Enhancing Cardiovascular Disease Prediction through Multi-Modal Self-Supervised Learning
Abstract: Accurate prediction of cardiovascular diseases remains imperative for early diagnosis and intervention, necessitating robust and precise predictive models. Recently, there has been a growing interest in multi-modal learning for uncovering novel insights not available through uni-modal datasets alone. By combining cardiac magnetic resonance images, electrocardiogram signals, and available medical information, our approach enables the capture of holistic status about individuals' cardiovascular health by leveraging shared information across modalities. Integrating information from multiple modalities and benefiting from self-supervised learning techniques, our model provides a comprehensive framework for enhancing cardiovascular disease prediction with limited annotated datasets. We employ a masked autoencoder to pre-train the electrocardiogram ECG encoder, enabling it to extract relevant features from raw electrocardiogram data, and an image encoder to extract relevant features from cardiac magnetic resonance images. Subsequently, we utilize a multi-modal contrastive learning objective to transfer knowledge from expensive and complex modality, cardiac magnetic resonance image, to cheap and simple modalities such as electrocardiograms and medical information. Finally, we fine-tuned the pre-trained encoders on specific predictive tasks, such as myocardial infarction. Our proposed method enhanced the image information by leveraging different available modalities and outperformed the supervised approach by 7.6% in balanced accuracy.
Paper Prompts
Sign up for free to create and run prompts on this paper using GPT-5.
Top Community Prompts
Explain it Like I'm 14
What this paper is about
This paper tries to predict heart problems—especially heart attacks—more accurately by teaching computers to learn from different kinds of medical data at the same time. The data includes:
- Heart images from MRI scans (CMR)
- Electrical signals of the heart (ECG)
- Basic medical information like age, lab tests, and lifestyle (tabular data)
The idea is to use smart learning methods that don’t need lots of hand-labeled examples, so the computer can learn from the large amount of unlabelled patient data available.
The main questions the researchers asked
- Can combining different types of medical data help predict heart disease better than using just one type?
- Can we “transfer” the detailed knowledge from expensive heart MRI images to cheaper and more common data like ECGs and basic medical info?
- Can self-supervised learning (where the computer teaches itself using clever tasks) improve predictions when labeled data is limited?
How they did it (in simple steps)
Think of this like training a team of “specialist readers,” each learning to understand different kinds of health data, and then teaching them to work together.
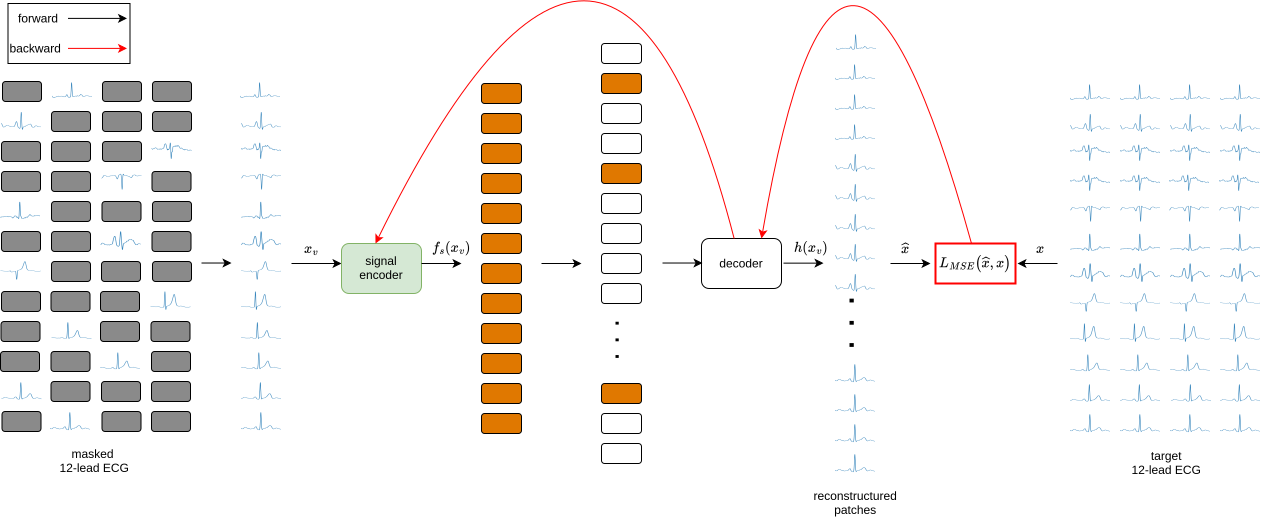
- Pre-train the ECG reader by solving puzzles:
- They used a technique called a “masked autoencoder.” Imagine covering parts of a long ECG signal and asking the model to guess the missing pieces—like completing a jigsaw puzzle. This helps the model understand ECG patterns without needing labels.
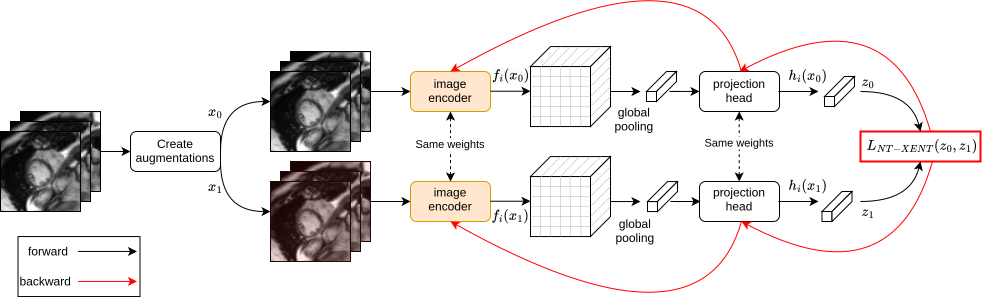
- Pre-train the image reader with spot-the-difference:
- For MRI images, they used “contrastive learning” (SimCLR). The computer looks at two slightly changed versions of the same image and learns that they are still the same heart. This teaches it what matters in the image.
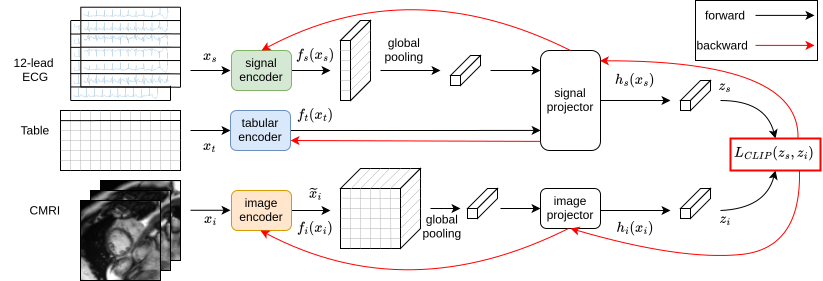
- Align the readers so they speak the same “language”:
- They used a method similar to how you learn to match pictures with captions (CLIP loss). The goal: make the ECG-plus-tabular data embedding line up with the MRI image embedding for the same person. In plain terms, they teach the cheaper data to “think like” the richer MRI data.
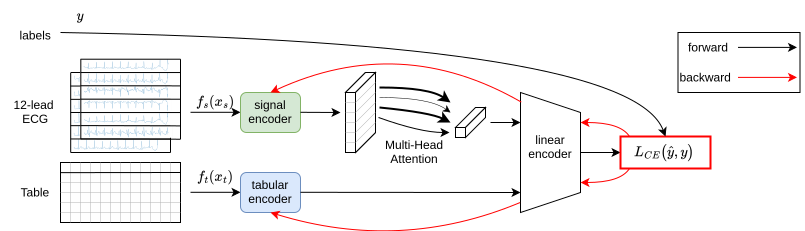
- Fine-tune for heart attack prediction:
- After the readers are trained, they add labels (who had a heart attack vs. who didn’t) and fine-tune the ECG-plus-tabular model to make the final prediction.
This approach lets the cheap, widely available data (ECG and tabular info) benefit from the detailed knowledge hidden in MRI scans—without needing MRI scans at prediction time.
What they found and why it matters
- Their method improved balanced accuracy by 7.6% compared to a standard supervised model using only ECG and tabular data.
- Balanced accuracy is important when the dataset has many more healthy people than sick people; it checks how well the model does on both groups fairly.
- The model learned better by using self-supervised learning (learning from puzzles and matching tasks) before using labels.
- Even when there aren’t many labeled examples, this training style helps avoid “overfitting” (where the model memorizes training data but doesn’t generalize).
In short: Combining different types of data and using smart, label-free training helped the computer make more reliable predictions about heart disease.
Why this research matters
- Better early detection: More accurate predictions can help doctors spot heart problems sooner.
- Uses common data: ECGs and basic medical info are cheap and easy to collect. If these can be improved using MRI knowledge, more patients can benefit.
- Works with limited labels: Hospitals often don’t have lots of well-labeled data. This approach still performs well.
Limitations and future impact
- The data comes from the UK Biobank, which may not represent all types of people (for example, different countries or age groups). The model needs testing on more diverse populations.
- Future work could try other learning methods, include more data types (like text notes from doctors), and explore how much knowledge can be transferred from MRI to other cheaper tests.
- If improved and widely tested, this approach could help personalized healthcare by giving doctors a fuller picture of a patient’s heart health using the data that’s easiest to get.
Collections
Sign up for free to add this paper to one or more collections.