VoxTell: Free-Text Promptable Universal 3D Medical Image Segmentation
Abstract: We introduce VoxTell, a vision-LLM for text-prompted volumetric medical image segmentation. It maps free-form descriptions, from single words to full clinical sentences, to 3D masks. Trained on 62K+ CT, MRI, and PET volumes spanning over 1K anatomical and pathological classes, VoxTell uses multi-stage vision-language fusion across decoder layers to align textual and visual features at multiple scales. It achieves state-of-the-art zero-shot performance across modalities on unseen datasets, excelling on familiar concepts while generalizing to related unseen classes. Extensive experiments further demonstrate strong cross-modality transfer, robustness to linguistic variations and clinical language, as well as accurate instance-specific segmentation from real-world text. Code is available at: https://www.github.com/MIC-DKFZ/VoxTell
Paper Prompts
Sign up for free to create and run prompts on this paper using GPT-5.
Top Community Prompts
Explain it Like I'm 14
Overview
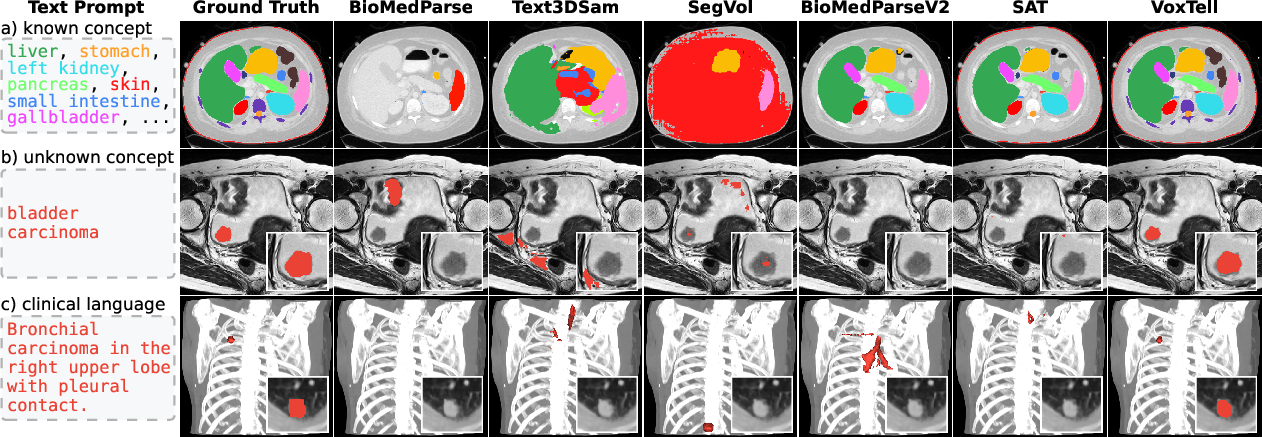
This paper introduces VoxTell, a computer program that can “read” text and use it to find and highlight specific parts inside 3D medical scans (like CT, MRI, and PET). Instead of clicking on points or drawing boxes, a doctor can type a simple word (“liver”) or a full sentence (“small tumor in the right upper lung”), and VoxTell will try to outline that area in the scan.
What were the researchers trying to find out?
The team wanted to answer a few simple questions:
- Can a single model segment many different body parts and diseases in 3D scans using only free text?
- Will it work across different scan types (CT, MRI, PET), even if it hasn’t seen that exact type in training?
- Can it understand real clinical language, including synonyms, misspellings, and detailed sentences like those in radiology reports?
- Will it still work on new or rare things it hasn’t been trained on?
How did they do it?
Think of a 3D scan like a stack of slices that make up a loaf of bread—the model needs to figure out which slices and areas belong to a body part or a lesion. VoxTell works in three main steps:
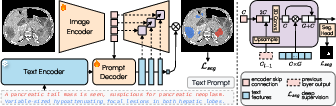
- Text understanding:
- A powerful “text encoder” reads the prompt (from one word to full sentences) and turns it into numbers the computer can use. This is like translating the doctor’s words into a guide for the model.
- Image understanding:
- A 3D “UNet-style” network looks at the scan at different zoom levels (coarse to fine), like starting with a map and then zooming in to street level. This helps the model understand both the big picture and details.
- Multi-stage vision–language fusion:
- Instead of waiting until the end, VoxTell mixes the text guidance with the image features at multiple steps while it’s decoding the image. Picture a teacher giving instructions not just at the end of an assignment, but at every step—this keeps the model aligned with the text. The paper calls this “multi-stage fusion.”
- It also uses “deep supervision,” which is like grading the homework at several checkpoints, not just at the final answer. That encourages the model to follow the prompt from early on.
Training and data:
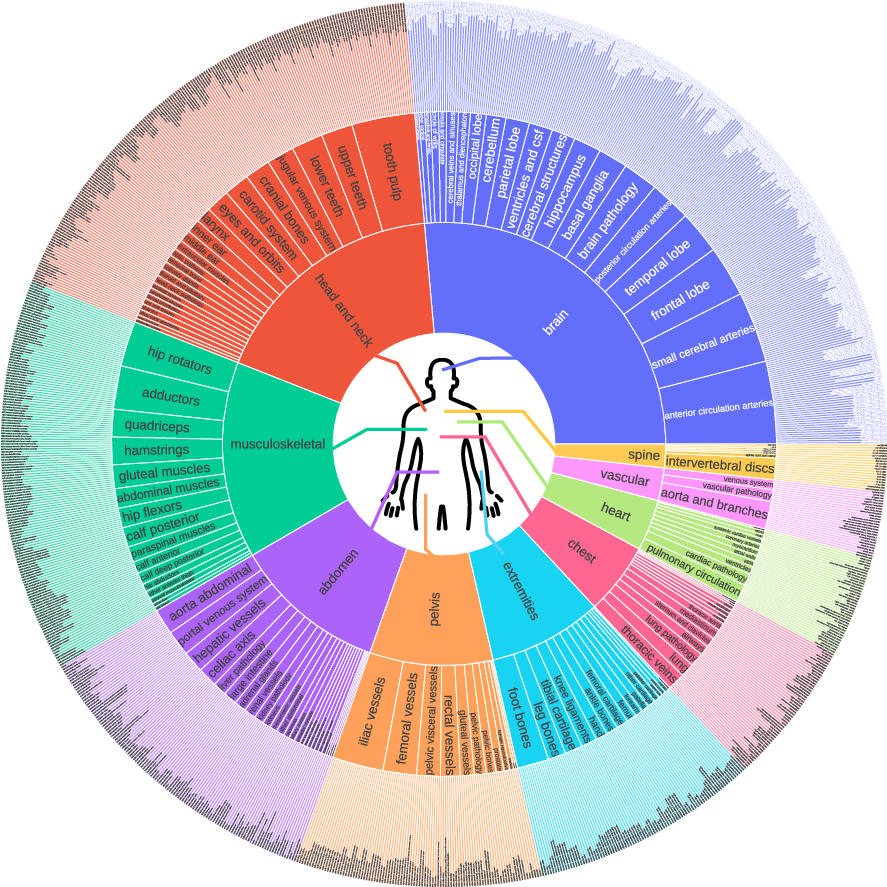
- VoxTell was trained on more than 62,000 3D scans across CT, MRI, and PET and over 1,000 different anatomical and disease concepts. The team also cleaned up and expanded the vocabulary, so it understands many ways to say the same thing (like “hepatic organ” for “liver”).
How they measured success:
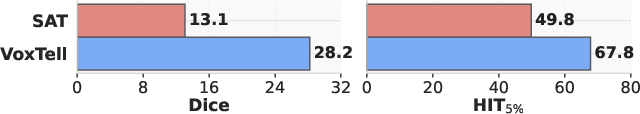
- They used the Dice score, which tells how much the model’s outline overlaps with the true outline. Imagine two shapes: if they overlap perfectly, Dice is 100; if they don’t overlap at all, it’s 0. Higher is better.
What did they find, and why is it important?
Here are the main results, explained simply:
- Strong performance across many structures and diseases:
- VoxTell beat other text-based segmentation models on many datasets in CT, MRI, and PET, including organs and lesions. It also did better than traditional tools that don’t use text.
- Works on new scan types and unseen concepts:
- Even when the model saw a familiar structure in a new modality (like MRI instead of CT), it still performed well.
- It could segment some structures and cancers it hadn’t been trained on (for example, esophageal tumors and bladder cancer), showing it can generalize.
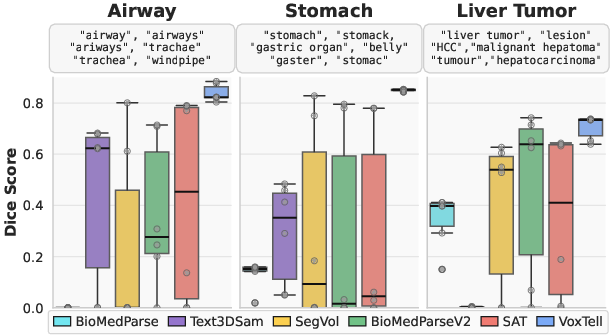
- Understands different ways of saying the same thing:
- VoxTell stayed accurate even with synonyms, rephrasings, and minor typos. That’s important because clinical language is varied and detailed.
- Handles real radiology report sentences:
- On a set of lung cancer patients, VoxTell used full sentences from reports (like “spiculated carcinoma in the left upper lobe with pleural contact”) to find the tumor, significantly outperforming other models.
- Instance-specific segmentation:
- On the ReXGroundingCT benchmark, which tests whether models can find exactly the right lesion mentioned in text, VoxTell scored higher than the previous top method. This shows it can use text to locate the specific instance described.
- Why it works:
- Ablation studies (controlled comparisons) showed that mixing text and image features at multiple stages and using deep supervision both make a big difference. Scaling up training also helps.
What does this mean going forward?
VoxTell brings us closer to “tell me what to find” medical imaging, where:
- Doctors and radiologists can use natural language to find and measure organs or lesions, saving time and reducing manual clicking.
- The model can link to existing clinical reports, helping automate parts of diagnosis and treatment planning (like outlining tumors for radiotherapy).
- It shows progress toward open-set segmentation: handling new concepts or scan types without being explicitly trained on them.
In short, VoxTell could make medical imaging more flexible and efficient by letting clinicians use plain language to guide powerful 3D segmentation, potentially improving care and speeding up workflows.
Knowledge Gaps
Knowledge gaps, limitations, and open questions
Below is a concise, actionable list of what remains missing, uncertain, or unexplored in the paper.
- Prospective clinical validation: No prospective or randomized studies demonstrate clinical utility (e.g., time savings, planning accuracy, outcome impact) compared to radiologist workflows; define task-specific acceptance thresholds and compare against expert interobserver variability.
- Safety mechanisms at inference: The model lacks calibrated uncertainty estimates, confidence maps, and explicit refusal/OOD detection for unsupported or ambiguous prompts; evaluate false-positive behavior on absent-target queries and implement safeguards.
- Robust handling of complex clinical language: Prompt robustness is shown for synonyms and minor typos, but not for negation, uncertainty (“likely,” “cannot rule out”), temporality (“interval growth”), multi-target conjunctions (“liver and spleen”), quantifiers (“three nodules”), or relational phrases (“lesion adjacent to pleura”); design tests and model components to parse these constructs.
- Multilingual and cross-institutional language generalization: Evaluation is limited to English; assess performance on other languages (e.g., German—given the institutional context), mixed-language reports, and institution-specific jargon, including domain adaptation strategies.
- Automated report-to-prompt pipeline: The method uses report-derived prompts but does not specify an automated extraction pipeline (coreference resolution, entity disambiguation, section filtering); benchmark end-to-end report parsing accuracy and its effect on segmentation.
- Instance-level segmentation beyond single findings: It is unclear how the model handles multiple instances (counting, distinct masks, selection by attributes such as size or laterality); evaluate multi-instance prompts and instance differentiation without additional fine-tuning.
- Negative and adversarial prompts: While negative prompts are sampled during training, there is no systematic evaluation of model behavior under adversarial phrasing, logically inconsistent instructions, or contradictory statements; develop adversarial and stress test suites.
- Vocabulary harmonization reliability: The LLM-based label expansion and cross-dataset harmonization may introduce semantic drift or errors; quantify mapping accuracy, human validation rates, and downstream impact of noisy or inconsistent label semantics.
- Label definition ambiguity and scope: The paper does not clarify how heterogeneous label definitions across datasets (e.g., “liver” including/excluding lesions) were resolved in training targets; assess sensitivity to differing ontologies and provide standardized definitions.
- Spatial grounding and orientation consistency: Robustness to varying DICOM orientation conventions, laterality inconsistencies, and patient positioning is not evaluated; audit orientation handling and quantify left/right errors across datasets.
- Generalization to additional modalities and acquisition types: Only CT, MRI, PET are tested; evaluate ultrasound, cone-beam CT, low-dose CT, contrast phases, and 4D/temporal sequences (e.g., dynamic PET, cardiac cine MRI).
- Performance on rare tail categories: Although gains are reported, rare or low-prevalence classes are not analyzed separately; provide per-tail metrics and sample efficiency comparisons (few-shot or zero-shot settings).
- Boundary and small-object accuracy: Metrics are limited to Dice and HIT5%; report boundary-sensitive metrics (e.g., Hausdorff distance), detection performance for small lesions, and clinical risk analysis for boundary errors.
- Calibration vs. scale: Ablation suggests benefits from multi-stage fusion and deep supervision, but disentangling architectural gains from pure scale (batch size 128 on 64 A100s) is incomplete; run controlled scaling experiments across baselines.
- Computational efficiency and deployability: Training requires substantial compute; profile inference latency, memory footprint, and explore compression/quantization/pruning for clinical deployment on commodity hardware.
- End-to-end text–vision co-training: The text encoder is frozen; investigate end-to-end co-training, domain-specific text pretraining, or multimodal alignment objectives to improve complex prompt understanding.
- Synergy with spatial prompts: The model is text-only at inference; evaluate hybrid interaction (text + points/boxes/scribbles) for more precise control, especially on challenging or ambiguous targets.
- Handling of multiple targets and compositional prompts: No experiments demonstrate segmentation for compound instructions (“segment liver and spleen, exclude lesions,” “only enhancing components”); enable set-based outputs and compositional operators (include/exclude/subset).
- Adaptation to unseen concepts: Unseen-class performance varies widely (e.g., bladder cancer 25.8 Dice); explore few-shot adaptation, concept expansion via weak supervision, and knowledge-graph/ontology guidance to improve open-set coverage.
- OOD dataset breadth and demographics: Although evaluations use unseen datasets, the breadth of institutions, scanners, patient demographics (age, sex, race), and disease prevalence is not analyzed; conduct stratified robustness and fairness audits.
- Preprocessing sensitivity: The effect of intensity normalization, resampling, cropping, and artifact handling on text-prompted performance is not reported; quantify preprocessing sensitivities and standardize pipelines.
- Error analysis and failure modes: Provide systematic analyses of common failure patterns (e.g., low contrast, motion artifacts, overlapping tissues), with targeted remediation strategies.
- Explainability and interpretability: There is no mechanism to visualize how text influences spatial predictions (e.g., cross-modal attention maps, gradient-based attribution); develop tools to make decisions auditable for clinicians.
- Clinical acceptance criteria: Define clinically meaningful thresholds per task (e.g., tumor contouring tolerances) and assess whether reported Dice values meet those criteria.
- Data and label provenance: The instance-level dataset partially derives instances via TotalSegmentator; quantify conversion errors and their effect on training and evaluation, and report provenance per class.
- Regulatory and reproducibility considerations: Clarify model weight availability, deterministic inference settings, and documentation needed for regulatory pathways; establish standard splits and benchmarks for reproducibility.
Practical Applications
Immediate Applications
Below are actionable use cases that can be deployed now, organized by sector. Each item notes key assumptions or dependencies that influence feasibility.
- Healthcare (Radiology)
- Free-text, report-driven lesion and organ segmentation within PACS and radiology viewers
- Example workflow: A radiologist types “spiculated carcinoma in the left upper lobe” or imports the report text; VoxTell returns a 3D mask aligned to the described finding (validated on ReXGroundingCT and a held-out SBRT cohort).
- Tools/products: PACS plugin; “Report-to-Mask” API; radiology viewer extension for text prompts.
- Dependencies: Integration with PACS/RIS; GPU inference; clinician-in-the-loop QA; site-specific validation; regulatory compliance for clinical use.
- Semi-automated contouring for radiation therapy planning
- Example workflow: Dosimetrist inputs “GTV of right lung primary” or “mediastinal nodes level 2R”; VoxTell produces candidate contours for review/edit.
- Tools/products: RT planning system add-on; “ContourAssist” pre-contour generator.
- Dependencies: Plan system integration (DICOM RTSTRUCT), QA policies, contouring consensus guidelines, liability/approval pathways.
- Cross-modality segmentation support (CT/MRI/PET) without retraining
- Example workflow: Segment “pancreatic tumor” in MRI even if trained predominantly on CT; useful for sites with mixed modality protocols.
- Tools/products: Modality-agnostic segmentation service “MultiMod-Seg.”
- Dependencies: Local performance audits across scanners/sequences; calibration for uncommon modalities; GPU memory for 3D volumes.
- Longitudinal lesion tracking and change detection driven by report language
- Example workflow: Extract prompts from prior and current reports (“left adrenal metastasis”); segment and compute volumetric changes.
- Tools/products: “LesionTracker” pipeline (report extraction → segmentation → volume trend).
- Dependencies: Reliable report parsing (NLP), consistent acquisition protocols, identity matching across timepoints.
- Healthcare (Operations and Quality)
- Report–image consistency checks (QA)
- Example workflow: Automatically verify that report claims (“no pleural effusion”) are consistent with segmentation outputs; flag discrepancies.
- Tools/products: “ReportQA” auditor for structured QA dashboards.
- Dependencies: Thresholding strategies; false-positive handling; governance for escalation; medico-legal considerations.
- Coding and billing assistance for imaging findings
- Example workflow: Segmentations derived from text prompts are mapped to codes (e.g., SNOMED/ICD) to support claim accuracy.
- Tools/products: “Findings-to-Code” assistant.
- Dependencies: Ontology alignment; payer policy; compliance and audit trails.
- Academia (Medical Imaging Research)
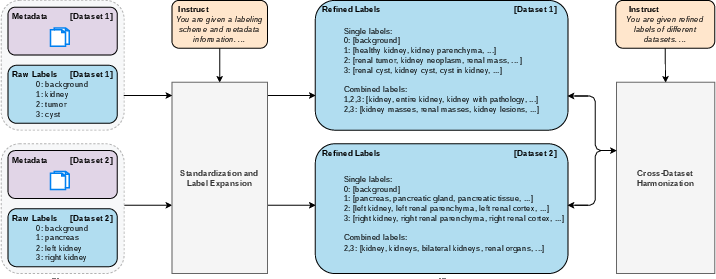
- Rapid dataset curation and label harmonization
- Example workflow: Use VoxTell’s vocabulary expansion/harmonization pipeline to unify labels across multi-dataset cohorts; accelerate annotation and re-labeling.
- Tools/products: “Vocab Harmonizer” scripts; semi-automatic relabeling toolkit.
- Dependencies: Domain expert validation of harmonized labels; provenance tracking; public dataset licenses.
- Open-vocabulary segmentation baselines for OOD evaluation
- Example workflow: Employ VoxTell as a baseline for rigorous OOD testing across new datasets/modalities; use its multi-stage fusion as a reference design.
- Tools/products: Benchmark recipes; reproducible evaluation scripts.
- Dependencies: Compute capacity; curated OOD datasets; standardized metrics.
- Education and Training
- Interactive anatomy and pathology teaching with free-text prompts
- Example workflow: Students query “hepatic vasculature,” “right rib cage,” “multiple sclerosis lesions,” and visualize 3D masks on real scans.
- Tools/products: Viewer for teaching hospitals; “AnatomyPrompt” classroom tool.
- Dependencies: De-identified teaching data; supervision by faculty; clear disclaimers.
- Policy and Governance
- Institutional AI-assisted segmentation guidelines and guardrails
- Example workflow: Draft SOPs for radiologist/dosimetrist oversight, error reporting, and validation steps when using text-prompted segmentation.
- Tools/products: Policy templates; validation protocols; bias/robustness checklists.
- Dependencies: Clinical validation data; stakeholder buy-in; alignment with regulatory guidance.
- Daily Life (Patient Engagement)
- Patient portal visualizations aligned to report text
- Example workflow: Patients view 3D masks of findings (“kidney cyst”) from their reports for education and shared decision-making.
- Tools/products: Patient-friendly viewer embedded in portals.
- Dependencies: Clinician approval workflows; health literacy considerations; secure data access; robust consent.
Long-Term Applications
The following applications require further research, scaling, or development before robust deployment.
- Healthcare (Clinical Decision Support)
- End-to-end report-grounded planning and documentation
- Vision-language pipelines that convert narrative findings into structured segmentations, measurements, and standardized reporting automatically.
- Potential products: “ReportGround” end-to-end pipeline (report extraction → segmentation → measurements → structured report).
- Dependencies: High-fidelity NLP extraction; evidence generation; regulatory approval; integration with EHR and reporting systems.
- Open-set, rare pathology segmentation across institutions and languages
- Robust performance on truly unseen categories across demographic, scanner, and language variations, including non-English clinical text.
- Dependencies: Multilingual text encoders; diverse training corpora; fairness/bias audits; domain shift calibration.
- Surgical, interventional, and intraoperative guidance
- Real-time text-driven segmentation for navigation (e.g., “arterial phase liver lesion near segment VIII”).
- Dependencies: Real-time inference; hardware (OR-grade GPUs); co-registration with tracking systems; safety certification.
- Healthcare (Population Health and Research)
- Hospital- and population-scale imaging phenotyping
- Auto-segmentation of large archives based on free-text queries (e.g., “coronary calcifications,” “vertebral fractures”) to study prevalence and outcomes.
- Dependencies: High-throughput compute; scalable storage; governance for secondary use; robust de-identification.
- Clinical trial automation
- Eligibility pre-screening using report-derived segmentations (e.g., “solitary lung nodule <3 cm”) and longitudinal response quantification.
- Dependencies: Trial-specific criteria validation; legal/ethical approvals; inter-site generalization.
- Software and Robotics
- Generalization of multi-stage vision–language fusion to non-medical 3D domains
- Voice/text-guided segmentation for industrial CT, materials inspection, geoscience volumes, and 3D mapping in robotics.
- Potential products: “OpenVLM-3D” adaptable segmentation frameworks.
- Dependencies: Domain-specific retraining; dataset availability; safety standards for industrial/robotic deployment.
- Integration with multimodal LLMs for reasoning-aware segmentation
- Systems that reason over clinical context (history, labs, protocols) to refine text prompts and segmentation targets dynamically.
- Dependencies: Reliable multimodal reasoning; hallucination safeguards; clinical governance.
- Academia (Methods and Benchmarks)
- Standardized benchmarks for report-grounded instance segmentation
- Larger, multi-institution datasets linking narrative findings to 3D masks across modalities and languages.
- Dependencies: Data-sharing consortia; annotation tooling; funding and governance frameworks.
- Methodological advances in multi-scale fusion and deep supervision
- Extending the architecture to jointly learn spatial grounding, uncertainty quantification, and causal language–vision links.
- Dependencies: New losses, calibration metrics, and interpretability methods.
- Policy and Regulation
- Regulatory pathways, reimbursement models, and liability frameworks for text-prompted segmentation
- Standards for validation, post-market surveillance, and coding (e.g., CPT) of AI-assisted segmentation outputs.
- Dependencies: Clinical trials; health technology assessment; consensus standards (DICOM extensions for prompt provenance; auditability).
- Daily Life (Consumer Health and Education)
- AR/VR tools for personalized imaging explanations
- Patients explore their own scans with narrated, text-driven overlays for pre-surgical counseling or chronic disease monitoring.
- Dependencies: Usability studies; accessibility; strong safeguards to prevent misunderstanding; clinician mediation.
Cross-cutting assumptions and dependencies
- Robustness and generalization: Performance depends on prompt quality, text encoder coverage (including multilingual/clinical jargon), and the match between local imaging protocols and training distributions.
- Safety and oversight: Clinician-in-the-loop review remains critical; deploy only with local validation, QA workflows, and clear disclaimers.
- Integration: Successful adoption requires seamless PACS/EHR/DICOM RTSTRUCT interoperability, secure data handling, and MLOps support.
- Compute and scalability: 3D inference can be GPU-intensive; throughput and latency must meet clinical workflow constraints.
- Governance: Privacy, consent, bias/fairness, and auditability need institutional policies and continuous monitoring.
Glossary
Below is an alphabetical list of advanced domain-specific terms appearing in the paper, each with a brief definition and a verbatim example from the text.
- Ablation study: A controlled experiment that systematically removes or alters components to quantify their impact on performance. "Ablation Study on TextâImage Fusion."
- Autosegmentation: Automated generation of image segmentation masks without manual delineation. "a classical autosegmentation tool without text prompts."
- Boltzmann-distributed attention: An attention mechanism whose weights follow a Boltzmann distribution to emphasize certain targets (e.g., small objects). "Boltzmann-distributed attention."
- Closed-set segmentation: Segmentation restricted to a predefined set of categories known at training time. "closed-set segmentation models"
- Cross-dataset harmonization: Standardizing and reconciling label definitions across multiple datasets. "cross-dataset harmonization"
- Cross-entropy loss: A probabilistic loss function commonly used for classification and segmentation, often combined with Dice loss. "cross-entropy losses"
- Cross-modality transfer: Generalizing a model’s knowledge across different imaging modalities (e.g., CT, MRI, PET). "strong cross-modality transfer"
- Deep supervision: Applying auxiliary losses at intermediate network layers to improve gradient flow and feature learning. "with deep supervision."
- Dice coefficient: An overlap metric for segmentation masks, measuring similarity between prediction and ground truth. "Dice coefficient."
- Encoder skip connection: A UNet link that concatenates encoder features with decoder features at corresponding scales. "encoder skip connection z_s"
- Foundation model: A large, broadly pre-trained model adaptable to many tasks via fine-tuning or prompting. "foundation models for medical images"
- Free-text prompt: An unconstrained natural-language query used to guide the model’s segmentation. "free-text prompts"
- Gross tumor volume (GTV): The visible or palpable extent of tumor used for radiotherapy planning. "gross tumor volume (GTV)"
- HIT: A hit-rate metric counting cases where predicted segmentation achieves at least 5% Dice. "HIT, the fraction of samples achieving Dice \ge 5\%."
- Instance-level annotation: Labels identifying and separating individual object occurrences of the same class. "instance-level annotations"
- Instance-specific segmentation: Segmenting a particular object instance described by text rather than a general class. "instance-specific segmentation from real-world text."
- Mask2Former: A transformer-based segmentation architecture extending MaskFormer with mask prediction across tasks. "Mask2Former-style architecture"
- MaskFormer: A transformer-based approach that uses queries to produce segmentation masks via feature fusion. "MaskFormer"
- Modality shift: A distribution change caused by switching imaging modalities or acquisition protocols. "modality shifts"
- Open-set generalization: The ability to handle classes or concepts not seen during training. "Towards open-set generalization"
- Open-vocabulary segmentation: Segmenting objects described by arbitrary text beyond a fixed label set. "open-vocabulary segmentation from free-form textual prompts"
- Out-of-distribution (OOD): Data whose distribution differs from the training set, used to test robustness. "out-of-distribution (OOD) images"
- Polynomial decay: A learning rate schedule that decreases following a polynomial function over training. "polynomial decay"
- Positron Emission Tomography (PET): A nuclear imaging modality measuring metabolic activity using radiotracers. "Positron Emission Tomography (PET)"
- Prompt decoder: A module that transforms text embeddings into guidance features for the vision decoder. "A transformer-based prompt decoder"
- Radiology report: A clinician-authored narrative describing imaging findings and interpretations. "radiology reports"
- ReXGroundingCT: A benchmark linking free-text report findings to 3D segmentations in chest CT. "ReXGroundingCT"
- Stereotactic body radiotherapy (SBRT): A precise, high-dose radiotherapy technique delivered in few fractions. "stereotactic body radiotherapy (SBRT)"
- Text-promptable segmentation: Producing segmentation masks directly conditioned on textual prompts. "text-promptable segmentation"
- TotalSegmentator: A widely used tool for comprehensive anatomical autosegmentation in CT. "TotalSegmentator"
- Transformer decoder: The component that processes queries attending to encoder memory to produce outputs. "transformer decoder"
- UNet: An encoder–decoder CNN with skip connections, widely used in medical image segmentation. "UNet-style backbone"
- Vision–language fusion: Combining textual and visual features so language can guide image understanding. "multi-stage visionâlanguage fusion"
- Zero-shot: Evaluating on tasks or classes without task-specific training examples. "Zero-shot segmentation performance"
Collections
Sign up for free to add this paper to one or more collections.