Colorectal Cancer Histopathological Grading using Multi-Scale Federated Learning
Abstract: Colorectal cancer (CRC) grading is a critical prognostic factor but remains hampered by inter-observer variability and the privacy constraints of multi-institutional data sharing. While deep learning offers a path to automation, centralized training models conflict with data governance regulations and neglect the diagnostic importance of multi-scale analysis. In this work, we propose a scalable, privacy-preserving federated learning (FL) framework for CRC histopathological grading that integrates multi-scale feature learning within a distributed training paradigm. Our approach employs a dual-stream ResNetRS50 backbone to concurrently capture fine-grained nuclear detail and broader tissue-level context. This architecture is integrated into a robust FL system stabilized using FedProx to mitigate client drift across heterogeneous data distributions from multiple hospitals. Extensive evaluation on the CRC-HGD dataset demonstrates that our framework achieves an overall accuracy of 83.5%, outperforming a comparable centralized model (81.6%). Crucially, the system excels in identifying the most aggressive Grade III tumors with a high recall of 87.5%, a key clinical priority to prevent dangerous false negatives. Performance further improves with higher magnification, reaching 88.0% accuracy at 40x. These results validate that our federated multi-scale approach not only preserves patient privacy but also enhances model performance and generalization. The proposed modular pipeline, with built-in preprocessing, checkpointing, and error handling, establishes a foundational step toward deployable, privacy-aware clinical AI for digital pathology.
Paper Prompts
Sign up for free to create and run prompts on this paper using GPT-5.
Top Community Prompts
Explain it Like I'm 14
Overview
What is this paper about?
This paper is about teaching computers to help doctors decide how serious colorectal cancer is by looking at microscope images of tissue. It focuses on doing this in a way that protects patient privacy and uses both “zoomed-in” and “zoomed-out” views of the tissue—because pathologists (doctors who study tissue) rely on both kinds of detail to make accurate decisions.
Goals and Questions
What were the researchers trying to do?
The paper set out to answer three simple questions:
- Can we train a strong cancer-grading model without moving patient data out of the hospital, so privacy is protected?
- If we let the model “look” at images at two zoom levels (fine cellular details and broader tissue patterns), does it get better at grading cancer?
- Will this approach reliably catch the most dangerous cases (Grade III), where missing them could delay urgent treatment?
How It Works
Key ideas explained in everyday language
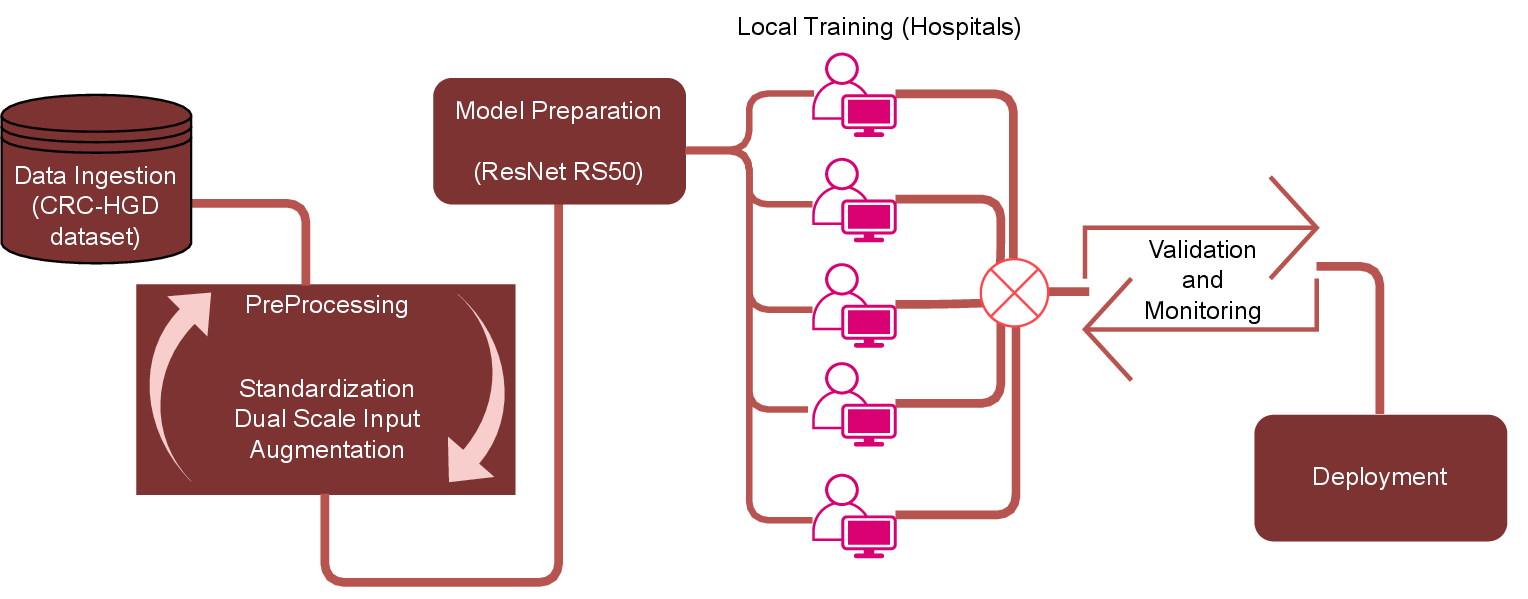
- Federated Learning (FL): Imagine a group project where each school keeps its own notes and never shares the actual notebooks. Instead, each school learns from its own notes and only sends back “lesson summaries” to build a shared, improved study guide. That’s FL: hospitals keep their patient images private, train locally, and share only model updates, not raw data.
- Multi-scale images: Pathologists switch between magnifications (like using a camera’s zoom) to see both big-picture tissue structures and tiny cell details. The computer model does the same: it “looks” at lower magnification for context and higher magnification for cell-level clues.
- Dual-stream network: Think of two cameras looking at the same scene—one wide-angle (tissue-level) and one close-up (cell-level). The model uses two image-processing “streams” (based on a popular image network called ResNetRS50) and then combines what each stream learned to make a better decision.
- FedAvg and FedProx: These are rules for how the different hospitals’ model updates are averaged and kept stable. “Client drift” (differences across hospitals due to different scanners or staining methods) can cause models to move in different directions. FedProx adds a gentle “pull” to keep everyone closer together, improving training stability.
What data did they use?
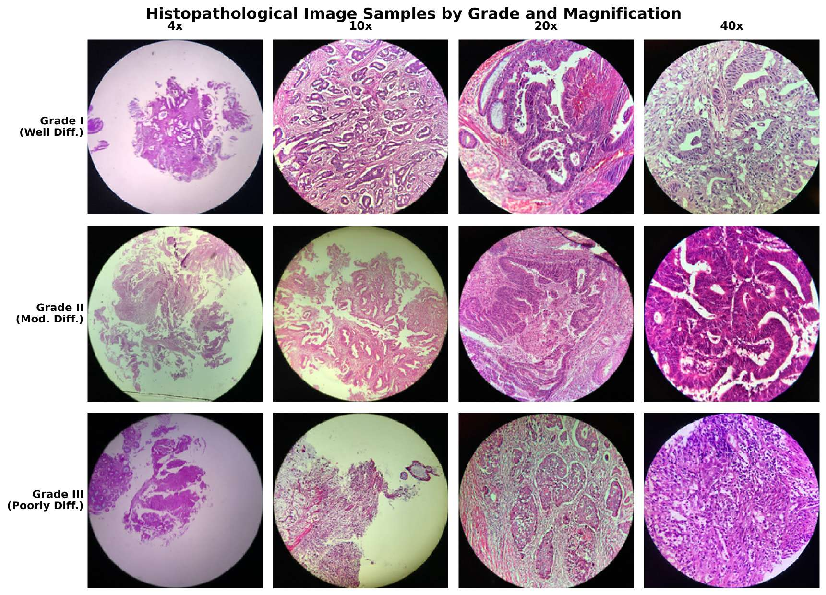
They used a dataset called CRC-HGD with 1,899 images of colorectal cancer tissue at four magnifications (4×, 10×, 20×, 40×). Each image is labeled as:
- Grade I (well differentiated): Looks most like normal tissue; least aggressive.
- Grade II (moderately differentiated): In between; most common.
- Grade III (poorly differentiated): Looks least like normal tissue; most aggressive.
What steps did the researchers take?
To make the training fair and strong across different hospitals, they carefully prepared the data and model. The main steps were:
- Standardize colors across images so differences in staining (how slides are colored) don’t confuse the model.
- Cut large slides into smaller image patches and remove background or blurry areas.
- Create two versions of each patch: one smaller (context) and one larger (fine detail).
- Balance the classes (since Grade III is rarer) so the model doesn’t ignore the most dangerous cases.
- Use data augmentation (like small color tweaks and blending images) to help the model generalize.
- Train across four simulated hospital “clients” for several rounds with local training at each hospital and global aggregation using FL.
Main Findings
What did they discover and why does it matter?
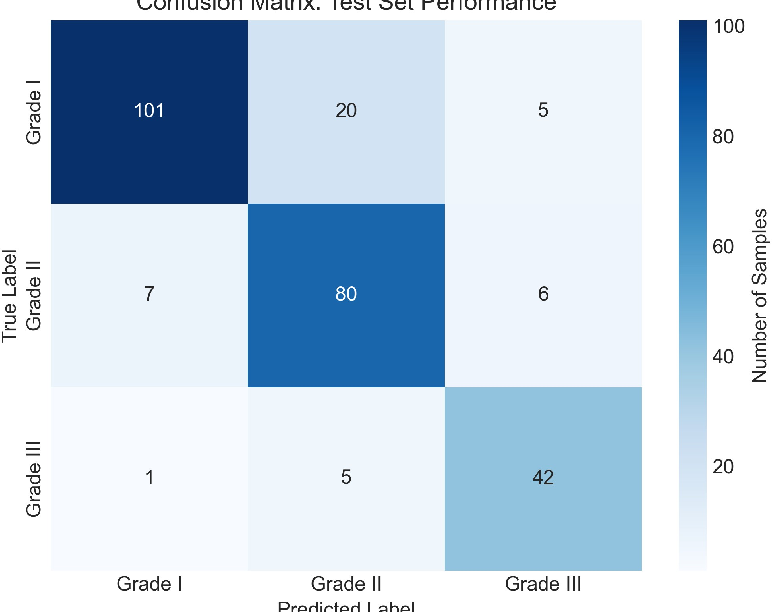
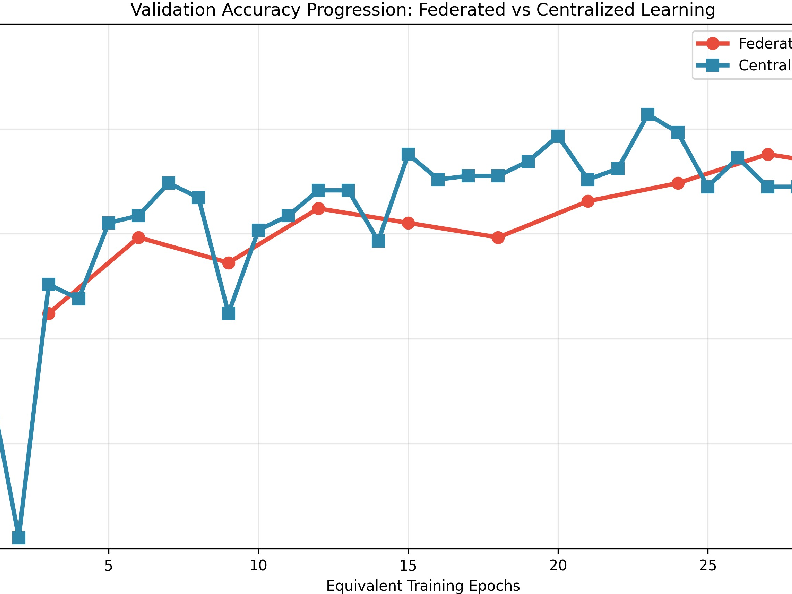
- Overall accuracy: The privacy-preserving federated approach reached 83.5% accuracy, slightly better than traditional centralized training (81.6%). This shows you don’t have to gather all data into one place to get strong performance.
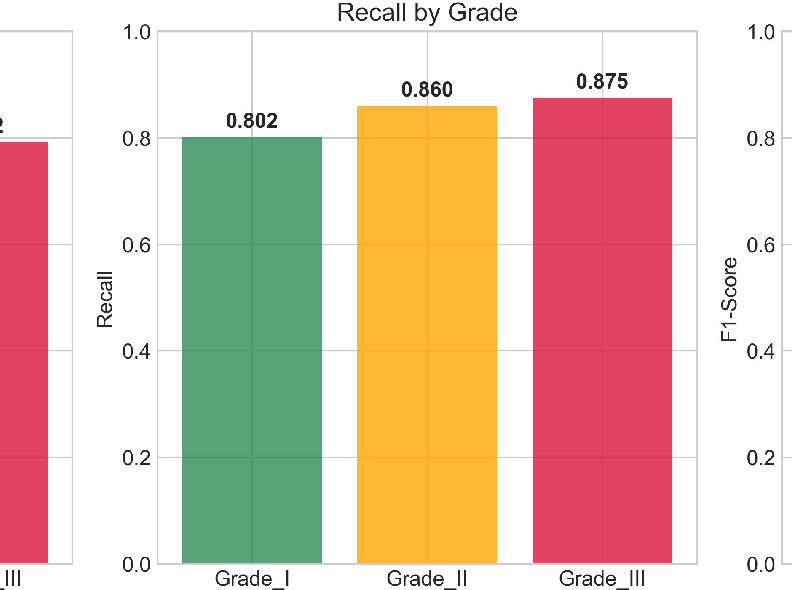
- Catching the most dangerous cases: The model had high recall (87.5%) for Grade III tumors. Recall is about finding as many true cases as possible. For aggressive cancers, high recall helps avoid missed diagnoses that could delay urgent treatment.
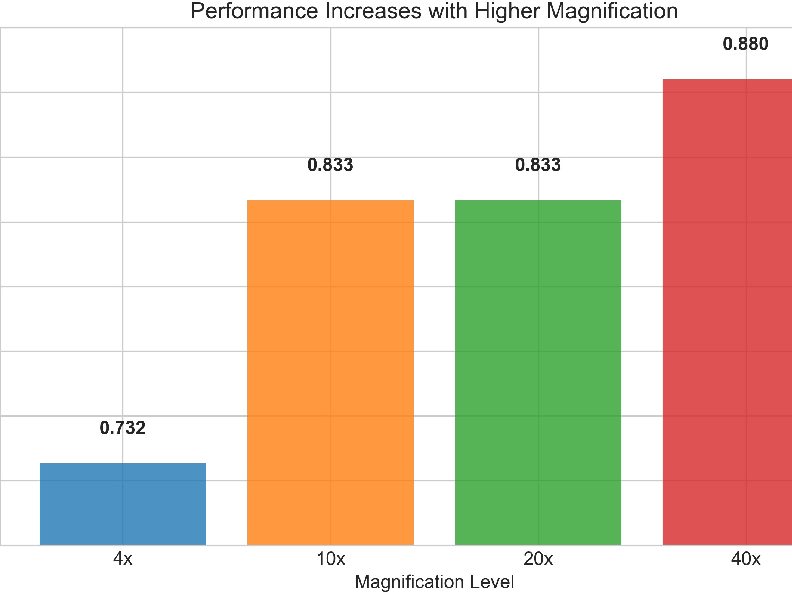
- Multi-scale helps: Accuracy improved with higher magnification (40× reached 88.0%), confirming that cell-level detail is crucial. But the model also did well at intermediate magnifications, showing that tissue-level context still adds value.
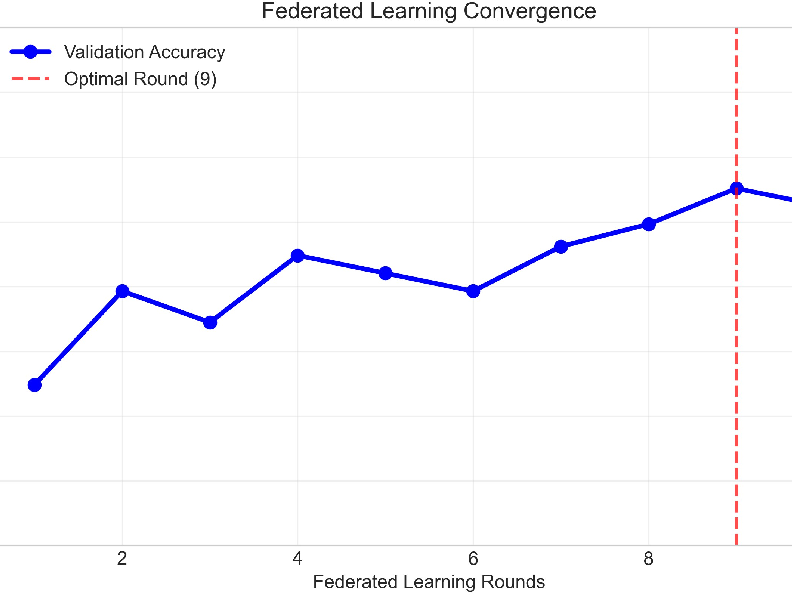
- Stable training across hospitals: Using FedProx kept different hospital models aligned, leading to smoother and more reliable training, even though hospitals’ data can differ.
Why It Matters
What is the impact of this research?
- Better patient privacy: Hospitals can collaborate to build strong models without sharing sensitive patient data, helping comply with privacy rules.
- Clinically useful focus: By prioritizing high recall for Grade III cases, the system aims to reduce dangerous false negatives—supporting safer, faster care decisions.
- Real-world readiness: The pipeline includes practical features like automatic checkpoints (save points), error handling, and flexible data input. That makes it easier to deploy in actual hospital settings.
- Path to future improvements: The approach can be expanded to more hospitals, new data types (like molecular information), and explainable tools (like heatmaps) so doctors can see why the model made a decision.
In short, this research shows a promising way to build smart, privacy-aware tools that support pathologists in grading colorectal cancer—catching the most dangerous cases more reliably while respecting patient confidentiality.
Knowledge Gaps
Below is a concise, actionable list of the paper’s unresolved knowledge gaps, limitations, and open questions that future work could address:
- Real-world cross-site validation: The federated setup was simulated with four clients; there is no evaluation with actual multi-institutional deployments (partial participation, variable connectivity, real hospital data flows, differing IT policies).
- Privacy and security guarantees: The framework lacks formal privacy protections (e.g., secure aggregation, differential privacy, homomorphic encryption) and does not present a threat model or resilience to gradient leakage, membership inference, poisoning, or backdoor attacks.
- Dataset scale and diversity: The CRC-HGD v2 dataset (1,899 images) is relatively small and may not capture institutional, demographic, and histologic diversity (e.g., rare subtypes, mucinous tumors, serrated patterns); external validation on independent, multi-center datasets is missing.
- Ground-truth reliability: The paper does not specify how grades were annotated (number of pathologists, consensus protocols, adjudication) or address inter-observer variability and label noise; uncertainty-aware labeling and probabilistic targets remain unexplored.
- Slide-level clinical inference: The pipeline focuses on patch-level classification without describing slide-level aggregation strategies (e.g., MIL, attention pooling, spatial priors) or reporting slide-level metrics aligned with clinical decision-making.
- Federated heterogeneity characterization: Non-IID data properties are unspecified (label skew, quantity skew, feature skew); there is no quantitative heterogeneity analysis or comparison of FedProx versus alternative methods (clustered FL, FedBN, personalization layers, meta-learning).
- Ablation studies on multi-scale design: There is no ablation comparing dual-stream versus single-scale baselines, alternative fusion methods (attention/gated fusion, cross-scale transformers), or contributions of each stream; inconsistency in “coarse/fine” resolution assignments needs clarification and testing.
- Magnification-specific granularity: Reported magnification effects are limited to overall accuracy; grade-wise performance per magnification, robustness to scale shifts, and the benefit of multi-scale fusion under varying zoom levels are not analyzed.
- Hyperparameter sensitivity: The choice of FedProx’s μ=0.01, number of rounds (10), local epochs (3), and optimizer settings lacks sensitivity analysis; no exploration of learning-rate schedules, early stopping, client sampling strategies, or aggregation weighting is provided.
- Communication and systems metrics: Communication cost (bytes/round), latency, model size, client memory/compute requirements, and scalability to larger client populations are not reported.
- Client-level fairness and personalization: Per-client performance, fairness across clients with different data sizes/domain characteristics, and personalization strategies (local heads, fine-tuning, proximal personalization) remain unassessed.
- Model calibration and uncertainty: Clinical deployment requires calibration metrics (ECE, Brier score), confidence intervals, and uncertainty quantification; none are reported.
- Statistical robustness: Improvements (e.g., +1.9% accuracy) lack statistical testing, multiple seeds, confidence intervals, or cross-validation; reproducibility across runs and sites is uncertain.
- Error analysis: There is no detailed analysis of failure modes (e.g., specific histologic patterns commonly misclassified, Grade II/III boundary errors), nor qualitative review with pathologists to identify systematic errors.
- Ordinal and cost-sensitive modeling: Tumor grades are ordinal, yet the model uses nominal classification; ordinal losses, margin-based approaches, or cost-sensitive training emphasizing Grade III recall were not explored.
- Explainability and clinician trust: No XAI outputs (heatmaps, attention maps) or human factors evaluation (pathologist-in-the-loop review, usability studies) are presented; integration into clinical workflows is unspecified.
- Domain generalization and adaptation: Robustness to stain, scanner, and protocol variation is only partially addressed via stain normalization and color jitter; federated domain adaptation or test-time adaptation methods are not evaluated.
- Preprocessing in FL: It is unclear whether stain normalization and artifact filtering are standardized across clients, locally tailored, or use shared references—raising questions about consistency, potential leakage, and cross-site reproducibility.
- Data partitioning details: The paper lacks transparency on client assignment (patient-level splits, magnification distribution per client), potential leakage across clients, and stratification to prevent overlap of patients/scans.
- Duplicate and artifact removal effects: Perceptual hashing and artifact filtering may remove informative patches; their impact on class distributions and performance is not quantified via controlled experiments.
- Pathology-native pretraining: ImageNet pretraining may be suboptimal for histopathology; comparisons to domain-specific pretraining (e.g., histopathology foundation models) were not conducted.
- Multi-modal extension: Although mentioned as future work, integration of molecular/genomic data and multimodal fusion architectures are not demonstrated or benchmarked.
- Regulatory and operational readiness: There is no assessment of regulatory compliance (HIPAA/GDPR in practice), audit logging, deployment latency, inference throughput, or end-to-end workflow integration in pathology labs.
- Claim of “comparable to human pathologists”: The paper asserts comparability without a human reader study or standardized benchmarking against pathologists; such a study is needed to substantiate this claim.
- Personalization to demographics and subtypes: Performance stratification across patient demographics, tumor subtypes, or clinical covariates is not reported, leaving potential bias and equity issues unexamined.
- Robustness to adversarial/poisoned data: The system’s resilience to adversarial examples or poisoned client updates is unknown; mitigation strategies (robust aggregation, anomaly detection) are not considered.
Practical Applications
Immediate Applications
The following are deployable now with existing tooling and modest integration effort, and they derive directly from the paper’s federated, multi-scale CRC grading framework and its preprocessing/engineering pipeline.
- Privacy-preserving CRC grading assistant for pathologists (Healthcare)
- Use case: Decision support to flag suspected Grade III (poorly differentiated) regions for urgent review; improves safety by prioritizing high-recall detection of aggressive tumors.
- Workflow/product: Plug-in to digital pathology viewers (e.g., Philips IntelliSite, Aperio eSlide, QuPath) that runs patch-based inference at 20×–40× magnification; outputs grade probabilities and triage alerts.
- Tools: Dual-stream ResNetRS50, stain normalization (Macenko/Reinhard), class-balanced sampling, MixUp, Softmax classifier.
- Dependencies/assumptions: On-prem GPU/CPU compute; access to WSI patches at higher magnifications; pathologist-in-the-loop validation; HIPAA/GDPR-compliant deployment; calibrated thresholds to minimize false negatives/positives.
- Multi-institution federated training pilots (Healthcare/Academia)
- Use case: Hospitals collaboratively update the model without sharing raw slides to improve generalization across staining/scanner differences.
- Workflow/product: FL orchestration with FedAvg+FedProx over 3–10 local epochs per round; periodic aggregation; local validation dashboards.
- Tools: FL frameworks (Flower, NVIDIA FLARE/Clara, FedML), Adam optimizer, FedProx, secure networking/VPN, MLflow for experiment tracking.
- Dependencies/assumptions: Data use agreements, institutional IT support, harmonized patch extraction, stain normalization protocols, non-IID handling via FedProx, audit logs.
- Magnification-aware grading guidance (Healthcare)
- Use case: Operational policy to prioritize 40× scans for grading when available, leveraging observed 88% accuracy at 40×.
- Workflow/product: SOPs for slide scanning; inference rules that adjust confidence by magnification.
- Dependencies/assumptions: Scanner capability and throughput; consistent magnification metadata; buy-in from lab operations.
- Slide triage and case prioritization (Healthcare)
- Use case: Auto-prioritize cases with high Grade III probability in the LIS queue; reduce turnaround for high-risk patients.
- Workflow/product: Integration with LIS/PACS (e.g., Epic Beaker, Cerner CoPath) to insert an “AI triage score” and routing rules.
- Dependencies/assumptions: LIS integration APIs; clinical governance approval; continuous performance monitoring; human override mechanisms.
- Histopathology preprocessing and QC toolkit (Software/Healthcare)
- Use case: Standardize slide preparation for ML across labs to reduce variability and improve downstream models.
- Workflow/product: Deploy stain normalization, Otsu tissue masking, artifact detection (Laplacian focus, pen-mark removal), perceptual hashing de-duplication.
- Tools: Open-source Python libraries (OpenSlide, scikit-image, timm), containerized microservices for preprocessing.
- Dependencies/assumptions: Access to WSI files in common formats (SVS, NDPI); local compute; pipeline observability and error handling.
- Academic benchmarking and reproducible research (Academia)
- Use case: Course labs and research projects replicating multi-scale FL on CRC-HGD; compare centralized vs. federated performance and non-IID stability.
- Workflow/product: Modular pipeline with configuration management, checkpointing, metadata snapshots; public leaderboards.
- Tools: GitHub repo, Docker/Conda environments, MLflow/W&B, CRC-HGD dataset.
- Dependencies/assumptions: Dataset licensing and IRB approvals where needed; reproducible seeds; institutional compute.
- Model monitoring and safety dashboards (Healthcare/Software)
- Use case: Track grade-wise recall/precision, drift across sites, and magnification-specific accuracy; trigger alerts if Grade III recall degrades.
- Workflow/product: MLOps dashboards for federated environments; bias/fairness reports by site and demographic.
- Tools: Evidently AI, MLflow, Grafana; custom confusion matrices and grade-level KPIs.
- Dependencies/assumptions: Data pipelines for aggregated metrics; agreements on privacy-preserving telemetry; well-defined KPIs.
- Privacy-compliant collaboration policies (Policy/Healthcare)
- Use case: Immediate policy templates enabling FL collaborations without raw data sharing (DPA, DUAs, SOPs for aggregation logs).
- Workflow/product: Governance playbook for FL; audit trails and incident response plans.
- Dependencies/assumptions: Legal counsel; institutional privacy officers; secure aggregation infrastructure.
- Training augmentation modules for pathology ML teams (Software/Academia)
- Use case: Drop-in augmentation components (MixUp, class-balanced sampling, color jitter) to improve generalization and minority-class performance.
- Workflow/product: Python package or notebooks with configurable augmentations tied to pathology-specific constraints.
- Dependencies/assumptions: Balanced sampling across grades; validation to avoid label noise; integration with existing pipelines.
- Vendor prototyping of privacy-aware pathology solutions (Industry/Software)
- Use case: Rapid prototyping of federated digital pathology products using the paper’s modular pipeline and FedProx-stabilized training.
- Workflow/product: MVPs for grading assistants; cross-site model updates; pilot deployments.
- Dependencies/assumptions: ISO 13485/IEC 62304 processes; QA, cybersecurity; clinical validation plans.
Long-Term Applications
These require further clinical validation, scaling, regulatory approval, or significant development to become dependable at scale.
- Regulatory-cleared AI for CRC grading with federated updates (Healthcare/Industry)
- Use case: FDA/CE-marked decision support that continuously improves via federated learning while maintaining safety controls.
- Workflow/product: Clinical-grade software with post-market surveillance, human-in-the-loop verification, locked update cycles and explainability features.
- Dependencies/assumptions: Multicenter prospective trials; robustness across scanners/stains/populations; risk management and traceability; regulatory submissions.
- National/international federated pathology consortia (Healthcare/Policy)
- Use case: Cross-silo FL networks among hospitals to boost performance without cross-border data transfer.
- Workflow/product: Secure aggregation, differential privacy, interoperability standards (DICOM-WSI, HL7 FHIR), governance councils.
- Dependencies/assumptions: Funding, legal frameworks for cross-jurisdiction collaboration, standardized metadata, secure networking.
- Cross-cancer generalization (Healthcare/Academia)
- Use case: Extend multi-scale FL to breast, prostate, lung, and other cancers; unify grading pipelines across tumor types.
- Workflow/product: Transfer learning with multi-stream encoders adapted to tumor-specific morphology.
- Dependencies/assumptions: High-quality labeled datasets; domain adaptation for stain/scanner heterogeneity; clinical validation.
- Federated personalization and clustered FL (Software/Healthcare)
- Use case: Site-specific personalization (e.g., clustered FL, fine-tuning heads) to address non-IID data and local staining practices.
- Workflow/product: Adaptive aggregation strategies; on-device personalization modules; fairness-aware optimization.
- Dependencies/assumptions: Research maturation, evaluation metrics for personalization vs. generalization, client compute budgets.
- Multi-modal federated prognostics (Healthcare/Academia)
- Use case: Combine histopathology with genomics/clinical data in federated settings to predict outcomes and therapy responses.
- Workflow/product: Secure multiparty computation, federated feature fusion; survival prediction tools.
- Dependencies/assumptions: Reliable data linkage; privacy-preserving cryptographic protocols; IRB/ethics approvals.
- Real-time edge inference on scanners (Healthcare/Industry)
- Use case: On-scanner or edge appliances performing patch-based grading during acquisition to accelerate workflow.
- Workflow/product: Hardware-accelerated inference (GPU/TPU), vendor SDK integration, streaming microservices.
- Dependencies/assumptions: Hardware support from scanner vendors; rigorous latency and reliability testing; hospital IT integration.
- XAI-integrated clinical decision support (Healthcare/Policy)
- Use case: Deploy saliency/attention maps and case-level explanations to build trust and support audits.
- Workflow/product: Explainability overlays in WSI viewers; standardized explanation reports for QA.
- Dependencies/assumptions: Clinically validated explanation methods; human factors studies; regulatory expectations for transparency.
- Trial stratification and precision oncology (Healthcare/Pharma)
- Use case: Use high-recall Grade III identification to accelerate patient recruitment and stratification in trials targeting aggressive CRC.
- Workflow/product: Screening pipelines linked to trial management systems; federated performance monitoring across sites.
- Dependencies/assumptions: Trial protocol alignment; patient consent; interoperability with clinical systems.
- Public health surveillance without data sharing (Policy/Healthcare)
- Use case: Aggregate privacy-preserving model outputs to monitor regional grade distributions and detect shifts in aggressiveness patterns.
- Workflow/product: Federated analytics dashboards; differential privacy for reporting.
- Dependencies/assumptions: Ethical frameworks; bias monitoring; standardized reporting pipelines.
- LIS/PACS/EMR integration at scale (Healthcare/Software)
- Use case: Seamless data flows for automated grading, triage, and documentation within EMR/LIS ecosystems.
- Workflow/product: APIs, message buses (FHIR), end-to-end MLOps for federated models.
- Dependencies/assumptions: Vendor cooperation; interoperability standards; sustained IT support.
- Safety, fairness, and liability frameworks for federated pathology AI (Policy/Industry)
- Use case: Define standards for performance auditing, site-level bias detection, and responsibility-sharing in federated deployments.
- Workflow/product: Certification schemes, audit tools, legal templates for AI-assisted diagnoses.
- Dependencies/assumptions: Cross-stakeholder consensus; insurer/regulator engagement; standardized metrics.
- Operational and reimbursement automation (Healthcare/Finance)
- Use case: Auto-populate grade fields and documentation to streamline billing and quality reporting.
- Workflow/product: Coding assistants integrated with EMR; audit trails to support reimbursement.
- Dependencies/assumptions: Payer acceptance; compliance with clinical documentation rules; robust error handling.
- Ecosystem of standardized FL pathology tooling (Software/Academia/Industry)
- Use case: Mature libraries for multi-scale FL in pathology, model registries, deployment blueprints, and test suites.
- Workflow/product: Open-source packages, reference pipelines, best-practice guides for stain normalization and artifact handling.
- Dependencies/assumptions: Community adoption; funding for maintenance; alignment with standards bodies.
Notes on cross-cutting assumptions and dependencies:
- Data quality and representativeness (grade distribution, magnification availability, staining variability).
- Secure infrastructure (encryption, access control, auditability) compliant with HIPAA/GDPR.
- Continuous monitoring for drift, bias, and safety; human oversight and clear escalation paths.
- Legal agreements and governance for federated collaborations.
- Sufficient compute resources at participating sites; robust networking for aggregation rounds.
- Thorough clinical validation and stakeholder training to ensure trust and effective use.
Glossary
- Adam optimizer: An adaptive gradient-based optimization algorithm commonly used to train deep neural networks. "Adam optimizer (lr , weight decay )"
- Attention maps: Visual explanations that highlight image regions most influential to a model’s prediction. "heatmaps and attention maps"
- Beta distribution: A continuous probability distribution on [0,1] used here to sample MixUp interpolation coefficients. "where with ."
- Class-balanced sampling: A sampling strategy that ensures minority classes are sufficiently represented during training. "class-balanced sampling ensured adequate inclusion of minority classes during training, preventing model bias."
- Client drift: Divergence of local model updates across clients due to heterogeneous data, harming global convergence. "stabilized using FedProx to mitigate client drift across heterogeneous data distributions from multiple hospitals."
- Clustered FL: A federated learning approach that groups clients by model similarity to improve personalization and stability. "introduced a clustered FL framework that groups clients by model similarity"
- Computational pathology: The application of computational methods and machine learning to analyze pathology data. "helped catalyze computational pathology."
- Convolutional neural networks (CNNs): Deep learning architectures designed for image analysis using convolutional layers. "Deep learning, particularly convolutional neural networks (CNNs), has transformed digital pathology"
- CRC-HGD dataset: A curated dataset of colorectal cancer histopathology images annotated by grade and magnification. "Extensive evaluation on the CRC-HGD dataset demonstrates that our framework achieves an overall accuracy of 83.5%"
- Cross-silo federated neural network: Federated learning across institutional silos, enabling collaboration without sharing raw data. "implemented a cross-silo federated neural network for dropout prediction across educational institutions"
- Digital pathology: The digitization and computational analysis of pathology slides. "With the rapid growth of digital pathology, machine learning has emerged as a powerful tool"
- Domain adaptation: Techniques to adapt models to distribution shifts across domains or institutions. "developing federated domain adaptation techniques"
- Dropout regularization: A neural network technique that randomly drops units during training to reduce overfitting. "dropout regularization"
- Explainable AI (XAI): Methods that make model predictions interpretable and transparent to users. "incorporating explainable AI (XAI) features"
- FedAvg: The standard federated averaging algorithm that aggregates client model weights. "FedAvg algorithm were used for global aggregation."
- Federated learning (FL): Collaborative model training across multiple clients without sharing raw data. "Federated learning (FL) offers a paradigm shift, allowing multiple institutions to collaboratively train AI models without exchanging raw data"
- FedProx: A federated optimization method that adds a proximal term to local objectives to improve stability under heterogeneity. "stabilized using FedProx to mitigate client drift"
- Heatmaps: Visual overlays indicating regions of high model activation or importance. "heatmaps and attention maps"
- Hematoxylin and eosin (H&E) staining: A standard staining technique in histology for visualizing tissue structure. "hematoxylin and eosin (H{paper_content}E) staining"
- Hierarchical context: Multi-level contextual information (e.g., patch-to-slide) used to improve classification. "leveraged hierarchical context with patch-based models for whole-slide classification."
- ImageNet: A large-scale image dataset used to pre-train vision models for transfer learning. "pre-trained on ImageNet (via the timm library)."
- Laplacian-based focus metrics: Measures derived from the Laplacian operator to detect blur and assess image sharpness. "Laplacian-based focus metrics and morphological heuristics"
- LoRA-based model adaptation: A parameter-efficient fine-tuning technique using low-rank updates. "using LoRA-based model adaptation and adaptive aggregation strategies"
- Macenko (stain normalization): A method for standardizing H&E staining across slides. "normalized using stain-standardization methods such as Macenko or Reinhard."
- Magnifications: Optical zoom levels (e.g., 4×, 10×, 20×, 40×) used to view tissue at different scales. "provided at four magnifications (4, 10, 20, 40)"
- MixUp: A data augmentation technique that blends pairs of images and labels to improve generalization. "MixUp~\cite{Zhang2017} was applied to improve generalization."
- Mitotic figures: Microscopic features representing cells undergoing division, indicative of tumor aggressiveness. "high magnifications (40) capture nuclear atypia and mitotic figures"
- Morphological heuristics: Rule-based image-processing checks leveraging tissue morphology to filter artifacts. "morphological heuristics"
- Multi-scale imaging: Analysis that integrates information from multiple resolutions or magnifications. "multi-scale imaging is central to pathology"
- Non-IID distributions: Client data distributions that are not independent and identically distributed, causing heterogeneity. "To address client drift from non-IID distributions, FedProx~\cite{Macenko2009} was integrated"
- Nuclear atypia: Abnormal nuclear appearance associated with malignancy. "high magnifications (40) capture nuclear atypia and mitotic figures"
- Otsu’s thresholding: An automatic method to choose a threshold that separates foreground and background. "Otsu’s thresholding on tissue masks"
- Pathologist-in-the-loop validation: Workflow where expert pathologists review and validate AI outputs. "pathologist-in-the-loop validation."
- Patch-based models: Models that operate on small image tiles extracted from larger slides. "patch-based models"
- Patch extraction: The process of tiling whole-slide images into fixed-size patches for analysis. "Patch Extraction:"
- Perceptual hashing: Hashing that captures perceptual similarity to identify duplicate or near-duplicate images. "perceptual hashing"
- Proximal term: A penalty term that constrains local updates toward the global model to reduce drift. "by adding a proximal term to each local objective"
- ResNetRS50: A ResNet variant optimized for performance and scale used as the backbone encoder. "dual-stream ResNetRS50 backbone"
- ReLU activation: A nonlinear activation function defined as max(0, x) used in deep networks. "ReLU activation"
- Softmax: A function that converts logits into class probability distributions. "employs Softmax to predict class probabilities"
- Stain-standardization: Normalization of staining variations to reduce inter-lab variability. "stain-standardization methods"
- timm library: A PyTorch library providing pretrained image models and utilities. "timm library"
- Whole-slide classification: Classification of entire digitized pathology slides rather than patches. "whole-slide classification."
- Whole-slide images (WSIs): High-resolution digital scans of entire pathology slides. "Whole-slide images (WSIs) were tiled into fixed-size patches."
Collections
Sign up for free to add this paper to one or more collections.