- The paper introduces a deep learning framework that segments and predicts biological spine age using 3D MRI data from over 17,000 individuals.
- The methodology employs nnUNet-based segmentation, DCNN age prediction, and bias correction, achieving an R² of 0.85 and an MAE of approximately 3.67 years.
- The findings demonstrate that the spine age gap is a clinically relevant biomarker linked to degenerative conditions and modulated by lifestyle factors.
Artificial Intelligence-Based Measurement of Human Spine Aging from MRI: Methods, Results, and Implications
Introduction
This work presents a deep learning framework for quantifying human spine aging using sagittal T2-weighted magnetic resonance imaging (MRI). The spine, comprising cervical, thoracic, and lumbar regions, exhibits both normal age-related degeneration and pathological changes, often discernible via imaging. Accurate estimation of biological spine age, distinct from chronological age, holds clinical promise for identifying individuals at higher risk for adverse spinal conditions and for tracking lifestyle or therapeutic interventions on spine health.
Methodology
A large-scale dataset of 18,070 3D whole-spine MRI series was compiled from 17,394 individuals (ages 25–84), sourced across 10 North American clinics and spanning 13 years. Rigorous eligibility filtering was performed using a data-driven approach: spine structural and degenerative conditions were vectorized from radiology reports (yielding 215-pathology features per case), aggregated by region and severity, reduced in dimensionality via UMAP, and clustered by HDBSCAN with a 15% population threshold to define "normal" versus "abnormal" aging profiles.
The model pipeline consists of three principal stages: semantic segmentation of the spine using a nnUnet-based architecture, masking to isolate spinal anatomy from surrounding tissues, and age prediction using a deep convolutional neural network (DCNN) composed of stacked 3D convolutional, batch-normalization, and pooling layers, followed by a dense prediction head.
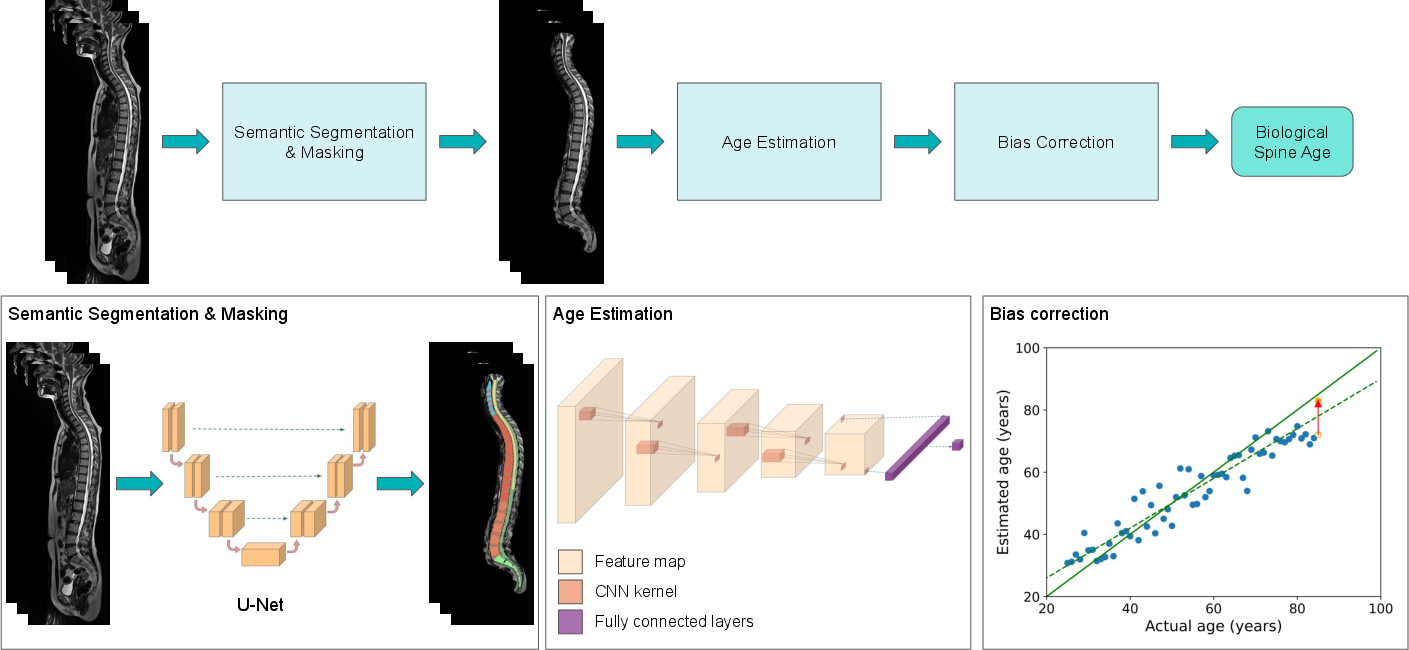
Figure 1: Schematic of pipeline: nnUnet-based segmentation, region masking, and DCNN-based age prediction with bias correction.
The predicted ages were subsequently bias-corrected using Cole’s method to mitigate regression-to-the-mean effects.
Experimental Protocol
The model was trained exclusively on "normal spine" MRI series, totaling 10,611 cases, with stratified splits by age and gender to form development, validation, and test cohorts. Model variants were compared in ablation studies varying training set size, region masking (cervical, thoracic, lumbar, or whole spine), and loss function (MSE vs. smooth-L1). Weighted mean absolute error (WMAE), MAE, and R2 metrics quantified performance, with repeat-scan stability assessed via intraclass correlation.
Quantitative Results
The primary DCNN trained on the entire spine and maximal sample size achieved R2=0.85 post-bias-correction, a substantial advance over prior classical ML approaches (R2=0.28) (2511.17485). MAE and WMAE were 3.67 and 3.60 years, respectively. Increasing sample size yielded monotonic improvements in R2, and lumbar-region-only models moderately lagged whole-spine models, underscoring the multi-regional nature of spine aging.
Repeat scans reflected an intraclass correlation coefficient of 0.73, indicating reasonable stability in predictions over time gaps averaging 1.6 years.
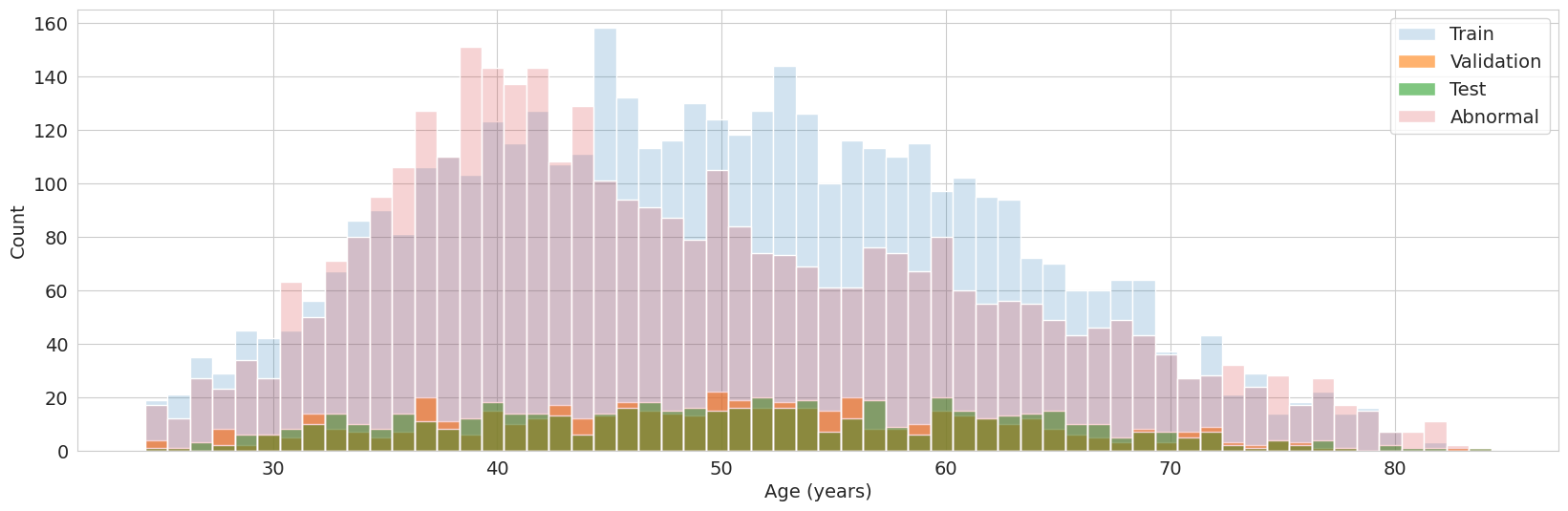
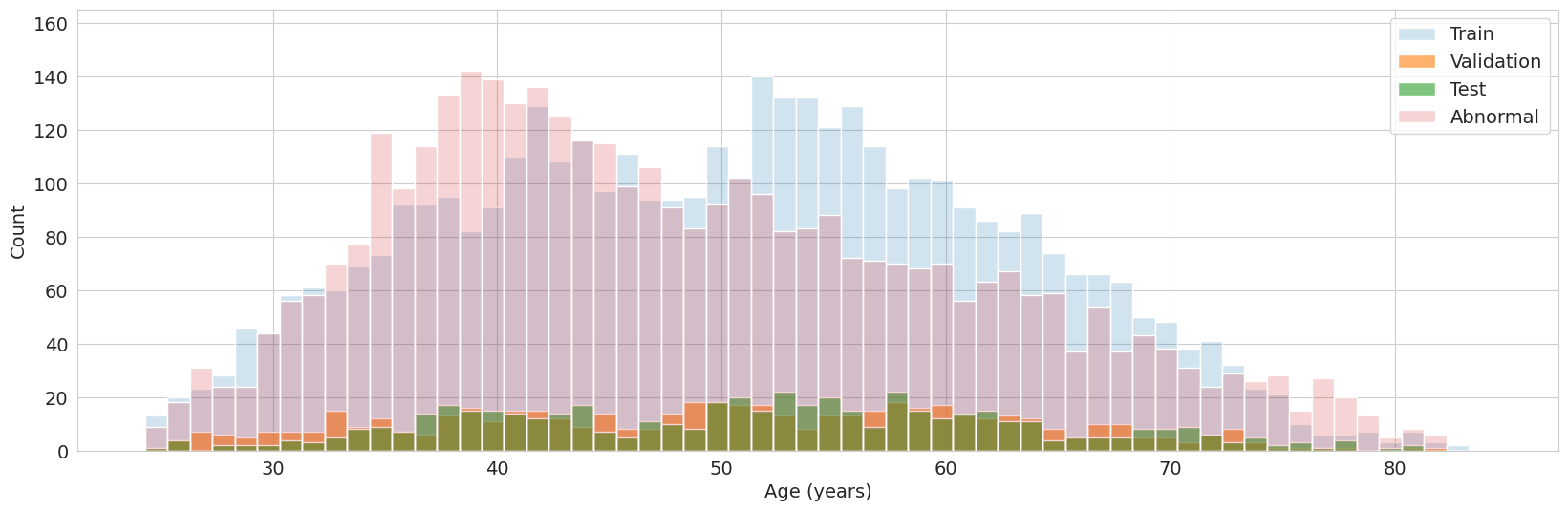
Figure 2: Distribution of male participants by age in training, validation, and test splits.
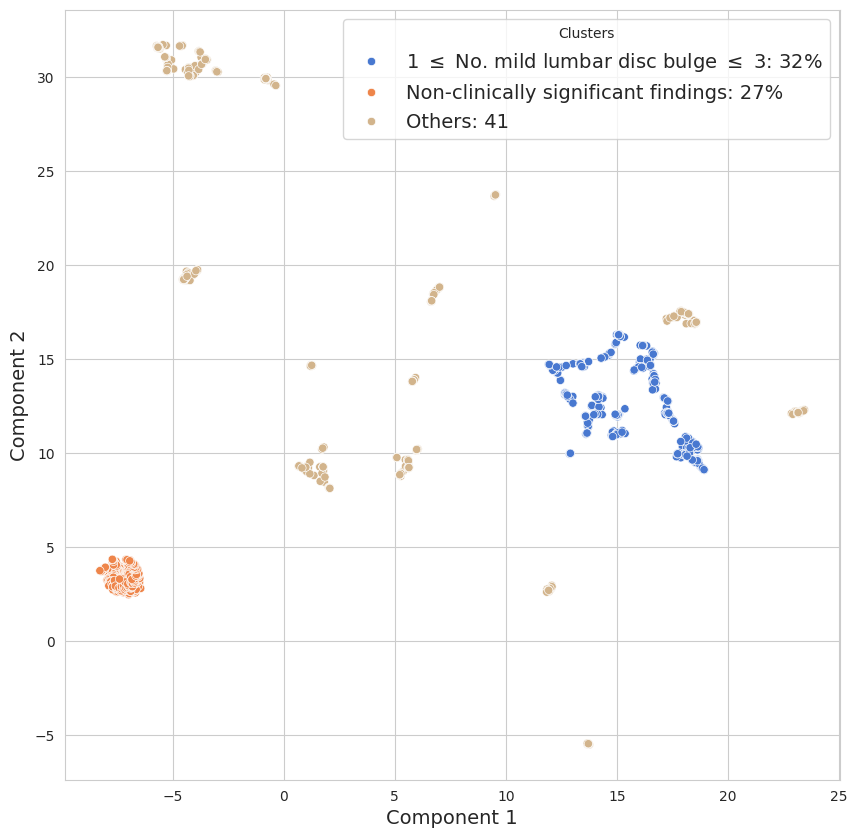
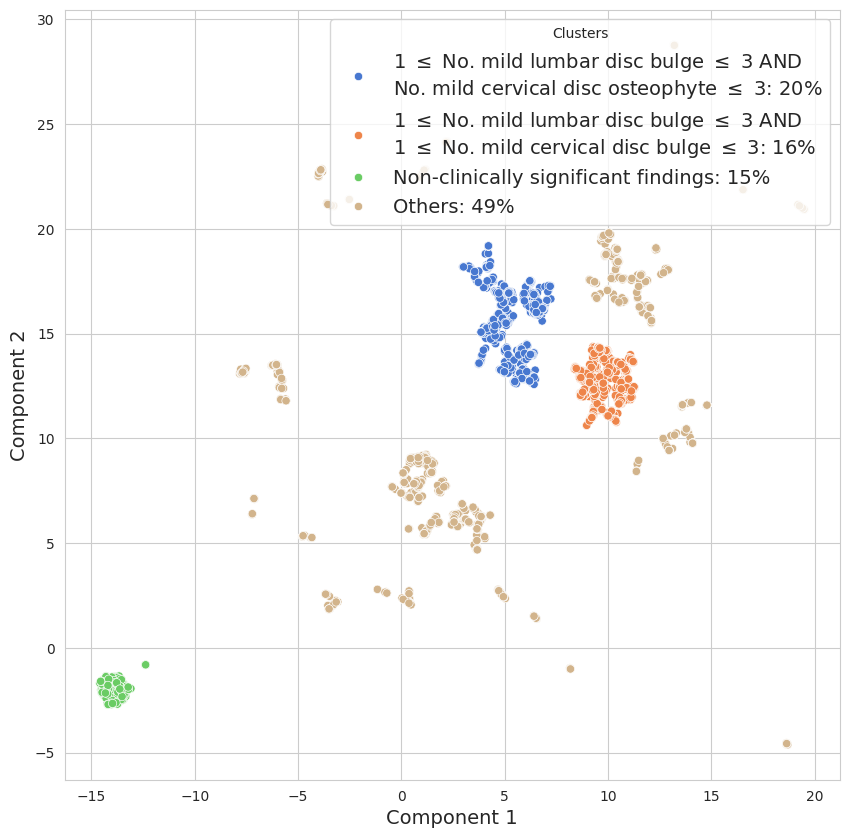
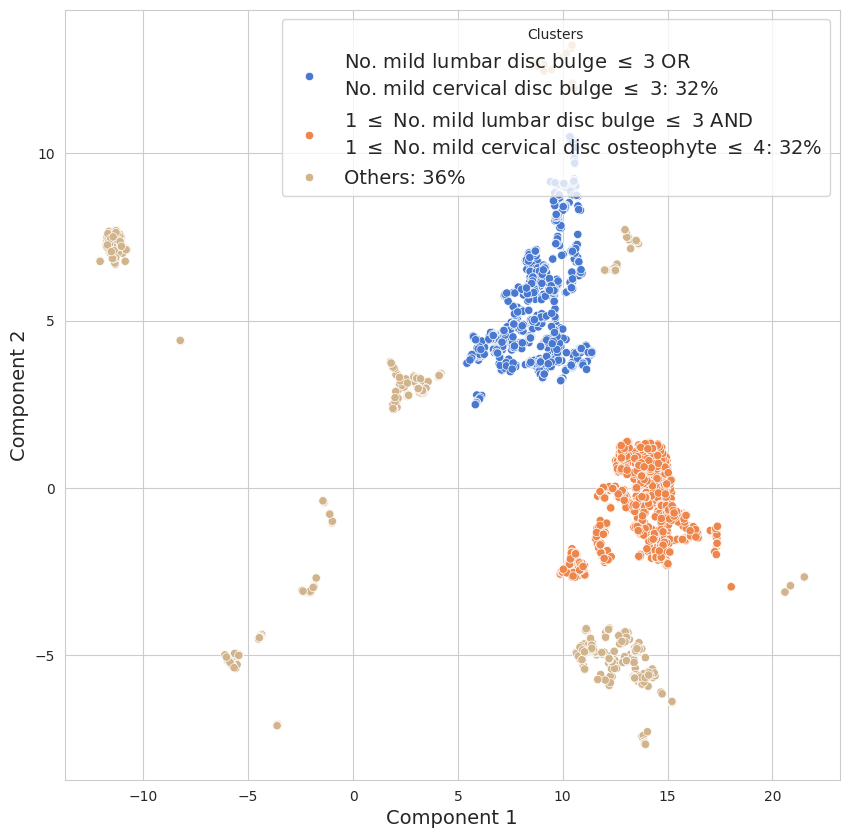
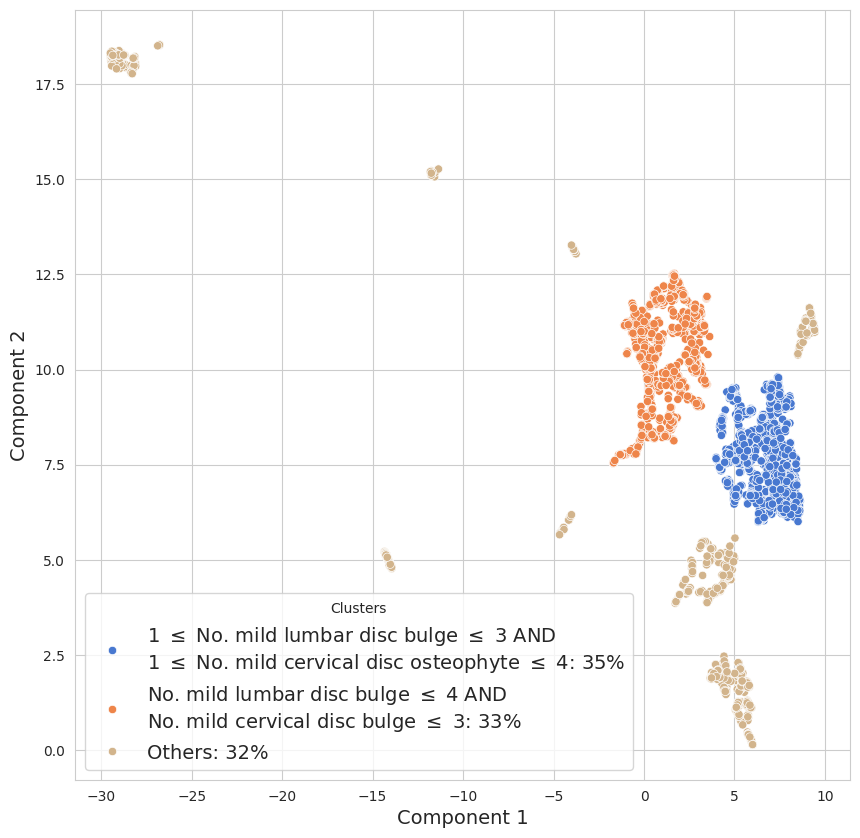
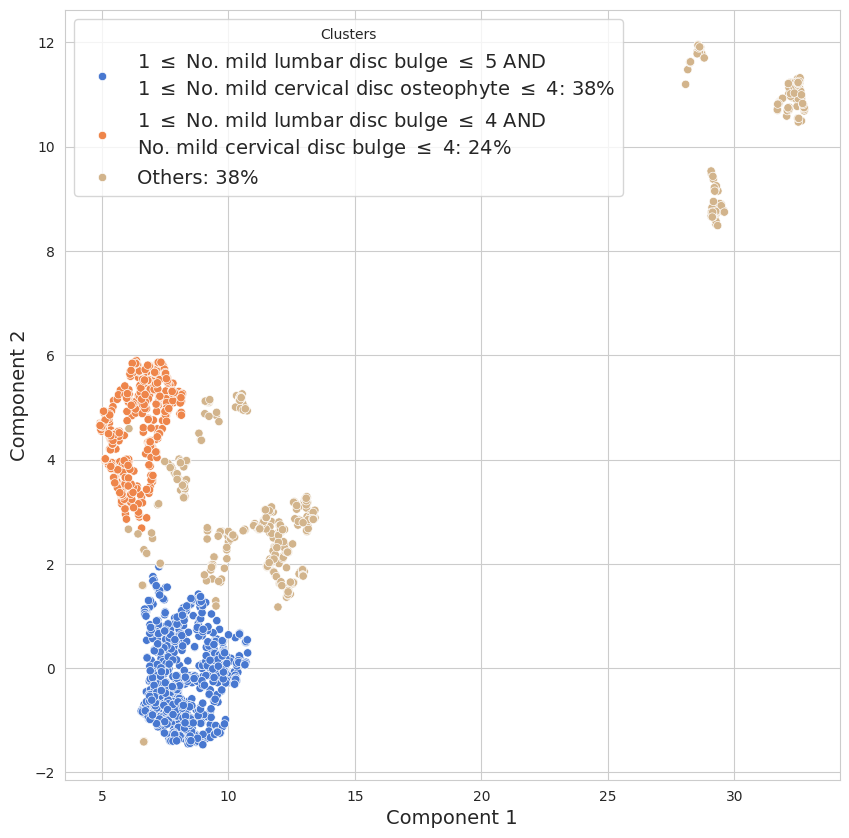
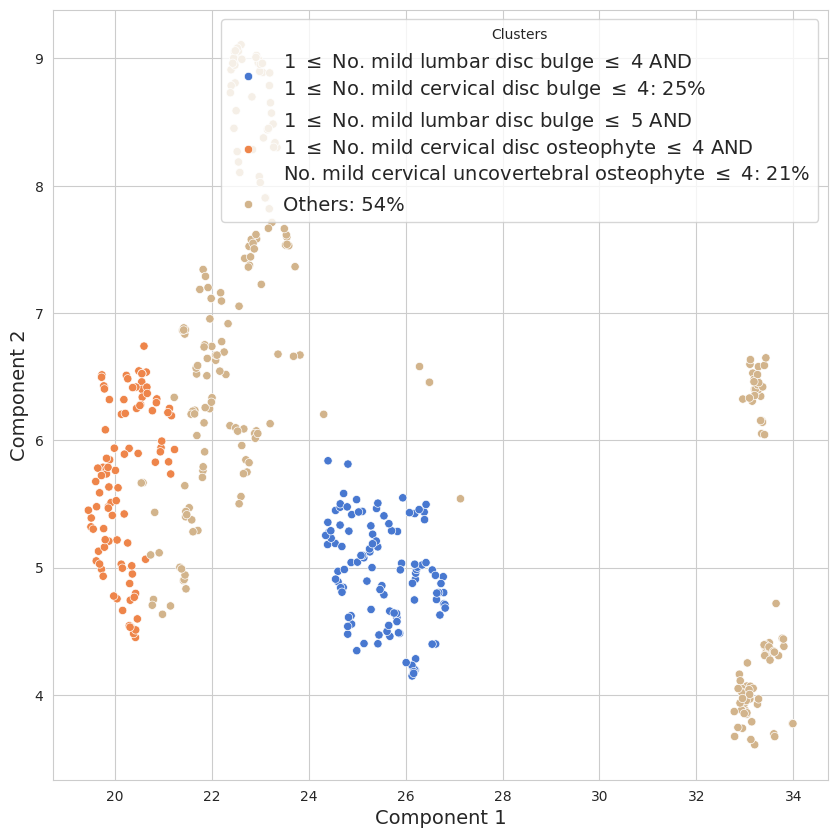
Figure 3: UMAP-based clustering of spine conditions in the 30-year-old age bracket—key to defining “normal” aging envelopes.
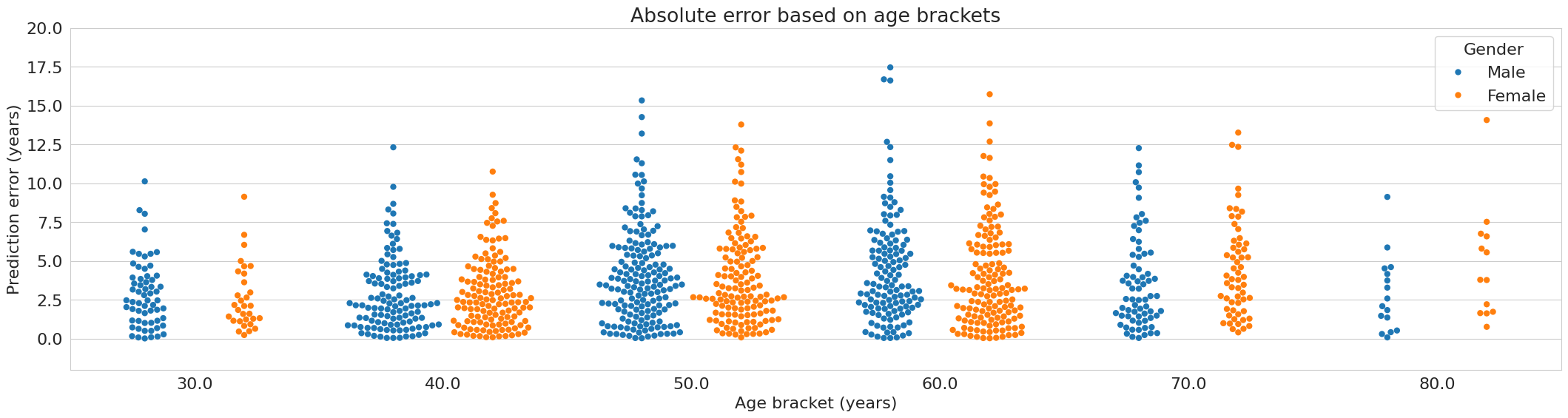
Figure 4: Absolute error of age prediction grouped by gender and chronological age bracket.
Model Interpretation
Grad-CAM visualizations demonstrate model focus on morphologic features such as disc bulges and vertebral curvatures, aligning with radiologic convention. The heatmaps highlight areas attended by the model for individual predictions, enabling both validation and nuanced error analysis. Expert review of large error cases revealed occasional segmentation failures and clinically plausible predictions for outliers.
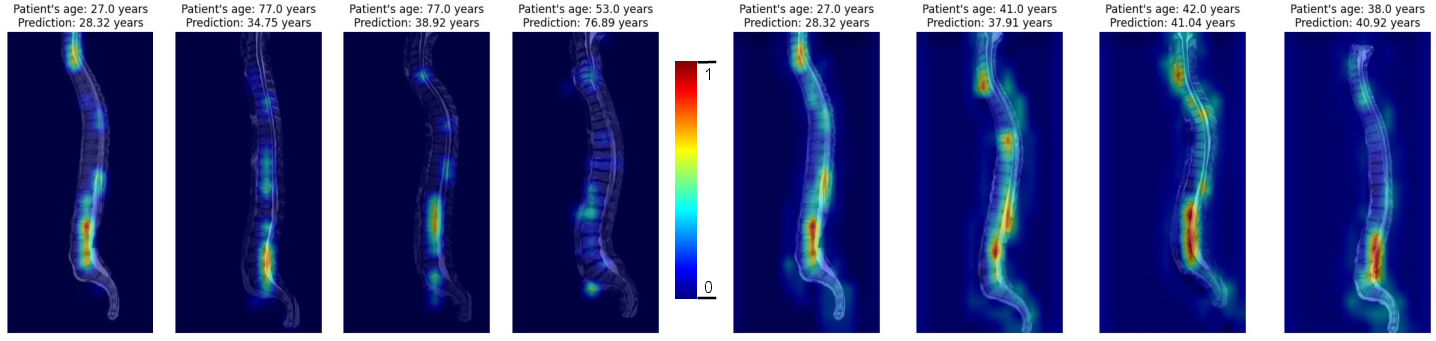
Figure 5: Grad-CAM attention maps on middle MRI slices across four subjects, revealing focus on disc bulges and vertebral features.
Spine Age Gap as a Clinically Relevant Biomarker
The “spine age gap” (SAG), defined as the difference between biological and chronological age, emerges as a biomarker tied to clinically relevant conditions. Severe disc bulges, osteophytes, fractures, and stenosis all correlated with increased SAG, with linear regression indicating that subjects with severe lumbar disc bulge had an average SAG increase of 2.96 years. Lifestyle factors such as smoking and physically demanding occupations were associated with increased SAG, whereas vigorous exercise showed a significant negative correlation.
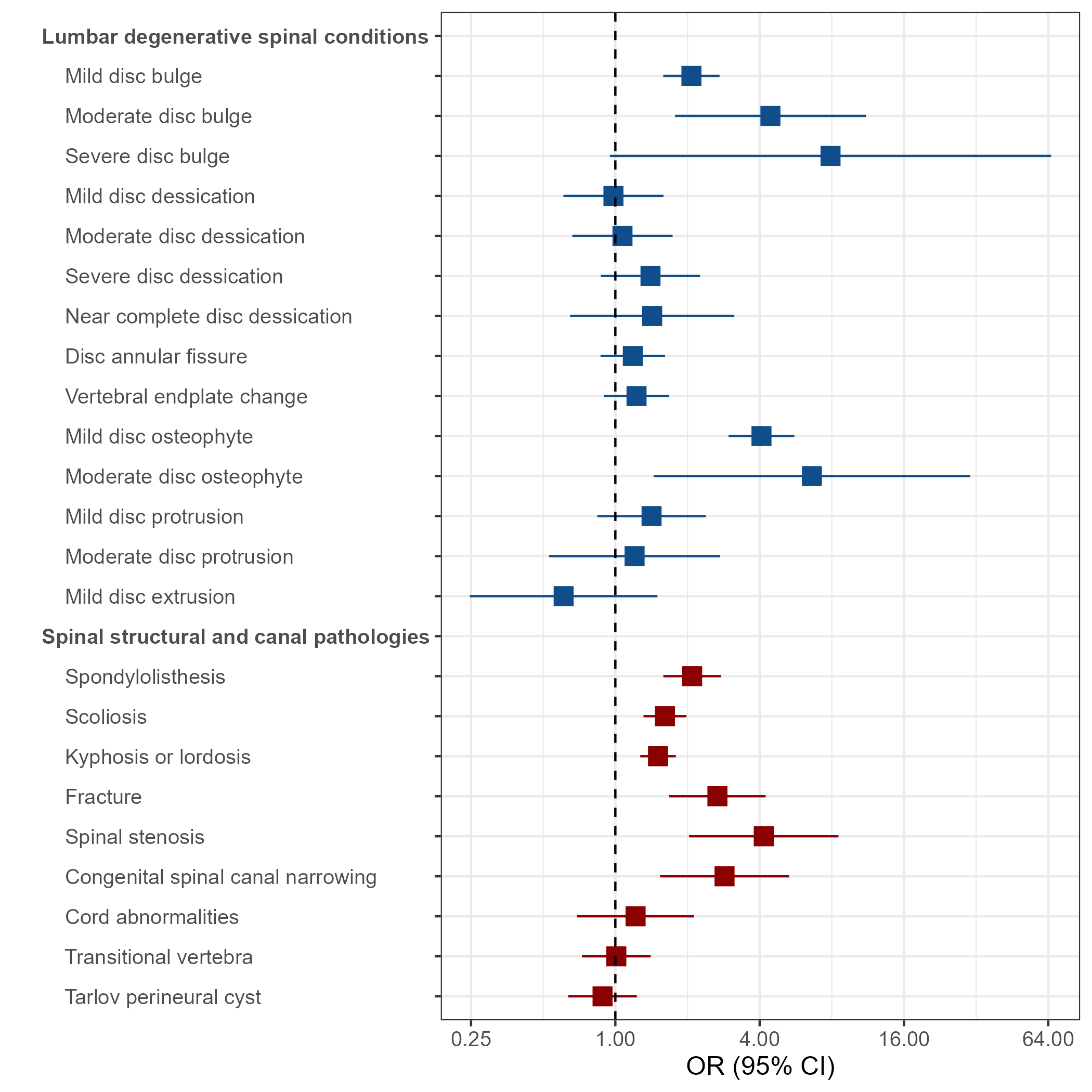
Figure 6: Odds ratios of lumbar degenerative and spinal structural conditions for subjects with large positive (>5 years) versus large negative SAG.
Furthermore, the effect of physically demanding work on SAG reverses with age, suggesting that in older individuals, continued physical activity may be indicative of better spine health.
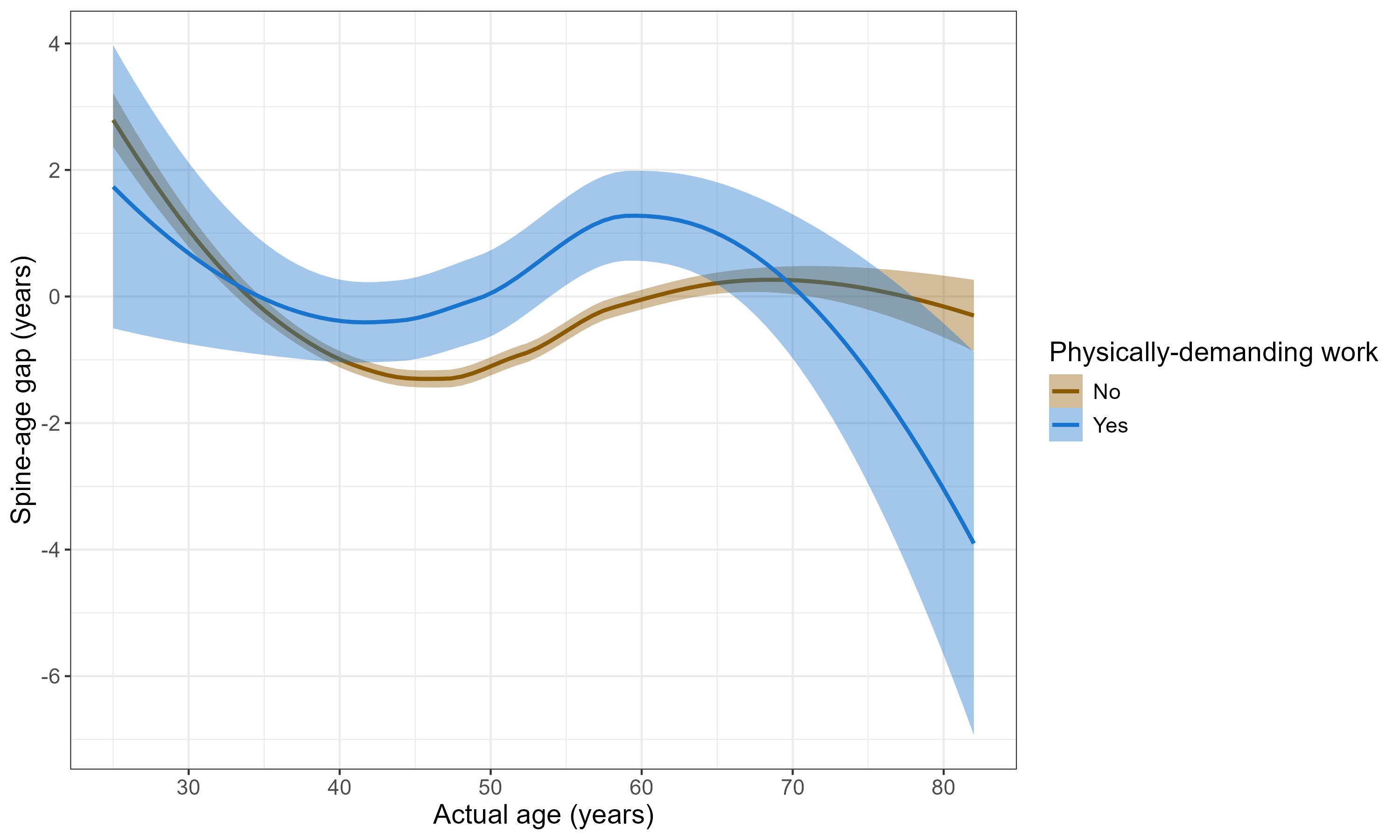
Figure 7: Mean SAG by chronological age, stratified by engagement in physically demanding work—a reversal in effect at older ages.
Implications and Future Directions
This framework, using deep learning on large-scale MRI datasets, establishes a precise, automated measure of spine aging with strong associations to both structural pathology and lifestyle metrics. The demonstrated accuracy and generalizability suggest near-immediate applicability in large-scale studies, prospective screening, and possibly in clinical decision support for spine-related disorders.
Potential future research directions include:
- Augmentation with rare/severe condition MRI datasets to enhance predictive robustness;
- Evaluation of alternative model architectures (e.g., vision transformers) for further performance scaling;
- Replacement of existing cluster-analysis normal/abnormal definitions with encoder-decoder dimensionality reduction;
- Extension to estimation of biological age in other organs (e.g., prostate, kidney, liver) for comprehensive “organ age” biomarkers.
Conclusion
This study introduces a rigorously validated DCNN framework for spine age estimation from T2-weighted MRI, leveraging advanced image segmentation and population clustering. The large-scale analysis demonstrates accurate prediction of spine age and characterizes the SAG as a biomarker intricately linked with structural degenerative conditions and lifestyle exposures. The approach represents a robust technical advance for imaging-based biological age estimation and offers a foundation for future interdisciplinary biomarker development and precision medicine in spinal health.












