- The paper presents a novel hybrid deep learning method that fuses ultrasound and lateral X-ray data to recover complete 3D vertebral shapes.
- It employs a two-stage VAE architecture with early and late fusion techniques to integrate synthetic and phantom data for detailed anatomical reconstruction.
- Quantitative results indicate significant improvements in metrics like Chamfer Distance and F1-score, demonstrating enhanced cross-domain generalizability and clinical potential.
Multi-Modal Shape Completion for Volumetric Spine Reconstruction: An Expert Review of "US-X Complete"
Introduction and Motivation
The clinical context of spinal procedures is defined by a tension between the real-time, radiation-free imaging offered by ultrasound and the incomplete anatomical depiction resulting from acoustic shadowing, especially in the vertebral body. Existing intraoperative solutions that fuse ultrasound with preoperative CT introduce registration challenges, confounded by posture-induced anatomical variance. Recent ultrasound-only deep learning reconstructions, while eliminating preoperative image dependencies, are inherently limited by the under-constrained nature of the inverse problem, particularly for the vertebral body. "US-X Complete: A Multi-Modal Approach to Anatomical 3D Shape Recovery" (2511.15600) addresses these limitations through a hybrid deep learning technique, coupling ultrasound with a single lateral X-ray for volumetric spine completion. This strategy targets intraoperative feasibility without the necessity for preoperative CT or complex intraoperative registration chains.
Data Generation and Multi-Modal Integration
The paper establishes a synthetic data generation protocol that produces paired ultrasound-like partial 3D vertebral segmentations and X-ray-consistent 2D projections. Ultrasound partial point clouds are generated through physics-aware ray casting on annotated CT meshes, closely emulating acoustic shadowing and varying probe orientations. X-ray observations are simulated by projecting vertebral segmentations onto a lateral plane and embedding these into a 3D coordinate system to ensure anatomical alignment.
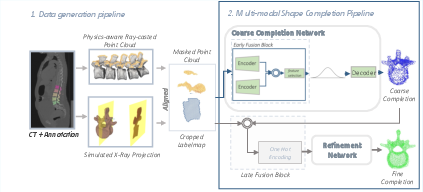
Figure 1: Schematic for paired synthetic data generation (ultrasound and X-ray), anatomical alignment in 3D space, and the two-stage network.
Key to the methodology is the construction of a unified multi-modal point cloud, embedding both modalities within a joint 3D representation space. In synthetic datasets, registration is inherent. For physical phantoms, the alignment pipeline concatenates ultrasound and X-ray data through a sequence of geometric heuristics based on bounding boxes and principal axis analysis to reflect anatomical consistency.
Deep Learning Pipeline: Coarse-to-Fine Multi-Modal Shape Completion
The shape completion architecture is structured in two stages: a coarse stage (capturing global anatomical priors) and a fine-grained refinement stage (focusing on high-frequency morphological details). Both stages are implemented as VAEs, jointly trained with KL and Chamfer Distance losses. Architectural innovations include:
- Early Fusion: Encodes modality-specific features using independent MLPs, concatenated and transformed into a shared latent space at the coarse stage.
- Late Fusion: At the refinement stage, the input concatenates the coarse prediction, ultrasound data, and X-ray data, each annotated with origin information, to enable the network to leverage the differing nature of each source.
The fusion strategy enables the model to utilize complementary information: the X-ray for global context and scaling, and the ultrasound for local, high-resolution surface details.
Experimental Setup: Phantom and Synthetic Data
Phantom validation was performed using two lumbar spine models (L1–L5), with robotic ultrasound acquisition and paired lateral X-ray/CBCT. Ultrasound segmentation relied on compounded volumes, and segmentation alignment across modalities exploited robot kinematics and mutual image registration protocols.
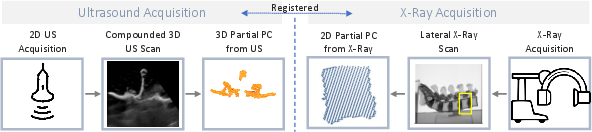
Figure 2: Experimental workflow for phantom-based validation with paired ultrasound and X-ray scans integrated into the unified 3D point cloud representation.
The phantoms employed included a 3D-printed model based on VerSe2020 annotations and a manufactured model with biomechanical tissue analogs.

Figure 3: Physical lumbar spine phantoms designed for clinical-like evaluation of the pipeline.
The learning procedure was conducted entirely on synthetic data, evaluating both on simulated datasets and real phantoms to assess both shape recovery precision and domain shift robustness.
Quantitative and Qualitative Results
The network significantly outperformed a recent ultrasound-only baseline [gafencu2024shape], achieving notable improvements in Chamfer Distance (CD), Earth Mover's Distance (EMD), and F1-score across synthetic and phantom datasets. In particular, the model delivered a mean CD reduction of 13.6 on vertebral body reconstructions in phantom data, with all statistical comparisons yielding p-values < 1e-6.
Qualitative assessment confirmed that X-ray integration greatly improved both the scale and morphology of the vertebral bodies, which are unseen in the ultrasound due to acoustic shadowing. The network demonstrated strong transfer from synthetic to real phantom data, indicating robustness to real-world imaging variability.
Bold Claims and Contrasting Results:
- Superior Cross-Domain Generalizability: The pipeline, trained exclusively on synthetic data, achieved robust inference on phantoms without further fine-tuning.
- Statistically Significant Improvements in Both Arch and Body: The approach improves arch and body completion, even when only the latter is directly informed by X-ray structure, suggesting effective multi-modal feature fusion.
Architectural Ablation and Fusion Strategies
Ablation studies isolated the roles of early and late fusion. Late fusion demonstrated the strongest enhancement in completion accuracy across metrics and anatomical regions, likely due to its capacity for modality-specific attention during point-level refinement. The combination of early and late fusion maximized completion performance, substantiating the effectiveness of hierarchical multi-modal integration.
Discussion: Implications and Future Directions
From a practical perspective, this work addresses a critical gap in intraoperative guidance: achieving volumetric, patient-specific anatomical reconstructions from real-time, low-radiation modalities without the logistics or risks of preoperative CT. This has immediate implications for workflow automation, navigation, and quantitative trajectory planning during spinal interventions.
Theoretically, the pipeline validates the utility of multi-modal feature fusion in medical image completion, where anatomical priors from one modality can anchor completion of occluded regions in another. The two-stage nature of the model supports extending this architecture to include even more contextual or semantic modalities.
Future work should address remaining challenges:
- Alignment Robustness: While the paper’s heuristic approach suffices for phantoms, clinical translation will require improved and possibly registration-free fusion to cope with anatomical variability and segmentation error.
- Whole-Spine, Context-Aware Completion: Moving from vertebra-wise operations to models that encode global spinal constraints could further enhance anatomical consistency.
Conclusion
"US-X Complete" (2511.15600) demonstrates a substantial advance in multi-modal, intraoperative anatomical shape completion. By fusing a single X-ray projection with real-time 3D ultrasound in a robust, VAE-based pipeline, it delivers significantly improved volumetric vertebrae reconstructions, overcoming fundamental limitations inherent in ultrasound-only solutions. The approach convincingly transfers from synthetic to practical settings, laying a foundation for broader adoption in intelligent, anatomy-aware surgical guidance systems. Future research should focus on further reducing dependency on precise intermodal alignment and incorporating global anatomical priors.


