- The paper presents MEIcoder, which decodes visual stimuli from primary visual cortex (V1) recordings by integrating neuron-specific most exciting inputs (MEIs).
- The methodology combines a six-layer CNN core with subject-specific readin modules and employs SSIM-based and adversarial losses for high-fidelity image reconstruction.
- The results show superior performance in low-data and low-neuron regimes, underscoring potential applications in neuroprosthetics and brain-machine interfaces.
Introduction and Motivation
The paper introduces MEIcoder, a biologically informed neural decoding framework designed to reconstruct visual stimuli from neural population activity, specifically targeting the primary visual cortex (V1). The motivation stems from the challenge of decoding high-fidelity images from limited and noisy neural data, a scenario common in primate and human studies where large-scale, high-throughput recordings are infeasible. Existing approaches either overfit in low-data regimes or rely on generative models that hallucinate semantically plausible but spatially inaccurate reconstructions. MEIcoder addresses these limitations by integrating neuron-specific most exciting inputs (MEIs), a perceptually motivated SSIM loss, adversarial training, and a parameter-efficient architecture.
MEIcoder Architecture
MEIcoder is composed of two main components: a shared core module and subject-specific readin modules. The readin module embeds neural responses into a latent space using MEIs and learnable neuron embeddings, while the core module—a six-layer CNN—maps these embeddings to reconstructed images. The architecture is designed to facilitate transfer learning across subjects with heterogeneous neural populations.
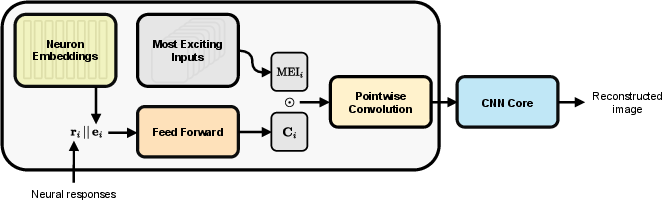
Figure 1: Architecture of MEIcoder, illustrating the flow from neural responses and neuron embeddings through context representations, MEI modulation, pointwise convolution, and the CNN core to image reconstruction.
The readin module leverages MEIs, which are images that maximally excite individual neurons, providing a strong computational prior aligned with biological coding properties. Context representations are generated by projecting neural responses and neuron embeddings, which are then pointwise-multiplied with MEIs to form neural maps. These maps are compressed via pointwise convolution to a fixed channel dimension, enabling compatibility with the core module regardless of the number of recorded neurons.
Training Objectives
MEIcoder is trained end-to-end using a composite loss function:
- SSIM-based reconstruction loss: Negative log-SSIM is used to prioritize perceptually relevant features over pixel-wise accuracy, outperforming MSE and perceptual losses based on pre-trained embeddings in terms of stability and artifact suppression.
- Adversarial loss: A discriminator CNN is trained to distinguish reconstructed images from ground-truth images, regularizing the decoder to produce naturalistic outputs and mitigating overfitting in low-data regimes.
Hyperparameter optimization and early stopping are performed using validation set performance on AlexNet-based identification metrics.
Experimental Evaluation
MEIcoder is evaluated on three datasets:
- Brainreader: Mouse V1, 8,587 neurons, 5,100 samples.
- SENSORIUM 2022: Mouse V1, 8,372 neurons, 5,084 samples.
- Synthetic Cat V1: Biologically realistic spiking model, 46,875 neurons, 50,250 samples.
MEIcoder is compared against five baselines: InvEnc, EGG, MonkeySee, CAE, and MindEye2. Evaluation metrics include SSIM, pixel correlation, and two-way identification using AlexNet features.

Figure 2: Reconstructions on the Brainreader (top) and SENSORIUM 2022 (bottom) datasets, demonstrating MEIcoder's superior fidelity and detail compared to baselines.
MEIcoder consistently outperforms all baselines across datasets and metrics, with the most pronounced gains in data- and neuron-scarce regimes. Notably, MEIcoder achieves high-fidelity reconstructions with as few as 1,000–2,500 neurons and less than 1,000 training samples, a strong claim substantiated by scaling experiments.
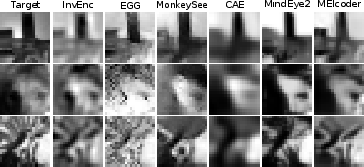

Figure 3: Synthetic Cat V1 reconstructions, highlighting MEIcoder's ability to generalize to large-scale, biologically realistic neural data.
Ablation and Scaling Analyses
Ablation studies reveal that MEIs are the primary driver of MEIcoder's performance; removing MEIs results in the largest degradation across all metrics, surpassing the impact of omitting neuron embeddings or substituting the SSIM loss.
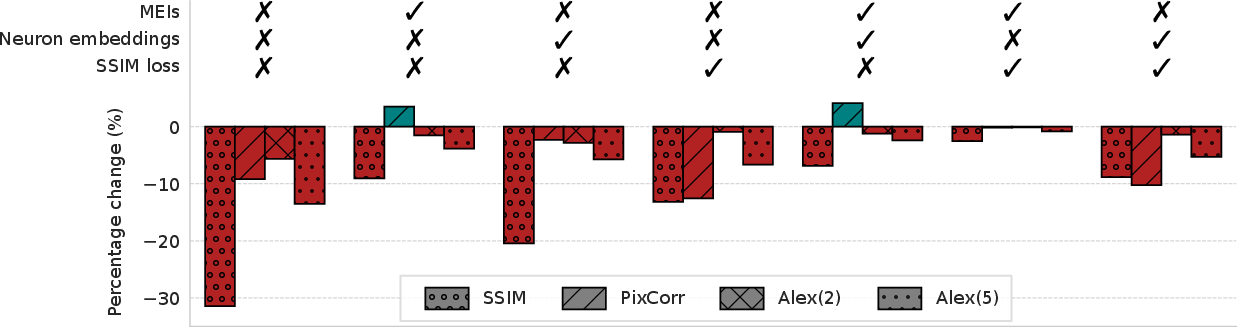
Figure 4: Ablation study quantifying the contribution of MEIs, neuron embeddings, and SSIM loss to overall performance.
Scaling experiments demonstrate that MEIcoder's performance saturates with increasing training data and neuron count, but qualitative improvements persist beyond metric plateaus, indicating limitations of current evaluation protocols.
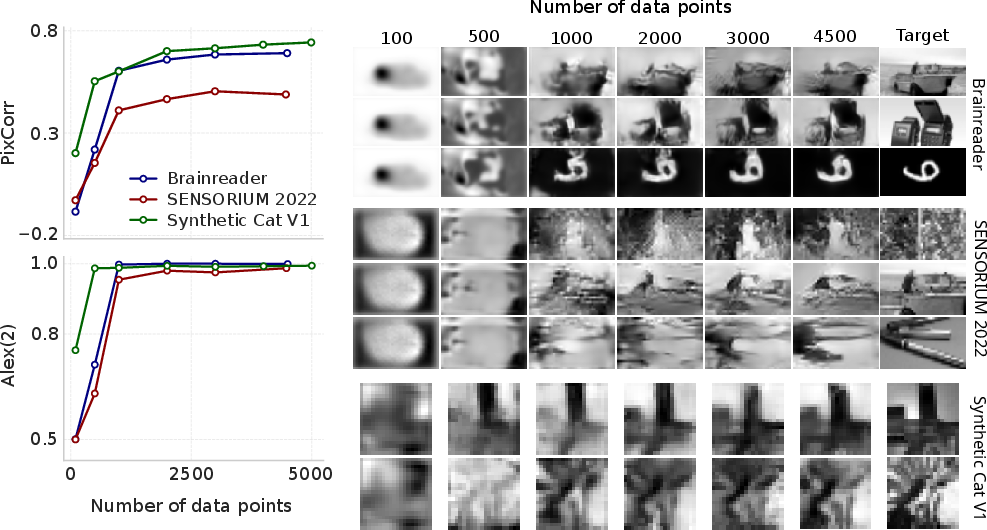

Figure 5: Relationship between MEIcoder's performance and the number of training data points, showing rapid gains with limited data.
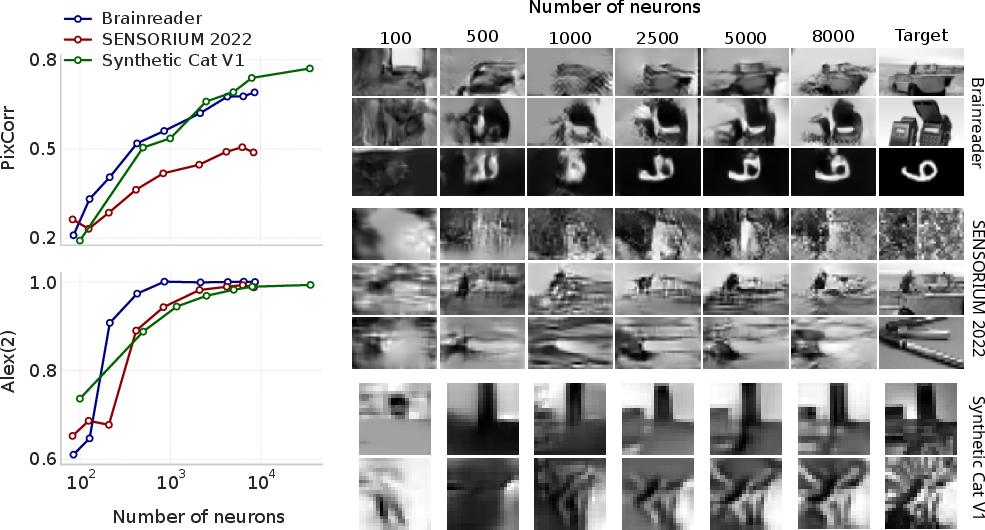
Figure 6: Relationship between MEIcoder's performance and the number of neurons, with high identification accuracy achieved at low neuron counts.
Interpretability and Concept-Based Analysis
MEIcoder's design enables interpretability via concept-based analysis. Non-negative matrix factorization (NMF) is applied to feature maps at each core layer, and sensitivity analysis quantifies the influence of individual neuron activations on visual concepts. The analysis reveals hierarchical refinement of features, specialization of neurons for local and global image properties, and identification of neurons with strong control over specific image regions, consistent with known V1 functional asymmetries.
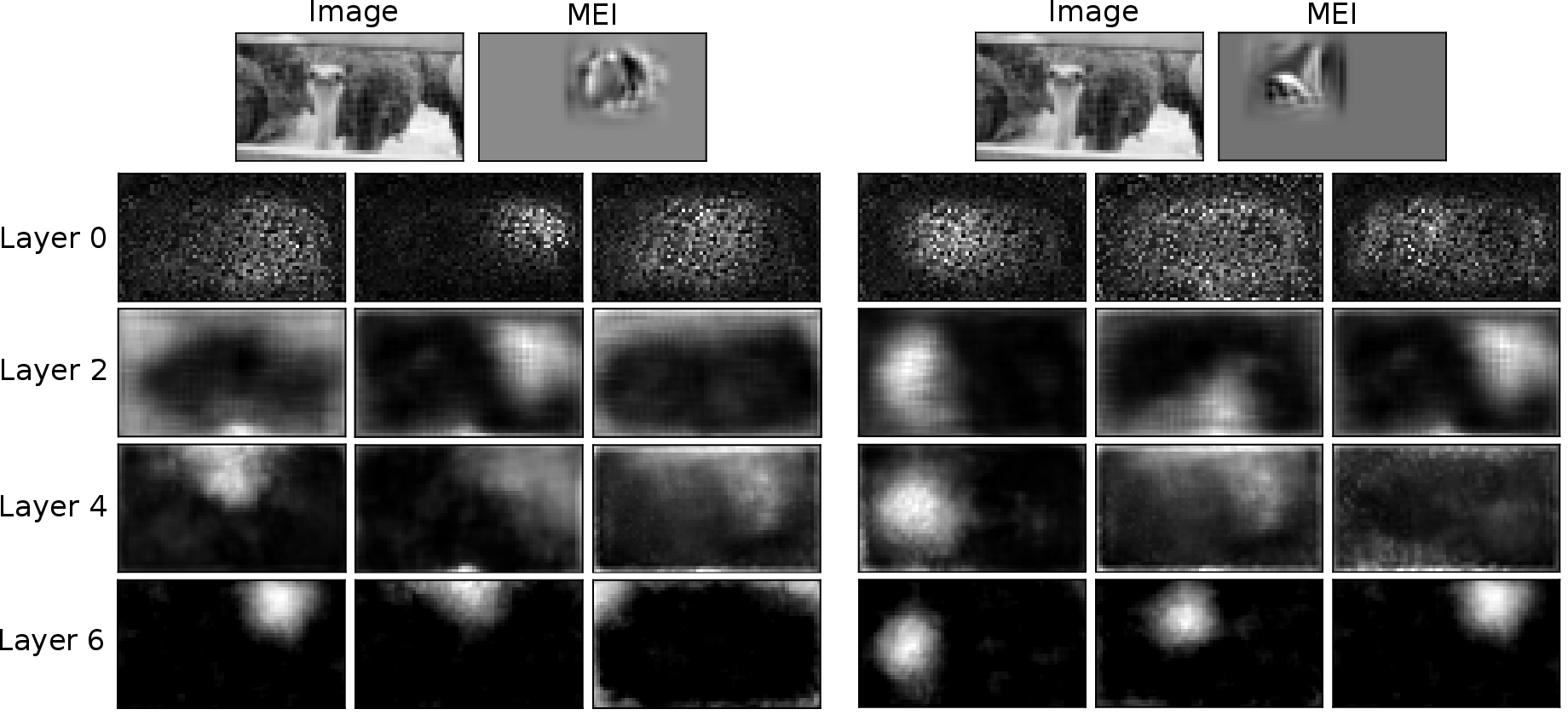
Figure 7: Input images, MEIs, and visual concepts with the highest activation gain for two example neurons, illustrating hierarchical feature specialization.
Robustness and Generalization
MEIcoder demonstrates robustness to noisy MEIs and generalizes to out-of-distribution artificial patterns, outperforming linear receptive field-based priors. Performance remains competitive even when decoding from highly nonlinear neurons, suggesting applicability to higher-order visual areas.
Practical and Theoretical Implications
MEIcoder's data- and neuron-efficiency has direct implications for brain-machine interfaces and neuroprosthetics, reducing the invasiveness and data requirements for reliable visual decoding. The framework's interpretability supports neuroscientific investigations into neural coding and population dynamics. The unified benchmark of 160,000+ samples facilitates standardized evaluation and future method development.
Future Directions
Potential avenues for future research include extending MEIcoder to higher-order visual areas, improving transfer learning across subjects, and integrating more sophisticated priors or concept extraction methods. Further work is warranted to refine evaluation metrics and explore clinical translation in human neuroengineering.
Conclusion
MEIcoder establishes a new state-of-the-art in decoding visual stimuli from V1 neural activity, leveraging biologically grounded priors and perceptually motivated objectives. Its superior performance in low-data and low-neuron regimes, robustness, and interpretability mark significant progress in neural decoding. The approach provides practical tools for neuroscience and neuroengineering, and the integrated benchmark supports ongoing research in this domain.






