- The paper presents the PASTA framework that uses a conditional diffusion model to translate MRI into PET images with enhanced pathology preservation.
- It employs a symmetric dual-arm U-Net and adaptive conditional normalization to integrate multi-modal data, thereby improving image fidelity and disease sensitivity.
- Extensive experiments demonstrate that PASTA outperforms GAN-based methods, achieving superior quantitative metrics and boosting diagnostic accuracy in neurodegenerative analysis.
Conditional Diffusion Models for Pathology-Aware MRI-to-PET Translation: The PASTA Framework
Introduction and Motivation
Translational imaging between MRI and PET is critical in neurodegenerative disease diagnostics, notably Alzheimer's disease (AD), where structural MRI reveals anatomical atrophy and FDG-PET maps metabolic decline. PET's clinical utility is hampered by cost and ionizing radiation; MRI is cheaper and widely available but less sensitive for early pathology. Existing generative models—primarily GAN-based—struggle to synthesize PET from MRI with adequate pathological specificity, often emphasizing generic structural fidelity at the expense of accurate disease signatures.
This work introduces PASTA, a pathology-aware, conditional diffusion framework for 3D MRI-to-PET synthesis. PASTA leverages a symmetric dual-arm U-Net architecture, extensive multi-modal conditioning (including anatomical, clinical, and pathology priors), a novel cycle exchange consistency training paradigm, and a computationally efficient 2.5D volumetric generation strategy.
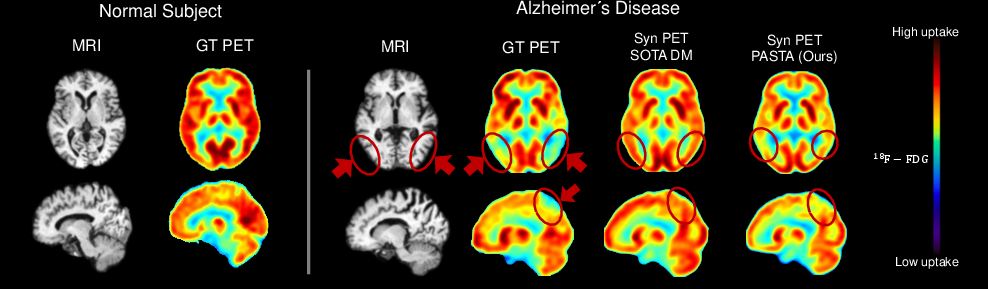
Figure 1: For AD, PET reveals reduced glucose uptake in the temporoparietal lobe. While SOTA diffusion models fail to recover this pathology when translating MRI to PET, PASTA preserves disease-relevant metabolic patterns.
Methodological Contributions
PASTA Architecture
PASTA fuses three key modules: the conditioner arm (anatomical encoder), the denoiser arm (diffusion model decoder), and adaptive conditional normalization (AdaGN). The conditioner arm (a U-Net) ingests MRI volumes, extracting task-aware features, while the denoiser arm reconstructs PET from noise under multi-level conditioning.
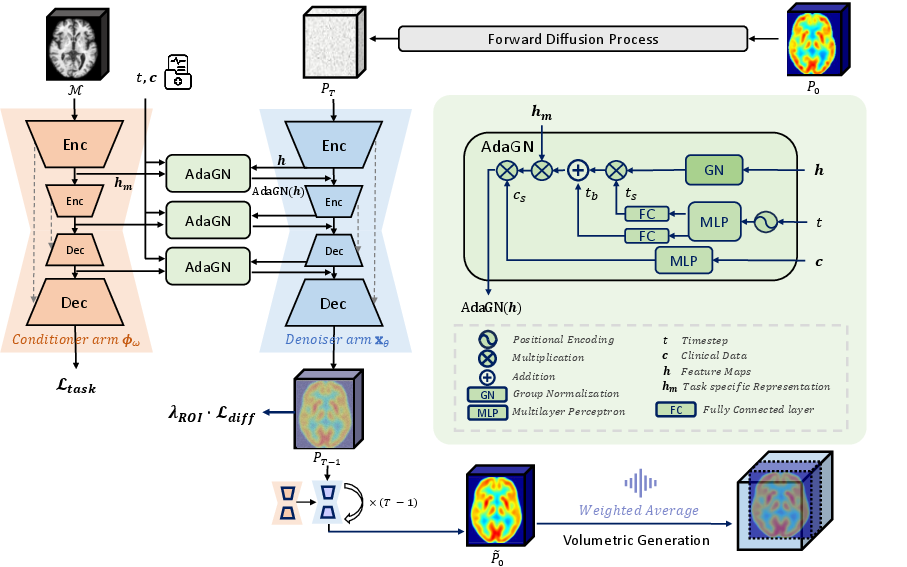
Figure 2: PASTA’s symmetric dual-arm structure: the conditioner arm (ϕω) and denoiser arm (xθ) interact via AdaGN to enforce multi-modal conditioning across the generation process.
Multi-modal fusion is realized through AdaGN layers, which simultaneously condition on timestep, clinical data, and the conditioner’s task-specific feature maps. Further, slice-aware AdaGN (SA-AdaGN) encodes slice position to enhance volumetric coherence.
Meta-ROI priors, derived from AD-associated hypometabolic regions, are injected as spatial loss weightings, directly biasing the generative process toward pathology-preserving outputs.
Cycle Exchange Consistency (CycleEx)
CycleEx generalizes standard cycle-consistency training: PASTA's dual arms are exchanged to learn both MRI-to-PET and PET-to-MRI mappings simultaneously, with the symmetric U-Net architecture enabling parameter reuse and information regularization.
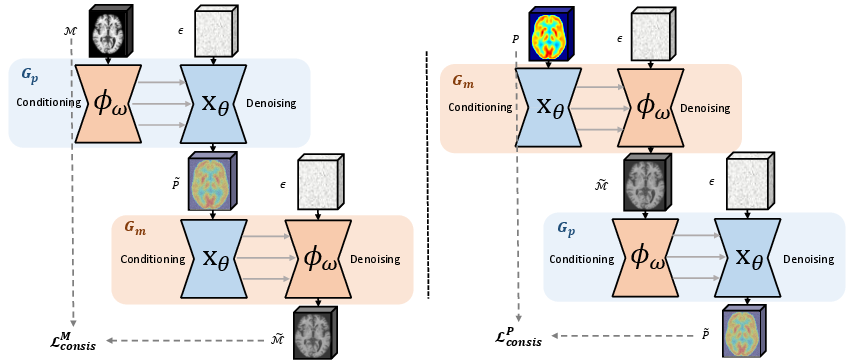
Figure 3: CycleEx: forward and backward mappings (Gp, Gm) ensure round-trip translation consistency by swapping conditioner and denoiser arms. This drives the extraction of modality-specific but pathology-coherent representations.
Efficient Volumetric Generation
For tractable 3D synthesis, a 2.5D hybrid strategy is employed: the network receives overlapping sequences of N slices and reconstructs PET slices along all anatomical axes, aggregating outputs to form a consistent 3D scan. This mitigates inter-slice inconsistency versus pure 2D approaches without the memory footprint of full 3D models.
Experimental Evaluation
Datasets and Setup
The framework is validated on 1,248 paired T1w MRI/PET scans from the ADNI database and 253 in-house clinical pairs, spanning cognitively normal, MCI, and AD subjects. MetaROIs are mapped to standard MNI152 space.
Robust data splitting preserves demographic and diagnostic balance; all results are averaged over held-out test sets and, for the in-house cohort, full 5-fold cross-validation.
Quantitative and Qualitative Results
Image Fidelity and Pathology Preservation
Compared to GAN-based (CycleGAN, Pix2Pix, ResVit), domain-aligned (RegGAN), and diffusion-based (BBDM, BBDM-LDM) baselines, PASTA sets state-of-the-art benchmarks on MAE, MSE, PSNR, and SSIM across all datasets. Specifically, on ADNI, PASTA attains MAE =0.0345, PSNR =24.59, SSIM =86.3%—statistically significant versus all baselines (p-values <0.0001).
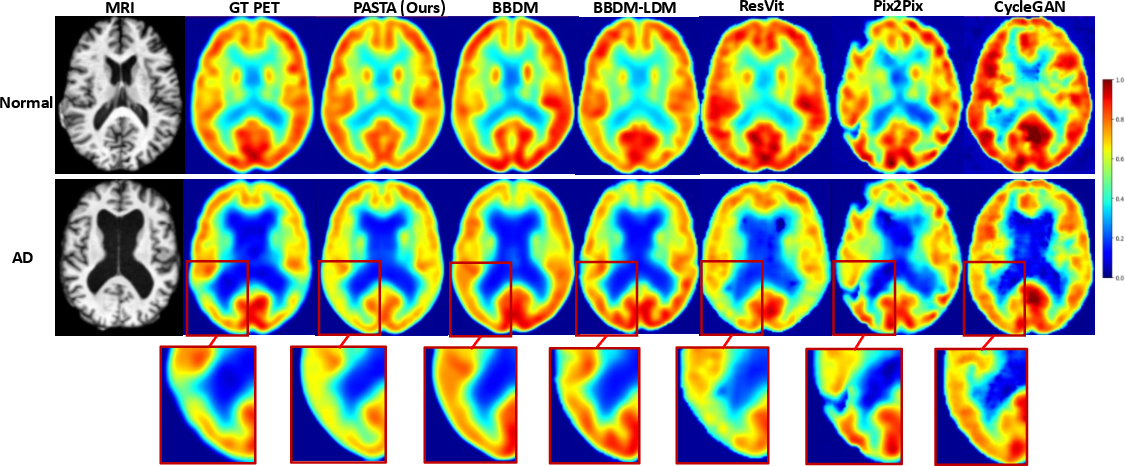
Figure 4: Normal and AD cases—PASTA uniquely recovers temporoparietal hypometabolism (bottom left/right), whereas baselines either miss or blur these neuropathological patterns.
Crucially, region-localized metrics restricted to MetaROIs further highlight superiority: in disease zones, PASTA achieves MAEROI=9.07×10−4, with SSIMROI=99.7%.
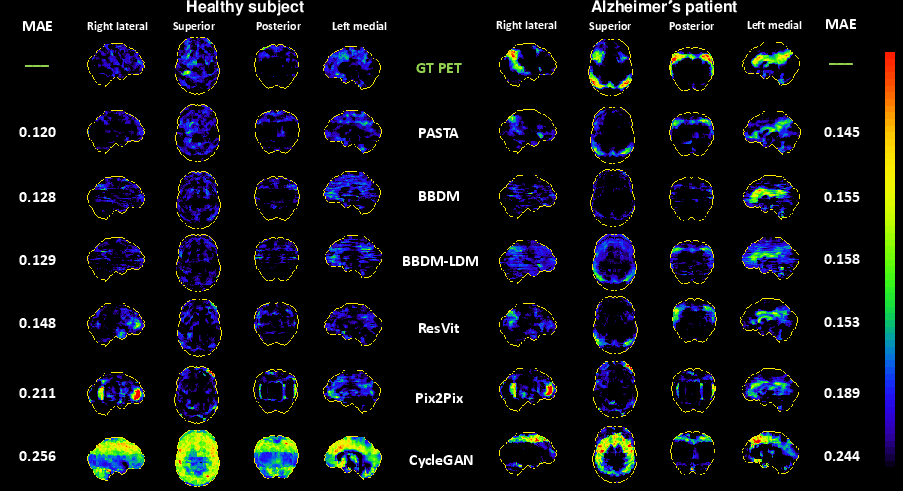
Figure 5: Neurostat 3D-SSP Z-score maps—PASTA's synthesized PET best preserves metabolic deficits in AD, matching ground-truth projections and minimizing quantitative error in key cortical areas.
Classification experiments using synthesized PET significantly boost AD detection accuracy relative to MRI. PASTA yields BACC =83.4%, F1 =80.0%, AUC =91.6%, notably outperforming both MRI and all other PET synthesis methods. Notably, inclusion of clinical conditioning improves fidelity and disease sensitivity, as evidenced by ablation studies on clinical variable impact (Table: Clinical Variable Sensitivity).
Ablation and Analysis
Ablations clarify the importance of CycleEx (12% MAE reduction), pathology-prior weighting, AdaGN-mediated conditioning, and the number of context slices N (optimal at N=15). SA-AdaGN and auxiliary classifier-consistency regularization slightly improve structural and ROI-based fidelity at a marginal global cost.
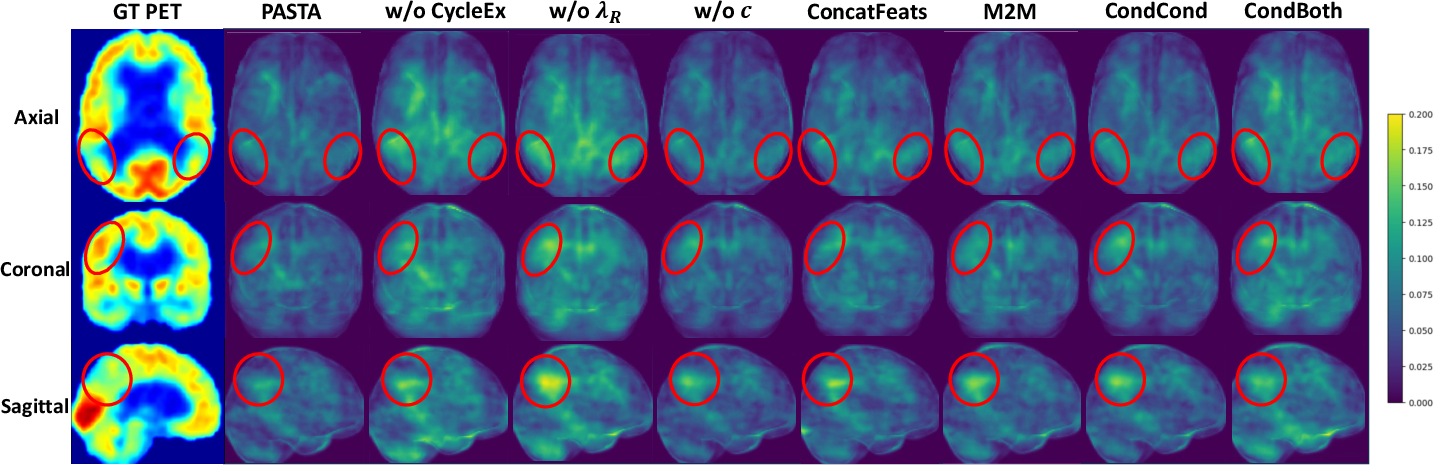
Figure 6: Error maps for ablation—removal of CycleEx or MetaROI priors increases artifact prevalence in AD-affected cortex (red circles), supporting their critical role in pathology preservation.
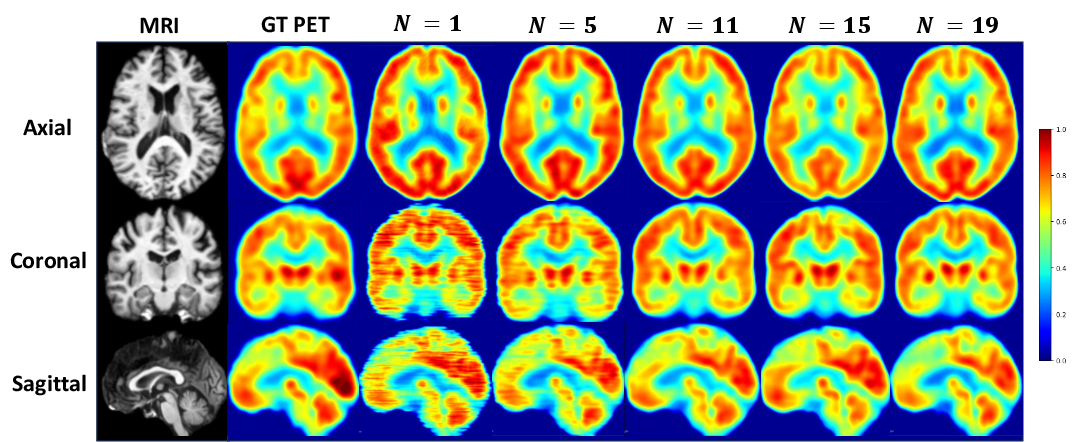
Figure 7: Input context size—N too small induces inter-slice artifacts; too large produces oversmoothing. PASTA optimally balances fidelity and consistency at N=15.
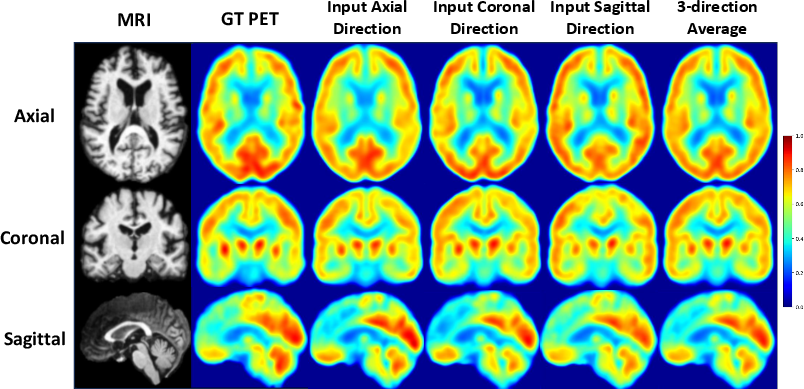
Figure 8: Consistency across anatomical axes—syntheses from axial, coronal, and sagittal slice inputs yield negligible cross-axis discrepancies; 3-axis averaging offers no substantial improvement, confirming robust volumetric integration.
Fairness
No statistically significant synthesis error is observed across age, gender, or diagnosis subgroups, confirming demographic robustness.
Computational Costs
While PASTA with CycleEx increases training time (3.7x per step, 2.3x GPU memory), inference remains efficient and the accuracy gains on challenging medical translation tasks justify the overhead.
Theoretical and Practical Implications
PASTA demonstrates that conditional diffusion models, when designed with strong pathology priors, comprehensive multi-modal conditioning, and efficient architectural regularization, can synthesize cross-modality medical images with clinically meaningful pathology retention. The findings challenge prior assumptions that high-fidelity generative translation will suffice for diagnostic purposes, highlighting the necessity of explicit disease-driven supervision. Moreover, PASTA’s modular design is applicable beyond the MRI/PET domain, inviting adaptation to other challenging modality pairs or scenarios with limited ground-truth data.
Conclusion
PASTA advances MRI-to-PET translation by deploying conditional diffusion architectures tailored for pathology preservation. By explicitly fusing anatomical, clinical, and region-specific priors and enforcing cross-domain cycle consistency, PASTA exceeds the structural and diagnostic performance of competing models. The pathology-aware synthesis paradigm presented here makes a substantial contribution toward the deployment of reliable, functionally informative cross-modality translation systems in neuroimaging and can serve as a blueprint for broader clinical AI development. Future work should pursue formal prospective reader studies, extension to other diseases, and more advanced multi-modal integration to further validate and generalize its clinical impact.







