evaluation for EDEN trial data by BFMM given G=1,…,6 clusters. Information criteria evaluation used Markov chains with 5000 Gibbs sampling iterations and 2500 burn-in.
The resulting subgroups are defined primarily by five biomarkers (RAGE, TNF-α, IL-6, PCT, IL-8) and renal function markers (BUN, creatinine), as well as treatment-related variables (PEEP, MV), with top 9 variables receiving weights ≥0.8. Notably, cluster membership uncovers treatment-covariate interaction: within one biologically defined group, patients randomized to trophic feeding experience a statistically significant reduction in gastrointestinal intolerance (p=0.0187), while overall and in other subgroups no effect is observed.
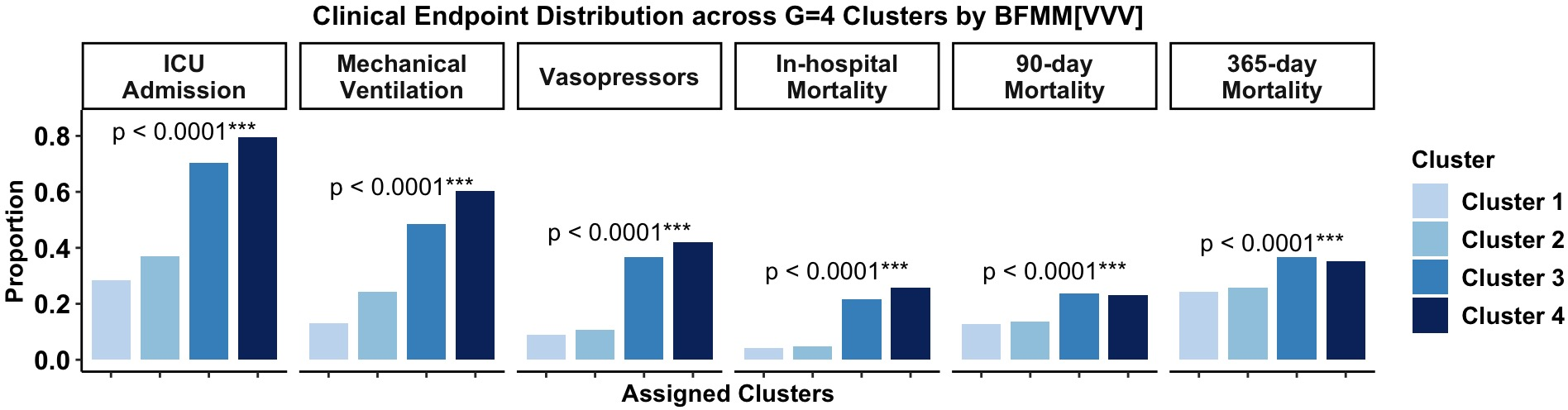
Figure 5: Distributions of the three clinical endpoints by BFMM[EEI] clustering results of EDEN trial dataset. The annotated p-values are based on Chi-squared test comparing the proportions among the three clusters in the top panel, and between the two feeding groups within each cluster in the bottom panel.
Discussion and Theoretical Implications
The Bayesian finite mixture approach unifies mixed-type clustering, handling complex dependencies, likelihood-based imputation for censored measurements, and feature selection in a principled manner. The inclusion of multiple covariance structures and adaptive hyperparameter calibration ensures both flexibility and resilience to the cluster correlation spectrum and missing data patterns.
Simulation and application studies support several strong claims of the framework:
- Consistent superiority in clustering accuracy (up to 0.98 ARI) and variable selection stability over state-of-the-art methods for mixed or censored data.
- Robust imputation for censored data, outperforming complete case and naive thresholding alternatives.
- Convergent and interpretable variable importance measures for both continuous and categorical domains.
- Empirical demonstration of subgroup detection revealing treatment effect heterogeneity otherwise masked in overall analysis.
These features have clear practical import for biomedical informatics and translational research, particularly in high-dimensional EHR mining and RCT subgroup identification for precision medicine.
Limitations and Future Directions
The model currently assumes multivariate normality for continuous data and conditional independence for categorical variables within clusters; relaxation to non-Gaussian or latent variable copula models would broaden applicability. MCMC scaling remains a bottleneck for very high-dimensional or large N applications, indicating a need for further algorithmic acceleration (e.g., variational or stochastic MCMC). Direct generalization to more flexible covariance parameterizations (e.g., all 14 parsimonious GMM structures) and improved automated hyperparameter selection for categorical variables are logical next steps.
Conclusion
This work establishes Bayesian finite mixture models with adaptive covariance structure, variable selection, and censored data modeling as a rigorous, interpretable framework for cluster discovery in heterogeneous biomedical datasets. The methodological design and empirical results demonstrate practical capability for identifying clinically meaningful subgroups and prognostic strata, and highlight the potential for detecting treatment heterogeneity critical to the advancement of precision medicine.


