- The paper introduces VISTA-PATH, an interactive model leveraging a large, ontology-driven dataset to perform class-aware pathology segmentation.
- It integrates visual, semantic, and spatial cues through cascaded cross-attention, achieving significant Dice score improvements over existing segmentation models.
- The method employs human-in-the-loop refinement to enhance clinical biomarker discovery and survival prediction, notably in colorectal cancer.
VISTA-PATH: Interactive Foundation Model for Pathology Image Segmentation and Quantitative Analysis
Introduction
The paper "VISTA-PATH: An interactive foundation model for pathology image segmentation and quantitative analysis in computational pathology" (2601.16451) introduces VISTA-PATH: an interactive, class-aware segmentation foundation model tailored for routine and complex histopathology images. Grounded in a large-scale, ontology-driven segmentation dataset and leveraging joint conditioning on visual evidence, semantic tissue descriptors, and spatial prompts, VISTA-PATH resolves multi-tissue heterogeneity and supports dynamic human-in-the-loop refinement, elevating segmentation from static prediction to interactive and clinically meaningful representation.
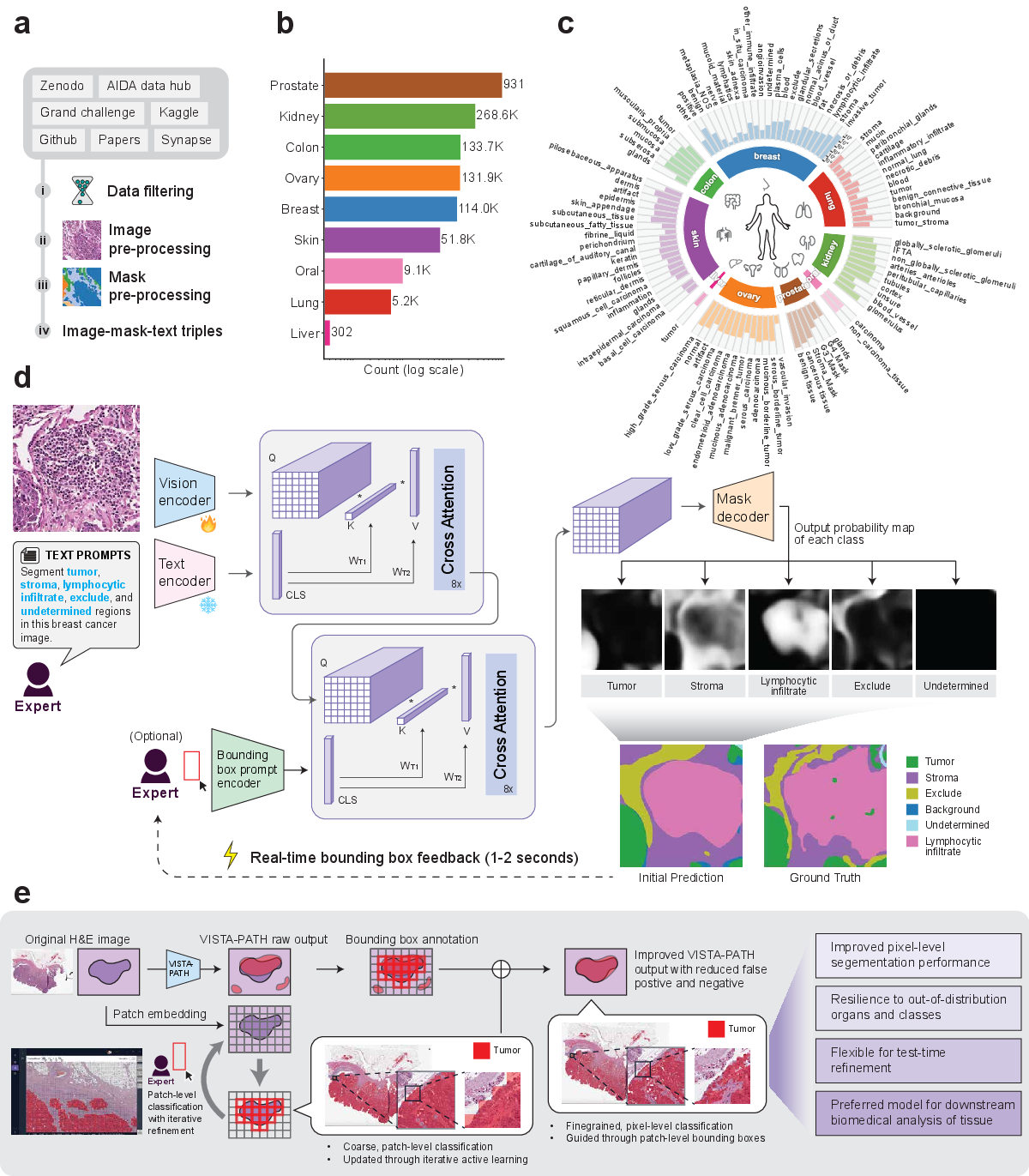
Figure 1: Overview of the VISTA-PATH dataset and architecture, highlighting data curation, semantic organization, model workflow, and interactive human-in-the-loop refinement cycle.
VISTA-PATH Data: Construction and Composition
VISTA-PATH Data aggregates 1,645,706 high-resolution pathology image–mask–text triplets spanning 9 organs and 93 tissue classes. This corpus assimilates resources from 22 publicly available datasets, encompassing tissue-level and instance-level dense annotations. The dataset is structured with a manually constructed tissue ontology, organizing class labels to enable both intra- and inter-organ analysis. Annotation quality is guaranteed via strict inclusion criteria and unified preprocessing—including rasterization of polygon-based masks into pixel-level formats and standardized image cropping and resizing pipelines.
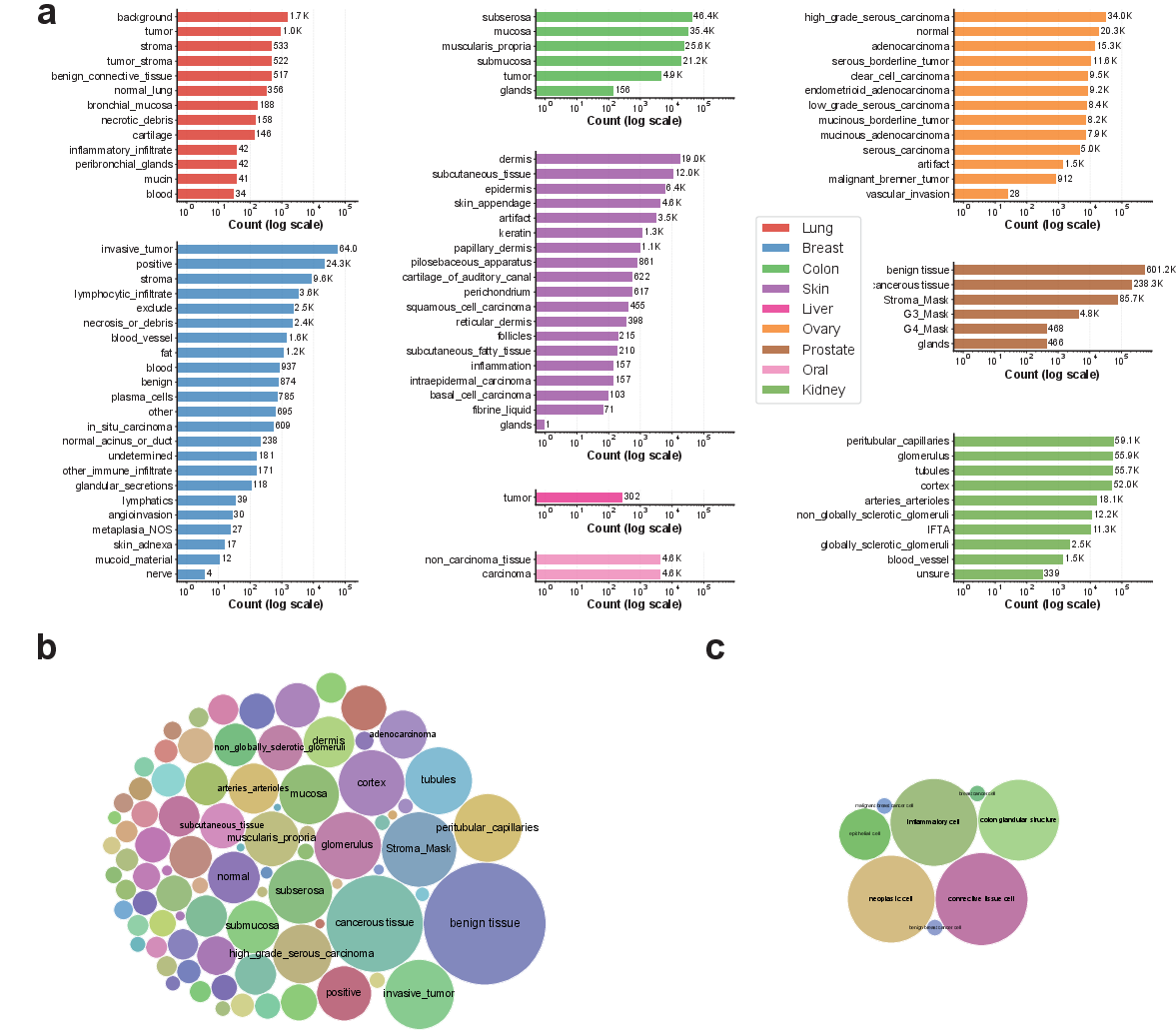
Figure 2: Distribution and semantic coverage of tissue classes in VISTA-PATH Data versus BiomedParse, illustrating the superior scale and diversity of VISTA-PATH.
External benchmarks comprise 66,355 samples from 13 organs and 82 tissue classes, derived from spatial transcriptomics-guided and single-cell–guided protocols (Visium HD, Xenium), as well as independent pathology segmentation datasets. These external datasets diversify annotation paradigms, allowing robust evaluation of model generalization across domain shifts.
Model Architecture and Training
VISTA-PATH integrates four principal modules: a trainable PLIP-based visual encoder, a frozen PLIP-based text encoder, a prompt encoder adapted from SAM for bounding-box spatial priors, and a Transformer-based mask decoder. The model fuses visual, semantic, and spatial information via cascaded cross-attention mechanisms, enabling context-aware, class-conditioned predictions. For each image–class pair, VISTA-PATH predicts a binary segmentation mask, optionally refined by bounding-box spatial prompts during training and inference.
The model is trained under class-conditioned supervision, utilizing mixed-precision and AdamW optimization on NVIDIA H200 GPUs. Bounding-box prompts are randomly generated and dropped during training to enhance robustness and avoid over-reliance on spatial priors.
Held-out and external evaluations demonstrate superior performance of VISTA-PATH compared to leading segmentation foundation models (MedSAM, BiomedParse, organ-specific Res2Net). Across 21 internal datasets, VISTA-PATH achieves a macro-average Dice of 0.772, compared to 0.698 (VISTA-PATH w/o box), 0.581 (MedSAM), 0.379 (BiomedParse), and 0.521 (Res2Net), with significant gains (2.7–30.9% Dice improvement) spanning all organ types. Notably, performance is robust even in the absence of bounding-box prompts, indicating strong semantic grounding.
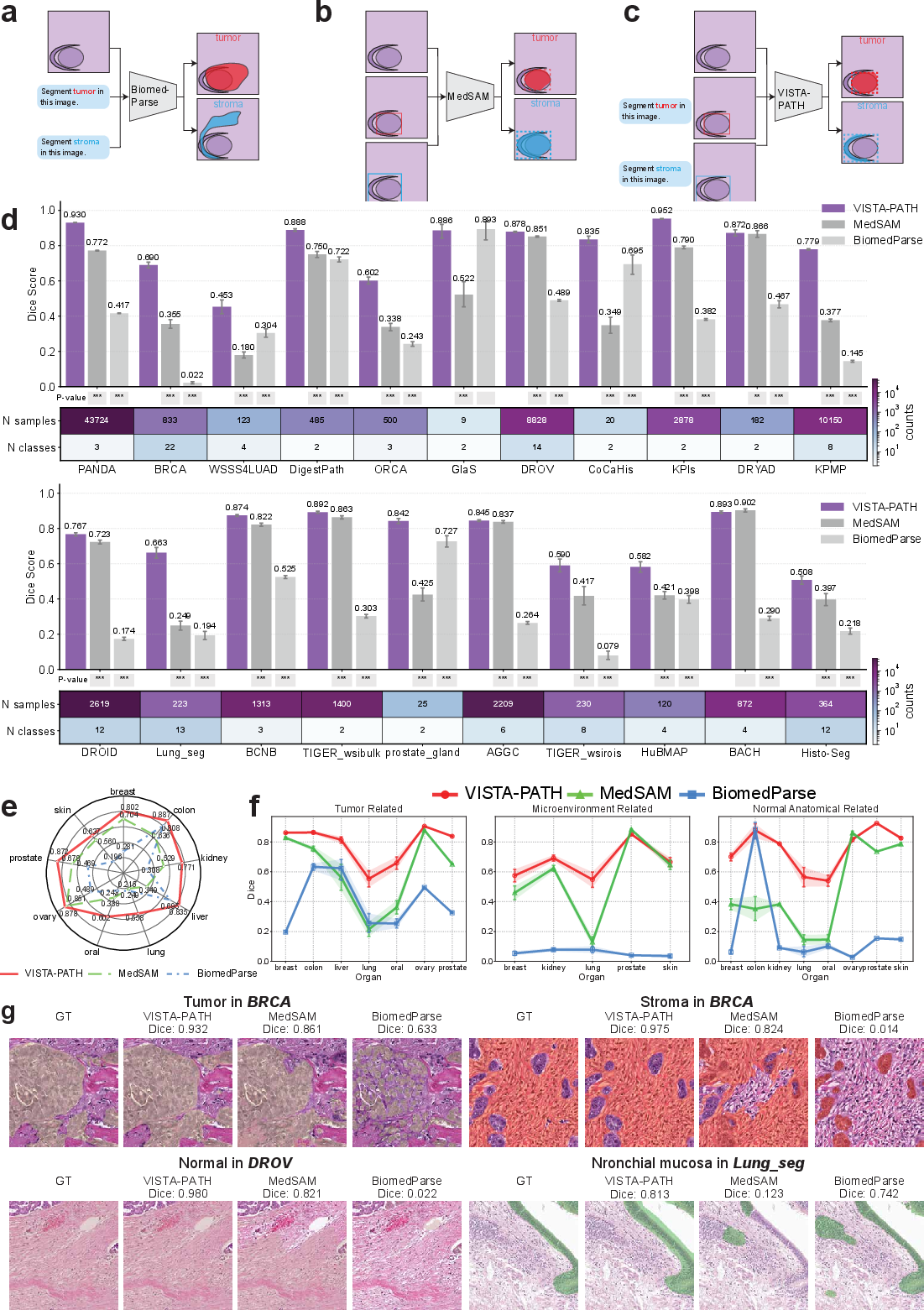
Figure 3: VISTA-PATH generalizes across annotation regimes on external datasets, outperforming baselines on pathology, Visium HD, and Xenium datasets across organs and tissue categories.
On external benchmarks, VISTA-PATH exceeds the second-best model by 3.3–16.3% Dice across 13 organs, with especially pronounced gains on heterogeneous cases (e.g., lung, kidney, colon, skin cancers) and multi-class spatial transcriptomics scenarios.
Interactive Human-in-the-Loop Refinement
VISTA-PATH supports a dynamic human-in-the-loop workflow for whole-slide refinement. Sparse expert bounding-box corrections (typically 10–2,000 patches) are rapidly propagated to global pixel-level segmentations through patch-based active learning, patch classifier retraining, and class-aware bounding-box prompt updates. Iterative refinement reliably increases Dice scores by 15.3–46.8% across nine evaluated datasets, outperforming patch-level segmentation and MedSAM baselines at all stages.
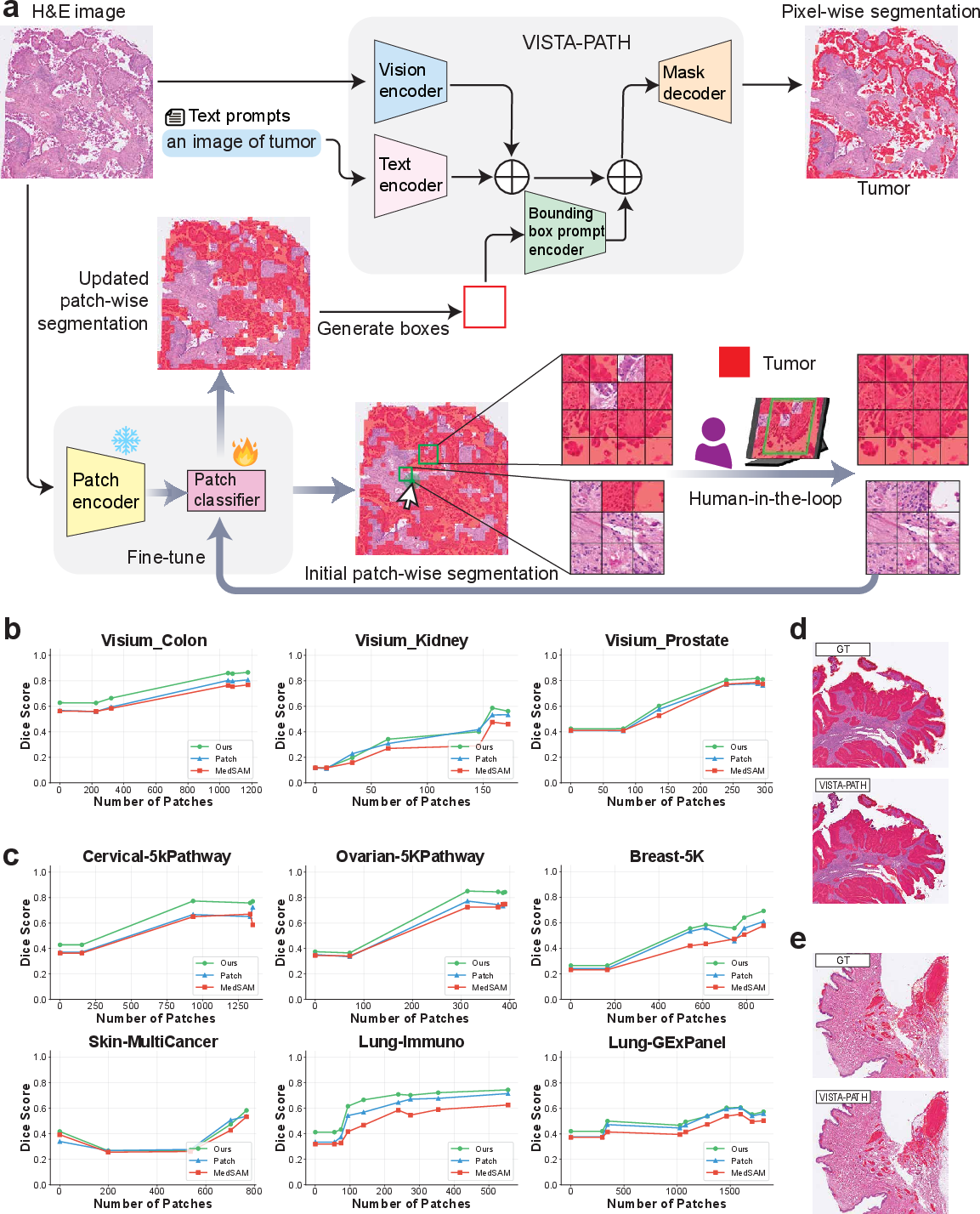
Figure 4: Human-in-the-loop workflow efficiently propagates sparse expert feedback to improved whole-slide segmentation, with performance gains visualized across Visium HD and Xenium datasets.
Downstream Clinical Impact: Tumor Interaction Score
VISTA-PATH segmentation outputs provide high-fidelity, class-aware tissue maps supporting direct biomarker discovery. The Tumor Interaction Score (TIS), combining patch-level and pixel-level tumor localization, quantifies tumor coherence and infiltration as interpretable measures of aggressiveness. Applied to TCGA-COAD colorectal cohort, VISTA-PATH enhances survival prediction C-index by 16.8–20.7% over standard multiple instance learning (ABMIL), with TIS achieving significant associations with patient outcome. The MedSAM-based pipeline and ablation studies demonstrate inferior performance, confirming the utility of VISTA-PATH’s semantic–spatial fidelity.
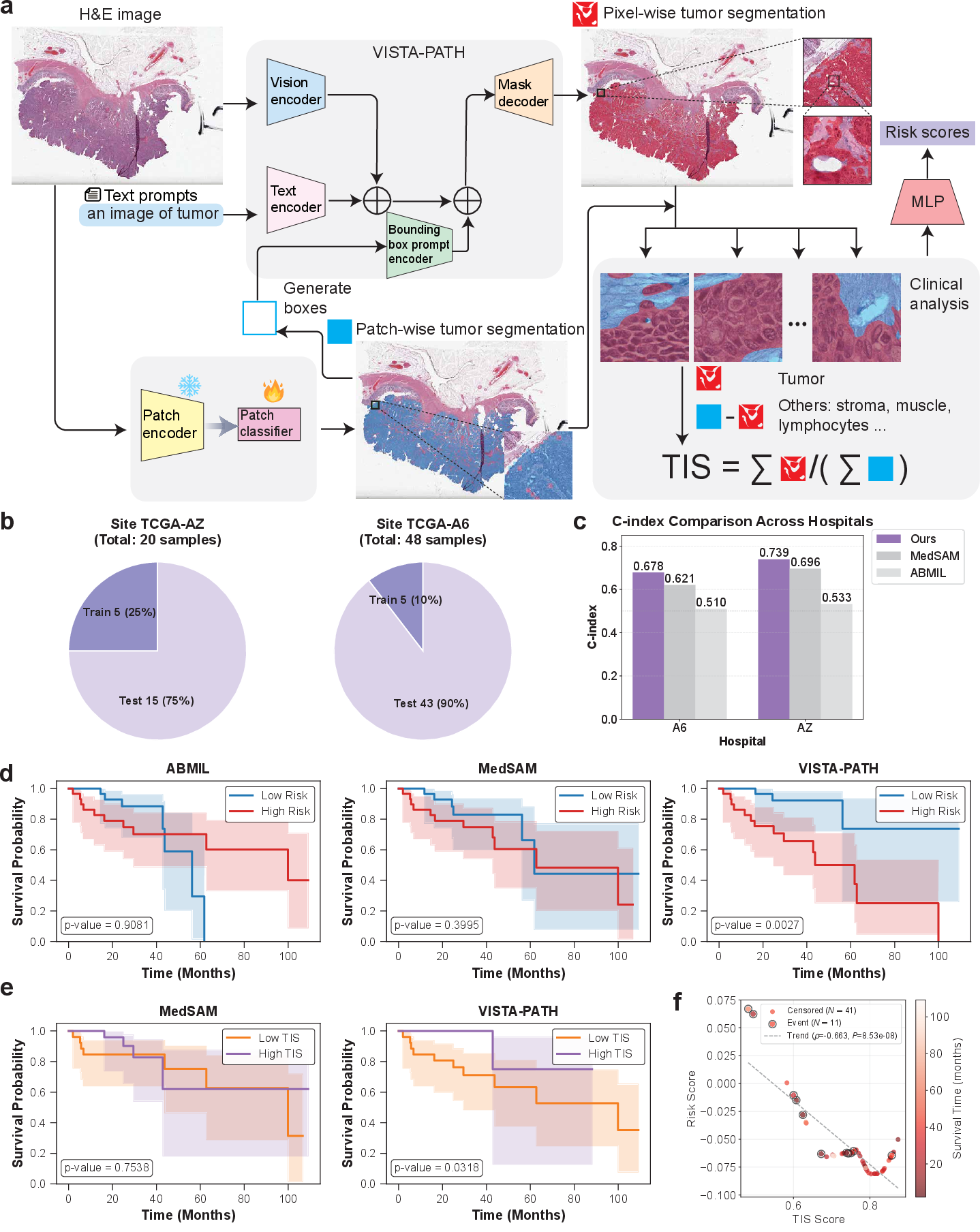
Figure 5: Survival prediction via VISTA-PATH-derived TIS shows strong C-index gains and interpretable risk stratification on the TCGA-COAD cohort.
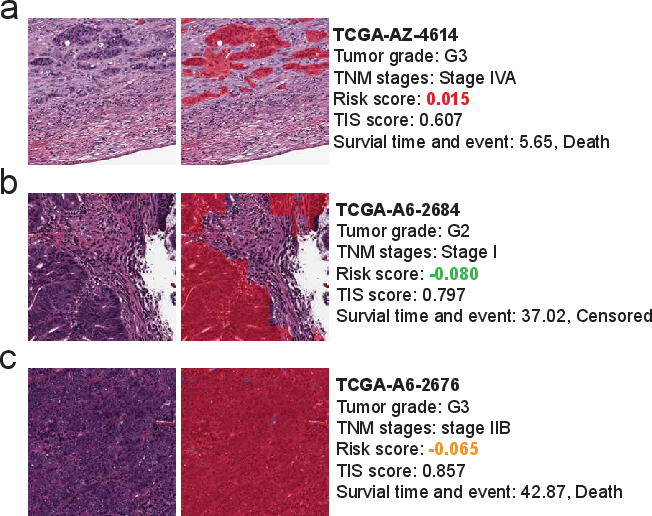
Figure 6: H&E patches with segmentation overlays illustrate the clinical utility of low, intermediate, and high TIS scoring for prognosis.
Implications, Limitations, and Future Directions
VISTA-PATH establishes an interactive segmentation paradigm where pixel-level predictions are jointly informed by semantic prompts and spatial priors, rather than static one-shot inference. This approach supports robust generalization to unseen tissues, annotation protocols, and molecular–spatial domains, while operationalizing segmentation into clinical biomarkers. The model’s design is compatible with agentic LLM-based systems, facilitating integration into multimodal diagnostic workflows.
The study acknowledges several open challenges: extending interactive protocols to richer expert input (e.g., region marking, uncertainty annotation), systematic, automated biomarker discovery from segmentation substrates, large-scale multi-institutional validation for real-world deployment, and optimization across magnification and scanner variability. Addressing these directions will be essential for translating interactive segmentation foundation models like VISTA-PATH into precision pathology practice.
Conclusion
VISTA-PATH advances pathology image segmentation from static algorithmic prediction to interactive, clinically grounded representation. The joint semantic–spatial modeling, large-scale ontology-driven data, efficient human-in-the-loop refinement, and translation to outcome-linked biomarkers collectively position VISTA-PATH as a preferred platform for computational pathology research and clinical translation. These contributions reframe segmentation as a core substrate for both expert reasoning and downstream quantitative analysis in digital pathology.







