- The paper introduces a novel Bayesian design that integrates patient pharmacokinetic data to model individual toxicity risks.
- It employs a two-stage process combining cohort-based dose escalation with personalized dosing based on predicted toxicity probabilities.
- Simulation studies show improved maximum tolerated dose selection and enhanced patient safety compared to conventional methods.
Precision Dose-Finding Design for Phase I Oncology Trials
The paper "Precision Dose-Finding Design for Phase I Oncology Trials by Integrating Pharmacology Data" (2509.05120) introduces a novel methodological framework for Phase I oncology trials that aims to refine dose-finding by incorporating pharmacokinetic (PK) data. This approach directly addresses inter-patient variability by using individual patient data to model toxicity risk, thereby advancing towards truly personalized medicine in clinical trial design.
Overview of the Precision Dose-Finding Design
The paper presents the Precision Dose-Finding (PDF) design, a Bayesian Phase I trial methodology that integrates patient-specific PK data into the dose-finding process. The PDF design consists of two stages: a training stage for model parameter updates using cohort-based dose escalation, followed by a test stage where doses are tailored to individual patients based on PK-predicted toxicity probabilities.
The framework assumes a one-compartment PK model characterized by individual parameters such as the volume of distribution (Vi) and the elimination rate constant (ki). The toxicity probability for each patient is calculated using these personalized PK metrics, distinguishing the PDF design from traditional population-level methods.
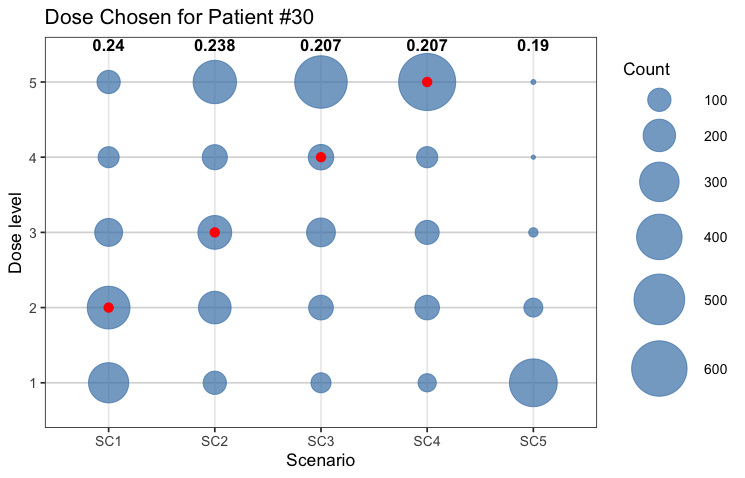
Figure 1: Dose assignment for the last patient treated in the precision dose-finding stage, illustrating personalized dose allocation with respect to individual toxicity profiles.
Methodological Details and Computational Approach
Bayesian Probability Model
The model utilizes the area under the concentration-time curve (AUC) as a covariate, which is derived from the patient-specific PK parameters. The toxicity probability pi for patient i is expressed as:
pi=logit−1(β0+β1×logVikidi),
where di is the administered dose, and β0, β1 are model parameters estimated using MCMC simulations. This individualized approach facilitates prediction of toxicity risk through posterior samples, maintaining robust inference throughout the trial.
Stage I: Cohort-Based Dose Escalation
The initial stage employs cohort-based dose escalation, where decisions are based on predictive toxicity probabilities derived from cumulative patient data. The approach adheres to traditional safety and escalation rules to balance dose exploration and patient safety effectively.
Stage II: Precision Dose-Finding
In the test stage, each new patient receives a dose tailored to his/her predicted toxicity profile, enhancing dosing precision. Selection of individual doses is informed by the posterior summaries of Vi and ki, ensuring personalized treatment optimization.
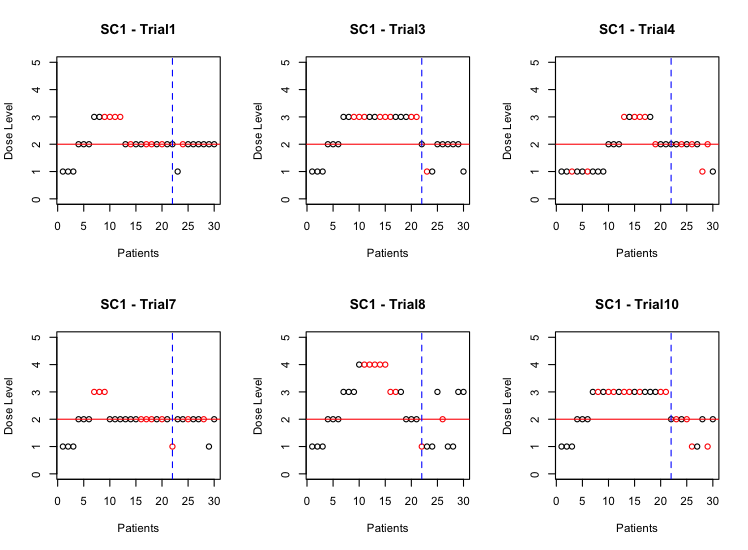
Figure 2: Dose assignments and DLT outcomes of six trials in scenario 1, showcasing dose escalation and precision dose-finding phases.
Simulation Study
Extensive simulation studies corroborate the PDF design's efficacy, demonstrating improved safety and dosing precision compared to the CRM. The simulations exhibit the strength of personalized dosing in reducing overdose risk and enhancing MTD identification accuracy, particularly in scenarios with high variability in patient responses.
Results
Results indicate that PDF efficiently allocates patients to appropriate dose levels, often achieving superior selection of MTDs with lower risk of toxicities. While the PDF approach aligns closely with CRM in identifying correct dose levels, it demonstrates a notable advantage in patient-specific safety profiles.
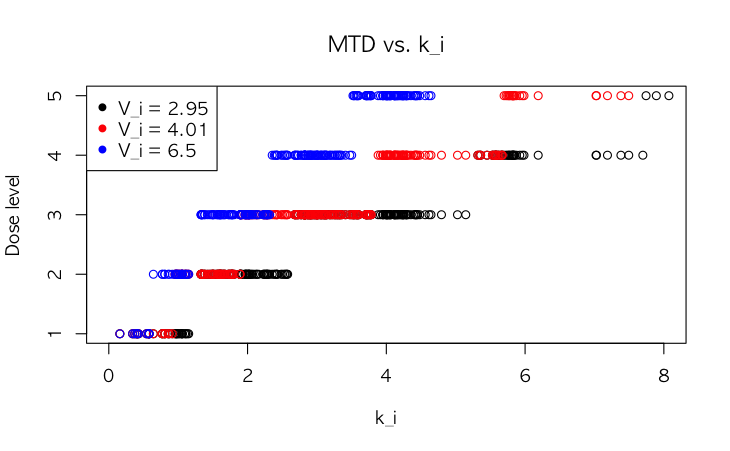
Figure 3: Examples of three volumes of distribution Vi with varying elimination rate constants ki and their corresponding predicted MTD values.
Discussion and Future Directions
The PDF design represents a significant step towards incorporating precision medicine principles into early-phase oncology trials. By leveraging individual PK data, this method provides a more accurate and personalized strategy for dose assignment, paving the way for innovations in clinical pharmacology and therapeutic customization.
Future research may focus on extending the PDF model to include more complex PK/PD relationships and dynamic dose adaptation strategies. Additionally, further investigation into the integration of machine learning approaches for real-time patient data analysis could enhance prediction accuracy and model generalizability.
Conclusion
The proposed PDF design effectively integrates individual PK data into the dose-finding process, offering a refined strategy for Phase I trials. With its Bayesian framework, the method supports personalized dosing while maintaining rigorous safety oversight, aligning early-phase studies with the ideals of precision oncology.


