Evolving Symbiosis, from Barricelli's Legacy to Collective Intelligence: a simulated and conceptual approach
Abstract: This report documents the work of our group (named SymBa) at the ALICE 2026 workshop in Copenhagen. Inspired by the pioneering work by Nils Aall Barricelli on symbiogenesis of numerical organisms (i.e., 1D cellular automata) in 1953 (70+ years ago!!), we discussed the role of symbiogenesis as mechanism contributing to the origins of life, open-endedness, and collective intelligence. We report replications of Barricelli's original work in 1D worlds, an extension to 2D symbioorganisms, and preliminary experimentation with DNA-norms. We discuss the implications of symbiogenesis for artifical life and artificial intelligence, and outline several opportunities for future works, both at the conceptual level as well as using different substrates (neural networks, neural cellular automata, etc.)
Paper Prompts
Sign up for free to create and run prompts on this paper using GPT-5.
Top Community Prompts
Explain it Like I'm 14
What this paper is about (big picture)
This paper looks at how “teaming up” can create new, more complex living-like systems—even inside a computer. The authors revisit and extend the ideas of Nils Barricelli, a pioneer from the 1950s who showed that simple digital “organisms” can copy themselves, mutate, and, most importantly, combine in helpful ways (symbiosis) to form bigger, smarter structures. The team rebuilds his experiments, expands them into two dimensions, and tries DNA-like rules to see how cooperation can lead to open-ended growth and collective intelligence.
What questions the researchers asked
The authors focus on two simple questions, asked in everyday terms:
- When do partnerships between simple parts become stable and lasting—turning two “things” into one new, better “thing”?
- Is teaming up (symbiogenesis) not just helpful, but maybe even necessary, for never-ending creativity and complexity in life and in artificial systems?
They also ask how these ideas relate to today’s AI and to our understanding of how life might have begun.
How they studied it (methods in plain language)
Think of a video game played on a grid of cells with easy, local rules—like a checkerboard where pieces move and copy themselves based on simple instructions. The researchers used several versions of this idea:
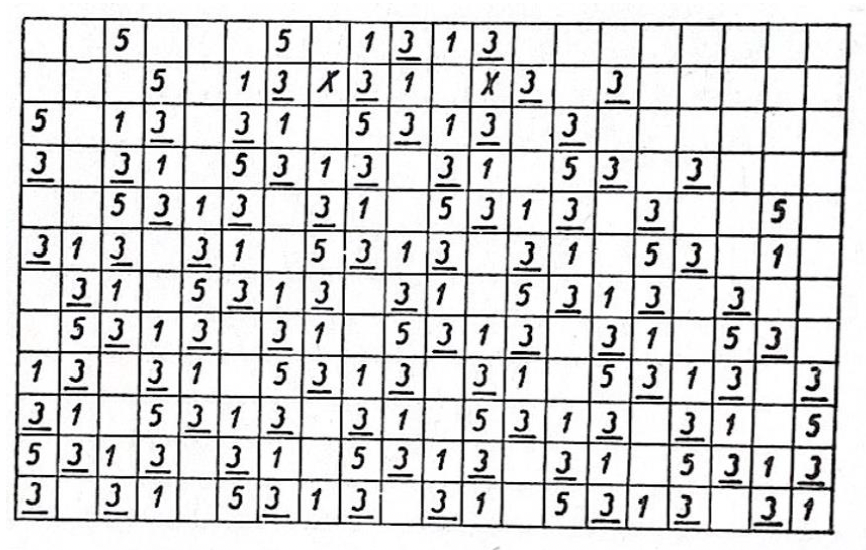
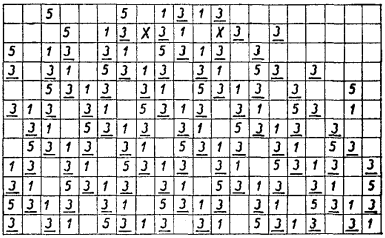
- Rebuilding Barricelli’s 1D “number world”:
- Picture a long line of boxes. Each box can hold a number (a “gene”) or be empty.
- Each number tries to copy itself to a new spot a certain distance away (like “move right by 4” if the number is 4).
- If two numbers try to land on the same spot, a “house rule” (called a norm) decides what happens—maybe they cancel out, combine, or produce a new number. These are like traffic rules for collisions.
- Over time, strings of numbers can form and act like digital organisms that copy themselves, repair damage, or even show parasitic behavior.
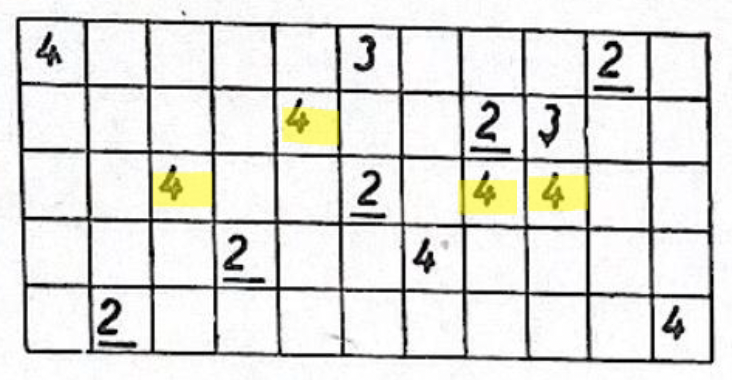
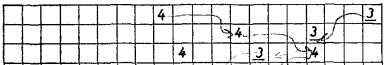
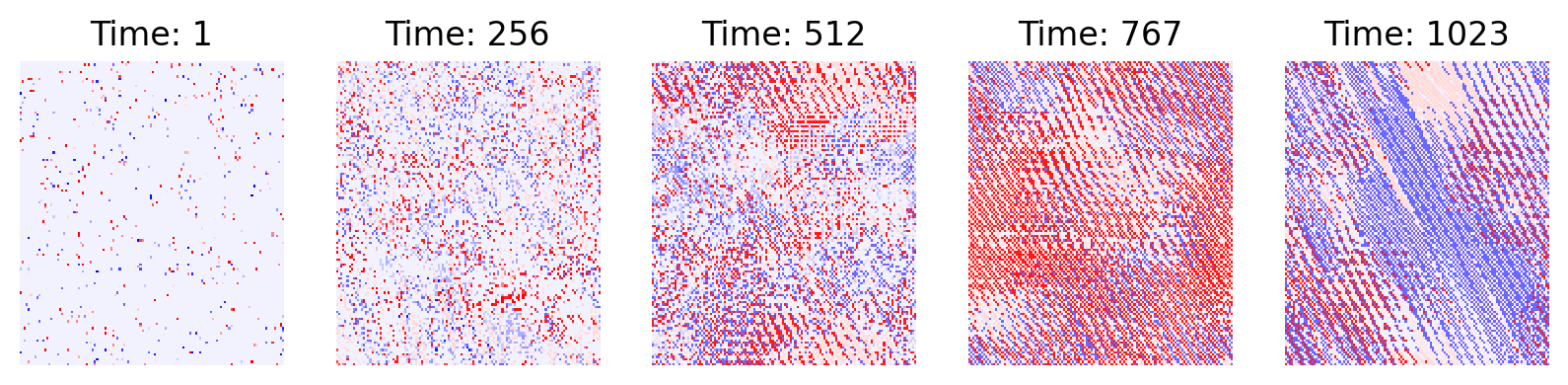
- Extending to 2D:
- Now the grid is a full map, not a line. Each cell holds a small “arrow” (a direction to move).
- The same copying and collision rules apply, but now patterns can spread in all directions, creating wave-like behavior.
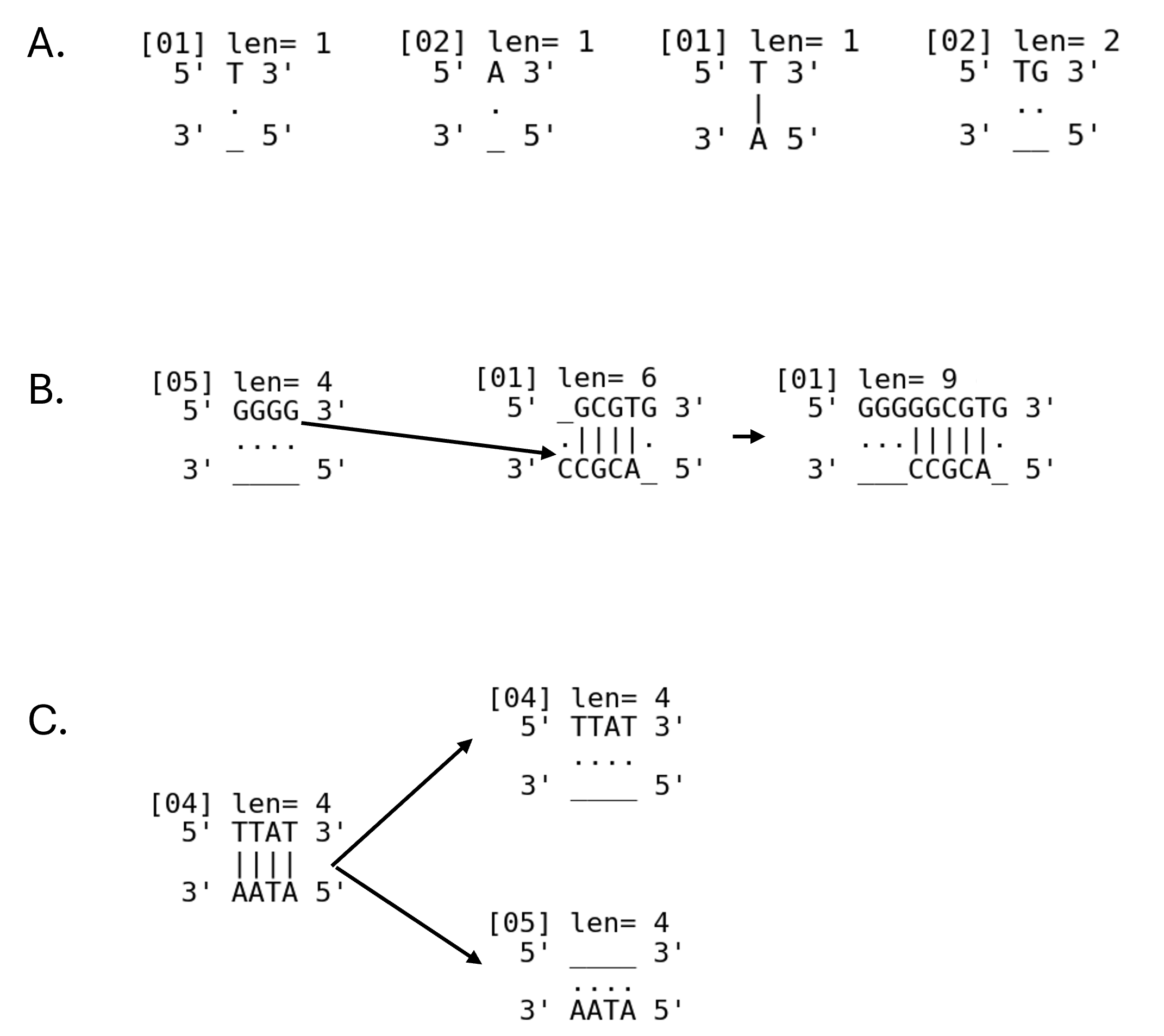
- Trying DNA-like rules (a minimal “DNA soup”):
- Instead of just numbers, the world has four “letters”—A, C, G, T—like real DNA.
- Three simple rules:
- 1) Elongation: strands grow by adding one letter at the end.
- 2) Complementary association: matching letters “stick” (A with T, C with G), letting strands align and merge if they overlap.
- 3) Split: when a double strand is complete enough, it can unzip into two single strands—like reproduction.
- They compared two setups: only growth (elongation) vs. growth + sticking + splitting.
- Measuring what happens:
- Counting how many cells are occupied and how often collisions happen—like taking the pulse of the digital world.
- Using simple information measures:
- Entropy: “How surprising or varied is the pattern?”
- Mutual information: “How much does one moment tell us about the next?”
- Testing robustness: poke a digital organism and see if it survives or repairs itself.
What they found and why it matters
Here are the main takeaways, explained with simple parallels:
- In Barricelli’s 1D worlds, cooperation matters:
- Just copying and mutating doesn’t go very far.
- Letting numbers “team up” creates longer, self-repeating sequences—digital organisms that can recombine, show parasitism, and even repair themselves.
- If the whole world uses one strict rule, things settle into simple loops. But mixing rules in different regions keeps the system lively for longer—more like an ecosystem than a mono-crop.
- In 2D, complex patterns spread:
- When the grid expands to two dimensions, patterns don’t just repeat—they form waves and moving fronts, similar to how living tissues or colonies expand.
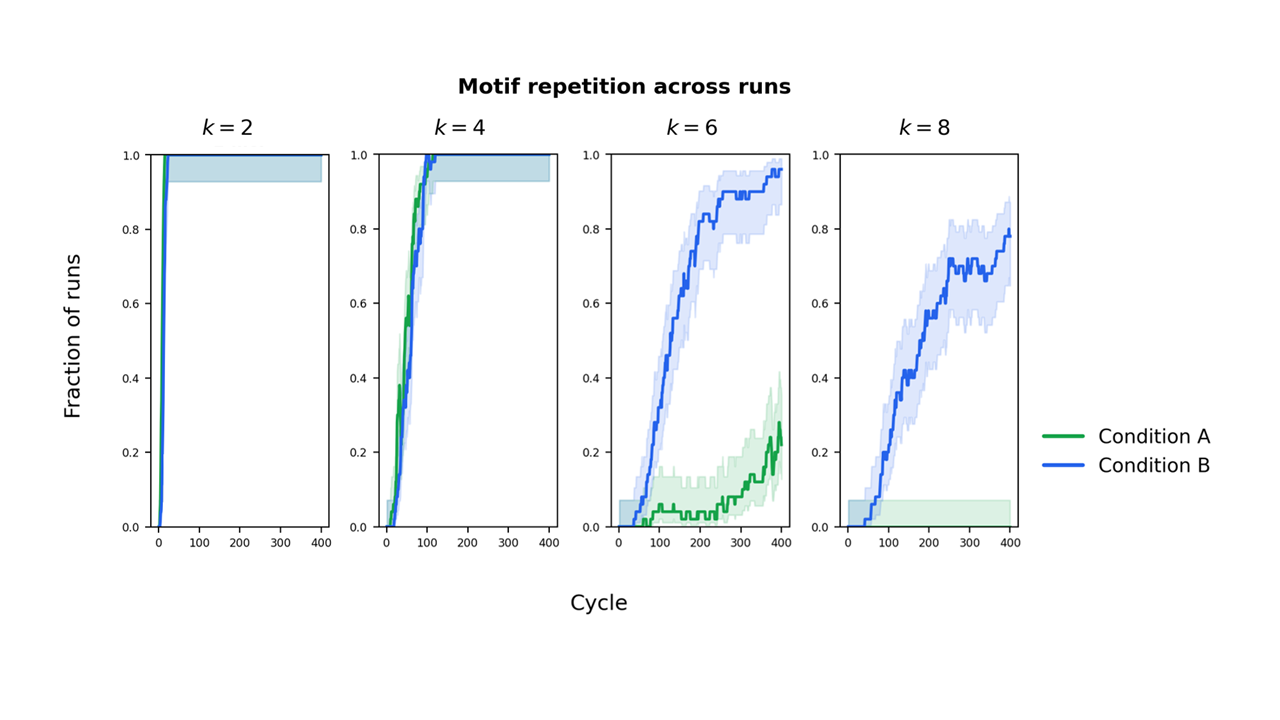
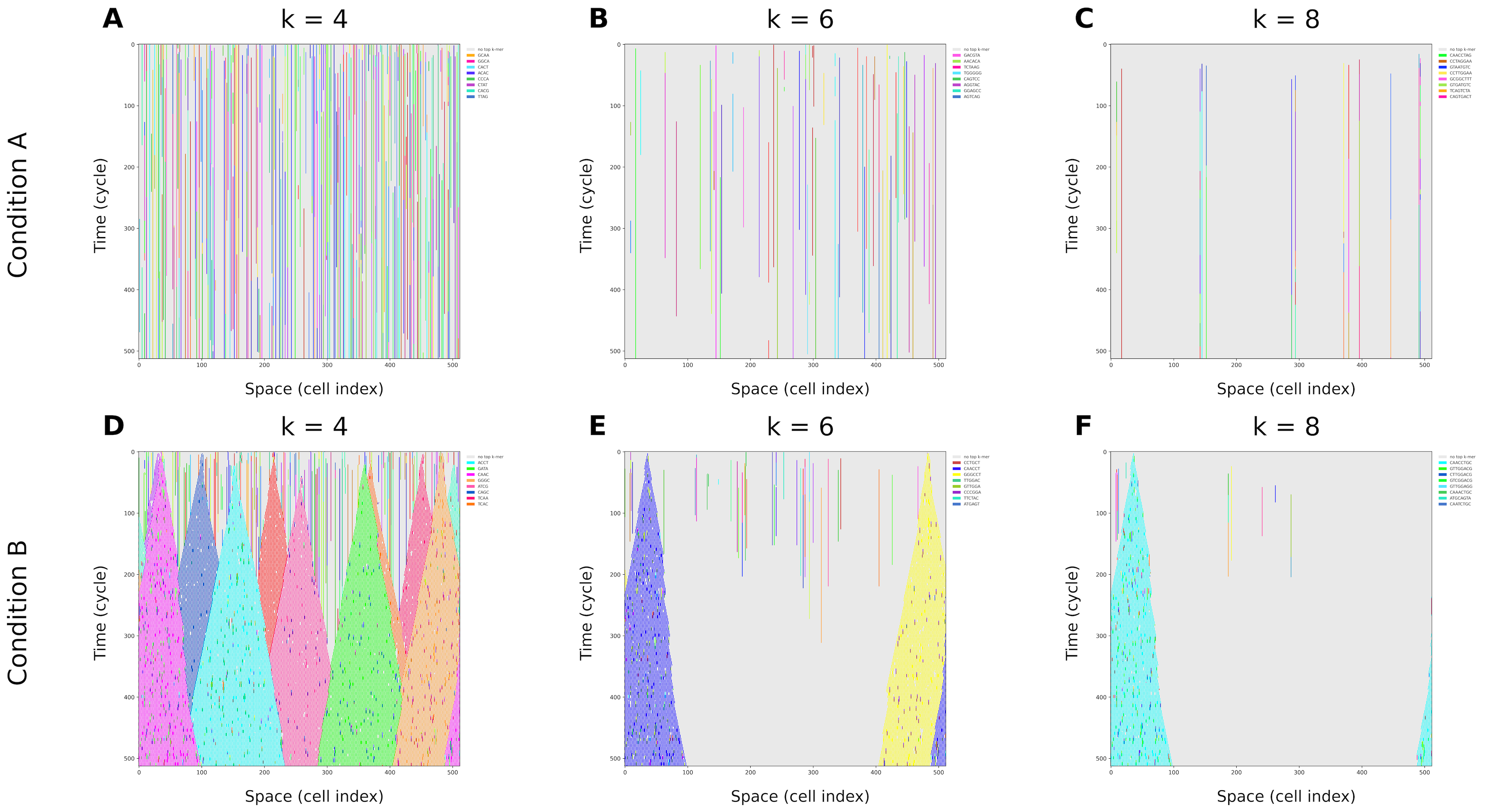
- DNA-like rules make a big difference:
- With only “grow by one letter,” long repeated pieces almost never show up.
- Add “stick to your complement” and “split,” and suddenly longer repeated chunks appear a lot more often, and they spread across space.
- In the 1D spatial version, you see clear “domains” (patches) where certain DNA-like motifs dominate and reproduce locally. Multiple different motifs coexist, compete, and interact—like an emerging ecosystem from simple parts.
- Information grows with structure:
- The information measures show how patterns become less random and more structured over time when symbiotic rules are allowed, hinting that cooperation helps organize and preserve useful information.
Why this matters:
- It supports the idea that life’s big leaps (like cells gaining mitochondria) may depend on symbiogenesis—permanent partnerships that create something new.
- It shows a path toward “open-ended” evolution in computers: not just repeating the same tricks, but discovering new ones as parts learn to cooperate.
What this could lead to (implications)
- Understanding life’s origins:
- If teaming up helps kickstart evolving systems, it may explain how early life moved from simple molecules to complex cells.
- Better AI and collective intelligence:
- Instead of building one giant model, we might design many simple agents or modules that can team up, split, and reorganize—growing abilities organically.
- This could make AI systems more adaptable and robust, like communities that survive shocks because members support and challenge each other.
- Designing resilient digital ecosystems:
- Mixed rules and local interactions can keep systems lively and creative, avoiding “one winner takes all” outcomes.
- Future steps suggested by the authors:
- Give sequences “functions” that affect resources or rules (like enzymes), letting organisms shape their environment.
- Explore the simplest possible setups (even simple true/false grids) to see when symbiosis first appears.
- Measure and test robustness more carefully: how well do digital organisms survive disturbances?
In short, the paper argues that cooperation isn’t just a nice add-on—it may be the engine that turns simple parts into complex, evolving systems. Whether we’re talking about early life or future AI, learning how simple things learn to “live together” could be the key to open-ended creativity and intelligence.
Knowledge Gaps
Knowledge gaps, limitations, and open questions
Below is a single, consolidated list of what remains missing, uncertain, or unexplored in the paper, framed to guide concrete next steps for future research.
- Lack of a formal, operational definition of “symbioorganism” and an automated detector for them; future work should specify detectable criteria (e.g., reproducible multi-gene patterns, persistence thresholds, lineage continuity) and provide algorithms to identify, track, and compare such entities across runs.
- No quantitative test of the permanence of symbiotic associations; design experiments that measure when and how transient cooperative interactions become obligate, irreducible units (e.g., by measuring dependency graphs, component survivability outside the collective, and time-to-reversion under perturbations).
- The central question “Is symbiogenesis beneficial or required for open-endedness?” remains untested; implement open-endedness metrics (e.g., unbounded growth in complexity/novelty, diversity turnover, innovation rate) and compare trajectories between systems with and without symbiogenetic norms.
- Insufficient exploration of parameter regimes and sensitivity analyses in 1D systems (e.g., norms A/B/C/D, initial density, mutation rates, boundary conditions); systematically map the phase space to identify regions that support sustained diversity and complex dynamics.
- 2D extension is demonstrated only qualitatively and with limited norms (0 and D); perform quantitative analyses (e.g., spatial autocorrelation, domain growth rates, motif lifetimes, lineage spread) across norms and initial conditions to establish what is genuinely novel about 2D dynamics.
- Absence of long-duration experiments and statistical replication for claims about “organized homogeneity” vs. sustained dynamics; run ensembles over much longer timescales, report outcome distributions, and test how heterogeneity of norms and patchy environments alter asymptotic behaviors.
- No explicit evaluation of emergent recombination, parasitism, and self-repair in the new implementations; reproduce and quantify these phenomena with standardized metrics (e.g., crossover detection, host–parasite dependency matrices, repair success rates post-damage).
- Lack of causal attribution for the role of symbiotic norms versus other factors; design ablation studies that isolate each norm’s contribution to evolvability, robustness, and complexity growth.
- The proposed robustness experiments for symbioorganisms are not executed; implement and report survival curves under controlled perturbations (single-collision intruders, stochastic norm noise, resource shocks) and analyze failure modes.
- Information-theoretic analyses are preliminary (Shannon entropy and pairwise mutual information only); extend to transfer entropy, active information storage, and local information dynamics, and scale these analyses to 2D substrates.
- No genealogy or ancestry-tracking instrumentation in the Barricelli systems; add token-tracing or lineage labeling to quantify inheritance patterns, cooperation histories, and the emergence of hierarchical organization.
- The interaction between spatial heterogeneity and norm heterogeneity—cited by Barricelli as prolonging dynamics—is not systematically investigated; construct patchy worlds with controlled norm mosaics and diffusion barriers to quantify their effects on diversity and open-endedness.
- Resource dynamics are abstract or absent (except in the DNA-CA, where budgets are very simple); incorporate explicit resource flows, costs of replication, and competition for limited substrates to evaluate selection pressures and ecological feedbacks.
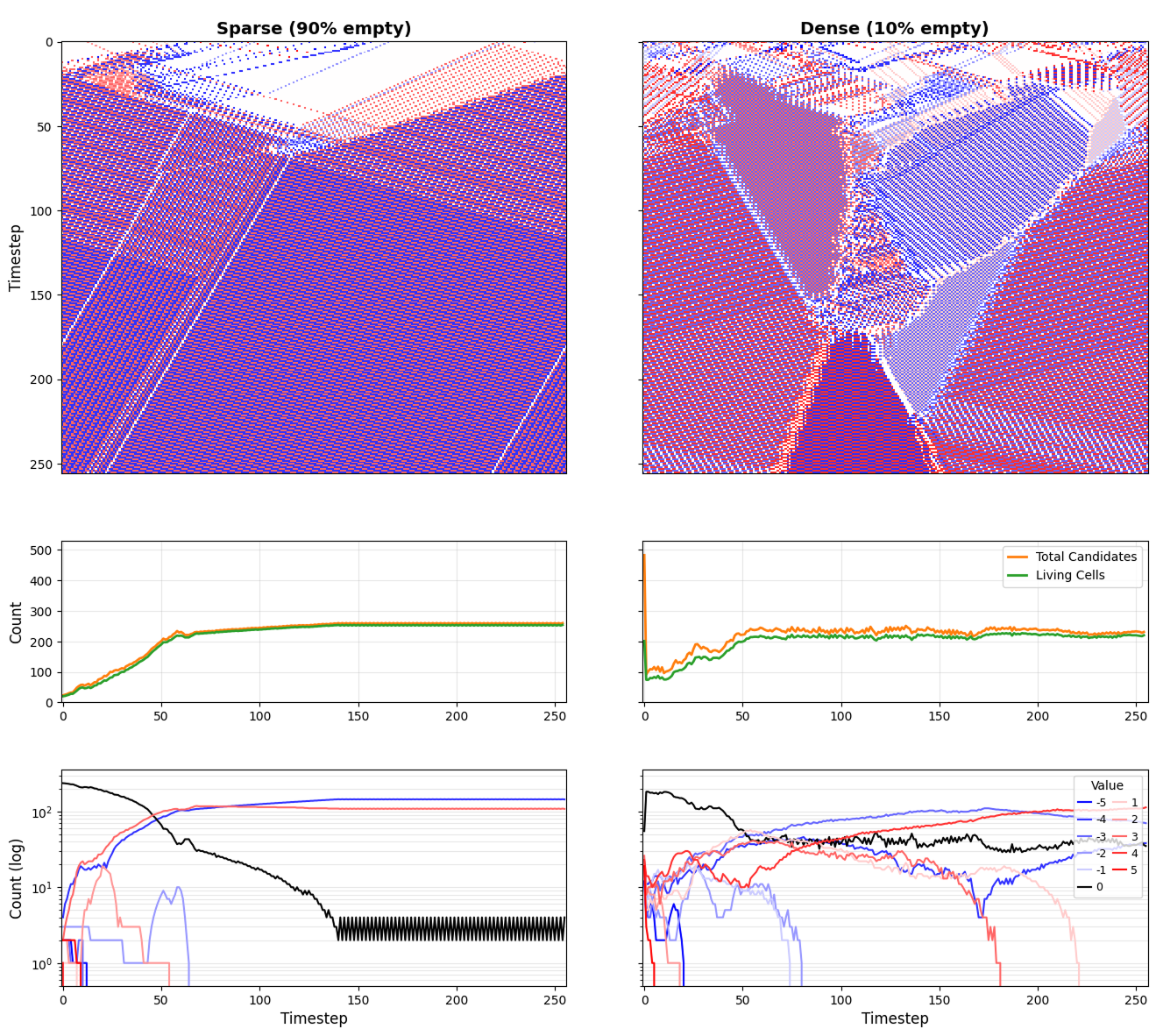
- The Boolean CA “gated Rule 110” demonstration is only illustrative; parameterize the gating norm, quantify minimal conditions for persistent “Boolean symbioorganisms,” and compare to ungated baselines to establish the necessity and sufficiency of the symbiogenetic gate.
- No cross-model comparisons to other ALife platforms (e.g., Tierra, Avida, Lenia) to isolate what symbiogenesis uniquely contributes; run controlled benchmarks with matched mutation/selection parameters and report differential outcomes in complexity metrics.
- The DNA-norm model is minimal and biophysically schematic; incorporate more realistic features (e.g., binding energies, mismatch penalties, secondary structure, kinetics, temperature/energy budgets) to test whether observed motif reuse and spatial heredity are robust to physical constraints.
- Motif-reuse metric ( with threshold ) is a coarse proxy for heredity and replication; complement with direct measures of strand-level reproduction, error rates, duplication fidelity, and heritable functional traits to avoid confounds from random repetition.
- Limited exploration of threshold sensitivity (e.g., ) and motif length choices; provide robustness checks across , , population sizes, and mutation rates, and report confidence intervals and effect sizes for all comparisons.
- The DNA-norm CA shows spatial domains but lacks demonstrated selection on function; introduce genotype-to-phenotype mappings (e.g., sequence-dependent catalytic sites, resource modulation) to test whether functionally encoded traits feed back to replication success.
- Unclear necessary and sufficient conditions for “split” (replication) in the DNA-norms; specify and vary completion thresholds, gap-filling criteria, and annealing requirements to determine minimal requirements for sustainable self-replication.
- No investigation of multi-part integration in the DNA-norm setting (composite genomes, modular assembly); add rules for assembling/routing multi-strand complexes and test whether symbiogenesis yields new higher-level reproductive units.
- Absence of coevolutionary dynamics (e.g., host–parasite arms races) in both numerical and DNA-norm systems; introduce parasitic sequences and measure reciprocal adaptation, diversification rates, and Red Queen dynamics.
- The link between symbiogenesis and collective intelligence remains conceptual; define task environments (e.g., distributed sensing/control, game-based selection), specify performance metrics, and test whether symbiotic collectives outperform solitary entities under identical resource budgets.
- No exploration of transfer across substrates (neural networks, neural cellular automata, program interpreters) despite being proposed; implement analogous symbiogenetic norms in these substrates and compare evolvability and open-endedness side-by-side.
- Missing evaluation of scalability and computational cost; report time/space complexity, parallelization strategies, and practical limits for long, high-dimensional runs to enable reproducible large-scale studies.
- Reproducibility is only partially addressed; provide full parameter sweeps, seeds, data outputs, and analysis scripts (including random seeds and versioned configurations) to enable independent verification of reported phenomena.
- Lack of theoretical analysis of when symbiosis becomes evolutionarily stable in these models; develop mathematical frameworks (e.g., game-theoretic/payoff matrices derived from norms, dynamical systems analysis) to predict transitions from transient cooperation to obligate integration.
- No explicit tests for emergence of multilevel selection and individuality; instrument and quantify when collectives exhibit causal closure, alignment of function, and agency-like behavior, and relate these to symbiogenesis events.
- Unclear generality across dimensions; evaluate whether 3D substrates or different neighborhood topologies change the prevalence or stability of symbiogenesis, and whether new phenomena (e.g., volumetric compartmentalization) emerge.
Practical Applications
Immediate Applications
Below are specific, deployable use cases that draw directly from the paper’s implemented systems (1D/2D Barricelli automata, DNA-norm “soup” and CA, robustness experiments, and information-theoretic metrics) and released code.
- Research toolkit for artificial life and evolutionary computation (Academia, Software)
- What: Use the open-source repo to replicate Barricelli’s 1D/2D systems, run DNA-norm simulations, and compute entropy/mutual information to study emergent replication, parasitism, and self-repair.
- Tools/workflows: Codebase as a baseline benchmark; add-on notebooks to quantify open-endedness (e.g., motif-repetition curves, mutual information matrices), robustness assays (inject disturbances, log survival rates).
- Assumptions/dependencies: Python environment and modest compute; researchers comfortable with CA/ALife paradigms.
- Curriculum modules and interactive labs on symbiosis and major transitions (Education)
- What: Classroom labs demonstrating how symbiogenesis yields complex structures from simple rules; illustrate mutualism, parasitism, and commensalism dynamics.
- Tools/products: Browser-based visualizers of 1D/2D runs; step-by-step assignments (e.g., compare single-rule vs patchy “heterogeneous norm” environments).
- Assumptions/dependencies: Educator-facing documentation; access to basic computing in classrooms.
- Generative art and public science exhibits from 2D symbiogenetic CA (Creative tech, Museums)
- What: Visualize symmetry, waves, and domain growth from symbiotic rules as dynamic artworks or explorable museum pieces.
- Tools/products: Real-time CA visualizer; exportable image/video pipelines; interactive kiosks.
- Assumptions/dependencies: GPU-accelerated rendering optional; licensing of code under permissive terms.
- Symbiotic operators for evolutionary algorithms and AutoML (Software, AI/ML)
- What: Incorporate “norm patches,” symbiotic crossover, and parasitism-inspired pressures to reduce premature convergence and promote modular composition.
- Tools/workflows: Plugins for evolutionary strategies/GP/NAS frameworks that: (1) run quasi-uniform “patch” selection pressures, (2) evaluate team-of-gene performance, (3) encourage emergent modular recombination.
- Assumptions/dependencies: Mapping of CA norms to algorithmic operators; evaluation on standard opt/NAS benchmarks.
- Testbeds for multi-agent interaction regimes (AI/ML, Robotics R&D)
- What: Use symbiogenetic gating and collision norms as simple, reproducible testbeds to study cooperation/competition balances and collective problem-solving.
- Tools/workflows: Integrate with PettingZoo or MARL libraries; run ablations on mutualism/commensalism/parasitism-inspired reward sharing; measure information flow.
- Assumptions/dependencies: Transfer from grid-world CA dynamics to multi-agent learning requires careful reward shaping.
- Open-endedness and emergence analytics for complex systems logging (Software ops, Research)
- What: Apply motif repetition and information-theoretic metrics (entropy, mutual information) to detect repeated interaction motifs and emergent structure in event streams (e.g., microservices traces, agent logs).
- Tools/workflows: A metrics library that ingests time-series/logs and outputs motif repetition curves and MI heatmaps over time.
- Assumptions/dependencies: Requires well-segmented, windowable logs; careful interpretation to avoid false positives.
- In silico pre-screening for DNA-based assembly protocols (Synthetic biology R&D)
- What: Use the DNA-norm “soup” to quickly assess how elongation, complementarity, and splitting rules impact k-mer reuse and duplex domain growth before wet-lab trials.
- Tools/workflows: Parameter sweeps on error rates, resource budgets, and overlap thresholds; identify regimes that maximize long-motif repetition.
- Assumptions/dependencies: Abstract rules simplify chemistry; results serve as coarse guidance, not direct lab recipes.
- Robustness benchmarking of self-repair dynamics (Software resilience, Cyber-physical R&D)
- What: Adapt the paper’s “inject a disturbance and measure survival” protocol to stress-test self-healing mechanisms in distributed systems or rule-based controllers.
- Tools/workflows: Fault-injection harnesses that emulate collisions; survival/time-to-recovery metrics analogous to symbioorganism robustness measures.
- Assumptions/dependencies: Mapping numerical “organisms” to services/modules requires instrumentation of real systems.
- Game mechanics inspired by symbiogenesis (Games/EdTech)
- What: Design games where players/agents can merge, parasitize, or form alliances, with emergent “organism-level” abilities and self-repair.
- Tools/products: Rule engines implementing symbiotic interactions; educational gameplay demonstrating ecological and collective-intelligence principles.
- Assumptions/dependencies: Balancing asymmetric interactions (e.g., parasitism that improves diversity) for fun and fairness.
- Conceptual training for organizational design and policy workshops (Policy, Management education)
- What: Use the paper’s continuum (mutualism↔parasitism) and “collective as individual” framing to design exercises on team-of-teams, mergers, and cooperative competition.
- Tools/workflows: Scenario simulations using simplified CA/agent models; debriefs linking results to institutional governance.
- Assumptions/dependencies: Translation from abstract models to organizational contexts requires facilitation and case mapping.
Long-Term Applications
These opportunities build on the paper’s methods but require further research, validation, scaling, or engineering integration.
- Wet-lab origin-of-life and protocell experiments guided by DNA-norm regimes (Healthcare/Basic science)
- What: Translate parameters that boost long-motif repetition and spatial lineage formation into microfluidic or surface-templated RNA/DNA experiments.
- Tools/workflows: Lab protocols testing overlap-mediated assembly and splitting; compare to in silico motif-repetition thresholds.
- Assumptions/dependencies: Accurate kinetic/thermodynamic models; enzymatic or non-enzymatic chemistries with controllable error rates.
- Self-assembling DNA computing and materials using minimal rules (Synthetic biology, Materials)
- What: Engineer strand-displacement systems where complementary association and splitting implement reproducible, modular assemblies and self-repair.
- Tools/products: DNA tile circuits or origami that “grow,” split, and recombine into higher-order structures; programmable self-healing materials.
- Assumptions/dependencies: Control over hybridization kinetics, error suppression, and energy budgets.
- Symbiogenetic swarm robotics with emergent “super-agents” (Robotics)
- What: Develop local interaction rules enabling robots to fuse capabilities (sensing/actuation/power), self-repair, and split, akin to CA symbioorganisms.
- Tools/workflows: Docking hardware, local comms, rule interpreters mapping “norms” to merge/split behaviors; field testing under disturbances.
- Assumptions/dependencies: Robust hardware interfaces and safety protocols; reliable local decision-making under uncertainty.
- Open-ended, modular AI via symbiotic composition (AI/ML, Software)
- What: Architect ecosystems of models/tools that can merge competencies, recombine (emergent crossover), and resist parasitism while sustaining diversity (heterogeneous “norm patches”).
- Tools/workflows: Orchestration platforms for agent mergers/splits; fitness via task wins (à la Barricelli’s game selection), MI-based emergence monitors.
- Assumptions/dependencies: New evaluation metrics, alignment/safety safeguards, and scalable credit assignment.
- Self-healing distributed systems inspired by symbiosis (Cloud/Edge, Energy)
- What: Apply “norm patches” to service meshes/microgrids so local rules favor cooperative replication of robust configurations and isolation of parasitic patterns.
- Tools/products: Controllers that detect repeated, high-fitness interaction motifs and promote them across the network; parasitism throttling policies.
- Assumptions/dependencies: Observability at the right granularity; provenance tracing to identify “organism-level” modules.
- Microbiome consortia engineering with controlled interaction types (Healthcare, Agriculture)
- What: Design community interventions that include mutualists and strategic “parasites”/commensals to improve stability and resilience, reflecting ecological findings referenced in the paper.
- Tools/workflows: In silico pre-screening with symbiosis-inspired dynamics; targeted consortia therapies for gut/soil.
- Assumptions/dependencies: Regulatory approvals; robust in vivo monitoring; ethical deployment.
- Market designs and agent-based finance with symbiotic mergers (Finance, Policy)
- What: Simulate markets with agents that can form composite entities when beneficial, exploring stability and innovation trade-offs under parasitism/competition.
- Tools/workflows: ABM platforms incorporating symbiotic composition norms; policy sandboxes for merger rules informed by “collective as individual” insights.
- Assumptions/dependencies: Behavioral calibration; regulatory appetite for experimentation.
- Security via coevolving, symbiogenetic red–blue teaming (Cybersecurity)
- What: Use symbiotic composition and parasitism dynamics to evolve defense coalitions that adapt to evolving threats with self-repair properties.
- Tools/workflows: Simulation arenas with DNA-norm-like recombination of tactics; selection via game outcomes as in Barricelli’s TacTix framing.
- Assumptions/dependencies: Careful containment; mapping abstract norms to TTPs (tactics, techniques, procedures).
- Urban/ecosystem planning anchored in interaction diversity (Public policy, Ecology)
- What: Use the paper’s evidence and cited works on unilateral interactions to design policies that mix mutualism, commensalism, and controlled antagonism for resilient communities.
- Tools/workflows: Planning simulations with “patchy norms” representing governance styles; measure stability with information-theoretic metrics over time.
- Assumptions/dependencies: Interdisciplinary data and validation; stakeholder engagement.
- Programmable matter and modular hardware with CA-like rule execution (Hardware, Manufacturing)
- What: Implement hardware modules that interact via local rules to assemble larger structures and heal after damage, inspired by 2D CA symbiogenesis.
- Tools/products: Smart connectors, embedded rule evaluators, local sensing; self-configuring assemblies for rapid deployment/repair.
- Assumptions/dependencies: Advances in low-power compute, standardized connectors, safe merging/splitting.
- Collective-intelligence governance and “superagent” institutions (Policy, Management)
- What: Formalize mergers/alliances of teams or orgs as symbiogenetic transitions to enhance problem-solving capacity, with metrics for integration and resilience.
- Tools/workflows: Decision-support dashboards tracking “emergent module” health via information flow and motif repetition; playbooks for integration.
- Assumptions/dependencies: Organizational readiness; translating technical metrics to human systems.
- AI safety via modular, evolvable coalitions (AI governance)
- What: Explore whether symbiogenetic, multi-agent architectures can align local incentives to produce higher-level aligned behavior, with parasitism detection and containment.
- Tools/workflows: Safety monitors for coalition composition; intervention norms to prevent runaway monoliths.
- Assumptions/dependencies: Formal alignment objectives; empirical evidence that composition reduces risk.
Cross-cutting dependencies and assumptions
- Abstraction gap: CAs and DNA-norm simulations are stylized; mapping to real-world biochemistry, robotics, or institutions requires careful translation and validation.
- Measurement: The provided metrics (entropy, mutual information, motif repetition) are useful but need domain-specific baselines to avoid overinterpreting “emergence.”
- Heterogeneity matters: Many benefits arise from quasi-uniform, patchy norms; real deployments need mechanisms to sustain rule diversity without fragmentation.
- Safety and ethics: Emergent composition and parasitism dynamics can have unintended consequences (e.g., in bio, AI, markets); governance frameworks must co-evolve with technology.
Glossary
- Active information storage: An information-theoretic quantity indicating how much past information a system retains that is useful for predicting its future state. "explore the concepts of transfer entropy and locations of active information storage"
- Amensalism: An ecological interaction where one species is harmed while the other is unaffected. "commensalism and amensalism"
- Annealing (DNA strands): The process by which complementary nucleic acid strands bind through base pairing to form double-stranded structures. "two strands to anneal when they share a sufficiently long contiguous complementary sequence"
- atan2: A two-argument arctangent function used to compute the polar angle from Cartesian coordinates. "by calculating it's polar angle with "
- Autocatalytic processes: Chemical reactions where a product acts as a catalyst for its own formation, enabling self-amplifying replication. "the capacity to reproduce by autocatalytic processes"
- Barricelli's collision norms: Local rules that resolve replication collisions in numerical cellular automata by specifying mutation or exclusion outcomes. "Barricelli's collision norms."
- Brainfuck family (BFF) programs: Programs written in Brainfuck-like esoteric languages used as a computational substrate to study emergent behaviors. "random Brainfuck family (BFF) programs"
- Causal closure: A condition where a system’s internal interactions sufficiently account for its behavior without external causal dependencies. "emerges when a collective achieves sufficient integration, causal closure, and alignment of function"
- Cellular Automata (CA): Discrete computational systems with grid-based cells updating by local rules, often used to study complex dynamics. "In his Cellular Automata (CA) models"
- Collective intelligence: The emergent problem-solving or adaptive capacity of groups arising from interactions among their components. "collective intelligence"
- Complementarity: The base-pairing rule that defines which nucleotides pair (A–T, C–G), enabling faithful template-directed replication. "Complementarity is defined by an involution:"
- Complementary association: A DNA-norm where complementary bases pair and overlapping fragments merge to extend strands. "Complementary association (pairing and overlap-mediated merging)."
- Connectionist perspective: An approach interpreting complex behavior as emerging from networks of interacting simple units. "From a connectionist perspective, such transitions can be interpreted as"
- Crossover: A recombination process where genetic material is exchanged between strands, facilitating variation. "emergent sexual recombination via crossover"
- DNA-norms: Local interaction rules inspired by DNA mechanisms (elongation, pairing, splitting) used to govern replication in simulations. "preliminary experimentation with DNA-norms."
- Duplex (double-stranded DNA): A two-stranded nucleic acid structure formed by complementary base pairing. "these duplexes split and place offspring into adjacent cells."
- Endosymbiosis: A symbiotic relationship where one organism lives inside another, often leading to integrated lineages. "modeled when endosymbiosis becomes advantageous."
- Evolutionary computation: A family of optimization and design methods inspired by natural evolution, employing variation and selection. "Example of evolutionary computation and embodiement proposed by Barricelli."
- Game-theoretic results: Analytical outcomes based on mathematical models of strategic interaction among agents. "Game-theoretic results shows also that when agents decide to merge"
- Genetic operators: Abstract procedures (e.g., mutation, crossover) that modify genetic representations in evolutionary algorithms. "Holland's work on genetic operators"
- Genetic programming: An evolutionary computation method where computer programs evolve to solve tasks. "in genetic programming"
- Hierarchical organization: Multi-level structuring where higher-level units emerge from coordinated lower-level components. "allowing hierarchical organization"
- High-order entropy: An entropy measure capturing dependencies beyond single-symbol statistics, used to assess emergent structure. "high-order entropy as a measure of emergent complexity"
- Holobiont theory: The view that hosts and their microbiomes form integrated ecological and evolutionary units. "holobiont theory"
- Holobionts: Host–microbiome collectives considered as unified functional entities. "functional systems sometimes described as holobionts"
- k-mer: A contiguous nucleotide sequence of length k, used to analyze motifs and repetition in genomes. "dominant -mer"
- Lenia: A continuous cellular automaton framework known for lifelike, self-organizing patterns. "in Lenia"
- Major evolutionary transitions: Transformative events where independent entities combine into new higher-level individuals (e.g., cells to multicellular organisms). "major evolutionary transitions"
- Metastable patterns: Temporarily stable configurations maintained by ongoing interactions rather than static equilibria. "metastable patterns of interaction"
- Mutual information: A measure of statistical dependence between variables or system states across time. "mutual information between states at all time-steps"
- Mutualism: A symbiotic interaction where both partners benefit. "mutualism, commensalism, amensalism, competition"
- Nucleotides: Molecular building blocks of nucleic acids (A, C, G, T) with specific pairing rules. "nucleotides have specific association rules"
- Open-ended evolution: Evolutionary dynamics that continually generate novel forms and increasing complexity without a fixed endpoint. "open-ended evolution"
- Organized homogeneity: A dynamical outcome where one simple structure dominates, leading to periodic, uniform behavior. "organized homogeneity"
- Overlap-mediated merging: The joining of nucleic acid fragments through alignment of complementary overlapping regions. "overlap-mediated merging"
- Parasitism: An interaction where one organism benefits at the expense of another. "parasitism emerges"
- Periodic boundary conditions: A topology where edges wrap around, making the space effectively loop (e.g., a ring). "periodic boundary conditions"
- Polarity (5' and 3' ends): Directionality of nucleic acid strands from the 5' to 3' ends, constraining growth and pairing. "a fixed polarity (a 5' and 3' end)"
- Quasi-uniform CA: A cellular automaton where different local rules apply in different spatial patches. "quasi-uniform CA where different rules are applied in different patches"
- Replicators: Entities capable of making copies of themselves, foundational to evolutionary processes. "kickstart the emergence of replicators."
- Rule 110: A one-dimensional cellular automaton rule known for computational universality and complex behavior. "rule 110"
- Self-organization: Spontaneous pattern formation driven by local interactions without centralized control. "self-organization and interaction patterns"
- Self-repairing organisms: Systems capable of restoring function or structure after damage through intrinsic processes. "self-repairing organisms would emerge"
- Serial Endosymbiosis Theory (SET): Margulis’s theory that eukaryotic organelles originated via sequential endosymbiotic events. "Serial Endosymbiosis Theory (SET)"
- Shannon entropy: A measure of uncertainty or information content of a random variable or system state distribution. "the
Shannon entropyis calculated" - Shift-based replication: A replication mechanism where numerical genes copy to positions shifted by their values, central to Barricelli’s CA. "shift-based replication and collision norms"
- Symbiogenesis: The formation of new evolutionary individuals by the integration of distinct organisms through symbiosis. "symbiogenesis"
- Symbioorganism: A higher-order self-replicating structure formed by interacting numerical genes in Barricelli’s framework. "Barricelli's symbioorganism."
- Symbiotic coevolutionary approaches: Methods where co-adaptation of interacting components is structured around symbiotic relationships. "symbiotic coevolutionary approaches"
- Symbiotic Organism Search (SOS) algorithms: Optimization methods inspired by symbiotic interactions among organisms. "Symbiotic Organism Search (SOS) algorithms"
- TacTix: A combinatorial game used by Barricelli to impose competitive selection pressures in evolution experiments. "TacTix"
- Token-tracing methodology: A technique to track ancestry and lineage of computational genes or tokens over time. "token-tracing methodology"
- Transfer entropy: A directional information measure capturing how much one process predicts future states of another. "transfer entropy"
- Turing-complete: Capable of performing any computation given sufficient time and memory, equivalent to a universal Turing machine. "render RNA Turing-complete"
- Universal constructor: A theoretical machine that can construct any machine, including a copy of itself, within a cellular automaton. "a universal constructor"
- Wilson confidence intervals: A statistically robust interval estimation method for binomial proportions. "95\% Wilson confidence intervals"
- Toroidal wrapping: A topology where opposite edges are connected to form a torus, avoiding boundaries. "toroidal wrapping"
- Well-mixed (spaceless) setting: A model assumption where interactions occur uniformly without spatial constraints. "well-mixed (spaceless) setting"
Collections
Sign up for free to add this paper to one or more collections.