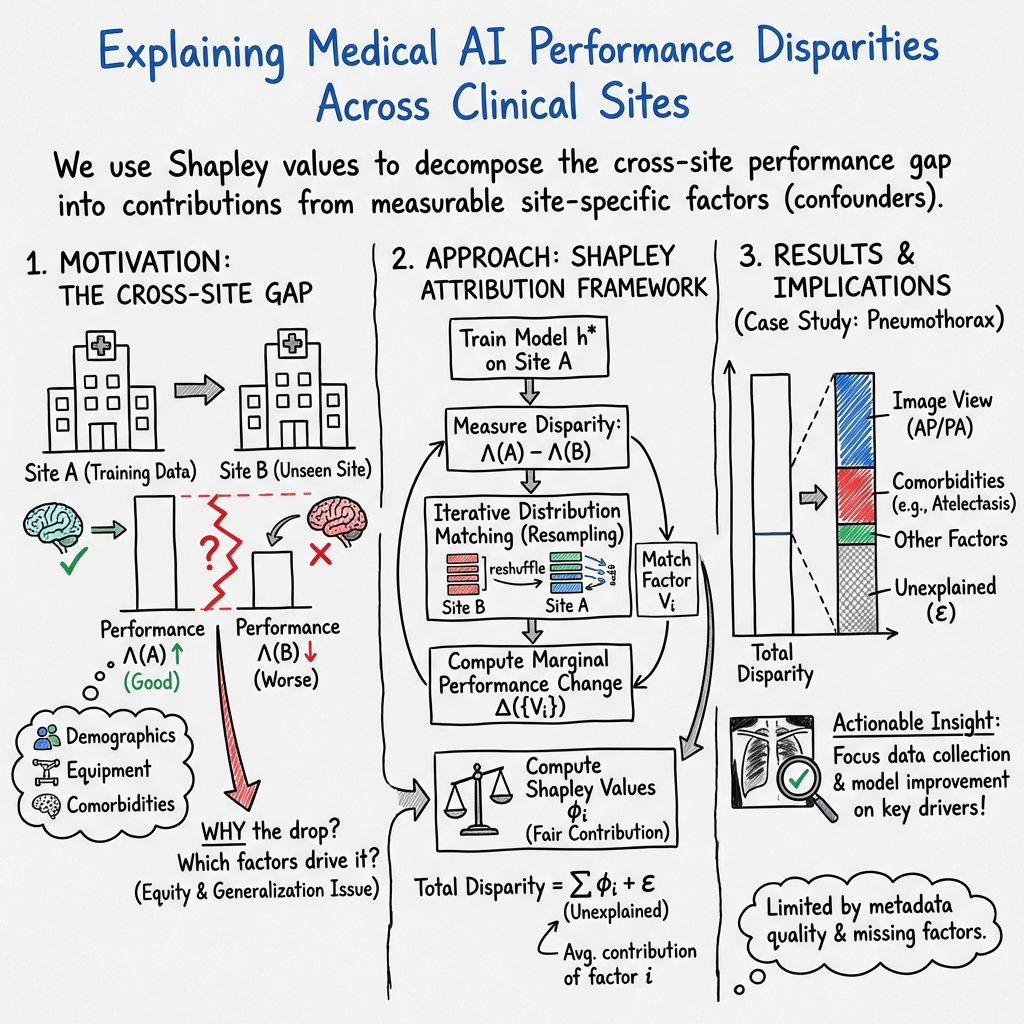
Explaining medical AI performance disparities across sites with confounder Shapley value analysis
Abstract: Medical AI algorithms can often experience degraded performance when evaluated on previously unseen sites. Addressing cross-site performance disparities is key to ensuring that AI is equitable and effective when deployed on diverse patient populations. Multi-site evaluations are key to diagnosing such disparities as they can test algorithms across a broader range of potential biases such as patient demographics, equipment types, and technical parameters. However, such tests do not explain why the model performs worse. Our framework provides a method for quantifying the marginal and cumulative effect of each type of bias on the overall performance difference when a model is evaluated on external data. We demonstrate its usefulness in a case study of a deep learning model trained to detect the presence of pneumothorax, where our framework can help explain up to 60% of the discrepancy in performance across different sites with known biases like disease comorbidities and imaging parameters.
Paper Prompts
Sign up for free to create and run prompts on this paper using GPT-5.
Top Community Prompts
Explain it Like I'm 14
Explaining the paper: “Explaining medical AI performance disparities across sites with confounder Shapley value analysis”
Overview: What is this paper about?
This paper looks at why medical AI systems often work well at one hospital but worse at another. The authors create a way to “break down” the performance drop and explain how much each difference between hospitals—like patient ages, scan types, or other diseases—contributes to that drop. They test their idea on AI that detects pneumothorax (a collapsed lung) in chest X-rays.
What were the researchers trying to find out?
The main questions are:
- Why do AI models for healthcare perform differently at different hospitals?
- Which specific differences between hospitals cause the performance drop?
- How much does each factor (like image view, patient age, or other illnesses) contribute to the difference?
- Can we use this information to make AI fairer and more reliable across different places?
How did they study it? (Simple explanation of the method)
Think of an AI model like a student who learned from one school (Hospital A) and then takes a test at another school (Hospital B). The student might do worse because the tests are different in various ways. The researchers wanted to know which differences matter most.
They used a fairness idea called “Shapley values” from game theory. Here’s an everyday analogy:
- Imagine a team wins a game. You want to fairly split the credit among players. You look at how much the team scores with and without each player in different lineups. A Shapley value is a fair way to assign each player their share of the credit.
- In this paper, the “players” are hospital differences (called “site factors”), like:
- Patient age
- Sex
- X-ray view (front, side, etc.)
- Image size/shape
- Comorbidities (other illnesses seen on the X-ray, like enlarged heart or atelectasis)
- The “score” is how well the model performs (measured by AUC, a number from 0 to 1 where higher is better).
Here’s what they did, step by step:
- Train an AI model on chest X-rays from one hospital.
- Test it on the same hospital’s data (best-case).
- Test it on a different hospital’s data (often worse).
- Measure the drop in performance between “same hospital” and “new hospital.”
- Then, for one factor at a time, make the new hospital’s data “look like” the original hospital on that factor. For example, if Hospital A uses more front-view X-rays and Hospital B uses more side-view X-rays, they re-balance Hospital B’s test set to match A’s proportion of image views.
- Test the model again and see how much the performance improves. That improvement is the amount of the drop explained by that factor.
- Because multiple factors can interact, they repeat this process in many orders and combinations, then use Shapley values to fairly split the total “blame” among all factors.
- They repeat the sampling many times to make the estimate stable.
In short: they “match” the data factor-by-factor, observe how the model’s score changes, and use a fair-sharing method (Shapley values) to assign how much each factor contributes to the performance gap.
What did they find, and why is it important?
They tested their method on three large US chest X-ray datasets from different hospitals: NIH (Bethesda, MD), SHC/Stanford (Palo Alto, CA), and BIDMC (Boston, MA). They trained a deep learning model (DenseNet-121) to detect pneumothorax and evaluated it across hospitals.
Key findings:
- Cross-hospital performance drops are real and can be significant.
- Their framework could explain, on average, about 27% of the performance difference. In some cases, it explained much more—up to about 60%.
- In two comparisons (SHC on BID and BID on SHC), their method explained very little. The likely reason: important metadata (like race, socioeconomic status, detailed scanner settings) wasn’t available in both datasets, so the method couldn’t analyze those factors.
- The two biggest contributors to performance gaps were:
- X-ray image view (AP/PA/lateral). Different views show the chest in different ways, and many AI models don’t train separately by view.
- Comorbidities (other conditions in the image). Some illnesses can confuse the model. For example:
- Atelectasis (part of the lung collapses) and
- Cardiomegaly (enlarged heart)
- were particularly important. These can make images look different and lead to false positives or confusion.
Why this matters:
- Instead of just saying “the model works worse elsewhere,” this method says “here’s how much worse, and here’s which differences are causing it.” That’s actionable.
- It helps developers focus on the most important fixes—for example, training separate models by image view or collecting more images with certain comorbidities.
- It can guide hospitals and researchers on what metadata to collect and share, making future models more reliable and fair.
What’s the big picture impact?
- For clinicians and users: You can understand when and why an AI might make more mistakes at your hospital.
- For developers: You can target data collection and model design to the factors that cause the biggest problems, instead of guessing.
- For dataset creators and regulators: You can encourage or require collection of key metadata that makes fair evaluation possible and highlight when multi-site testing is truly diverse.
Limitations to keep in mind:
- Missing or sparse metadata limits what can be explained. In some datasets, important details like race or detailed imaging parameters weren’t available.
- Labels can be noisy (for example, different hospitals use different methods to label images), which can affect results.
- The method matches factors one by one (not all at once), which is practical but may miss complex interactions.
- More studies are needed across other diseases and image types.
Takeaway
Medical AI often performs differently across hospitals because hospitals differ in patients, imaging styles, and other conditions. This paper offers a fair and practical way to measure how much each difference contributes to the performance gap. The results show that factors like X-ray view and comorbidities are major drivers. By identifying and quantifying these factors, the approach helps make medical AI safer, fairer, and more reliable in the real world.
Collections
Sign up for free to add this paper to one or more collections.