- The paper presents a deep learning method using U-net architecture for fully automated muscle ultrasound analysis.
- It demonstrates high correlation with manual and semi-automated systems in measuring fascicle length and pennation angle.
- The study reduces processing time and enhances reproducibility, paving the way for real-time diagnostics in clinical settings.
Fully Automated Muscle Architecture Analysis Using Deep Learning
Introduction
The study presents a deep learning approach for the analysis of muscle architecture from B-mode ultrasound images. Utilizing a U-net-based neural network architecture, the system identifies superficial and deep aponeuroses as well as muscle fascicle fragments, streamlining what has traditionally been a manual or semi-automated process. This paper offers a comprehensive methodology, training data, and implementation to enable broad application and adaptation by other researchers in the field of musculoskeletal analysis.
Methods
The implementation centers around utilizing the U-net model, renowned for its efficacy in medical image segmentation tasks. Training was performed on a labeled dataset consisting of ultrasound images acquired from diverse muscle groups and populations, using supervised learning to accurately detect aponeuroses and muscle fascicles.
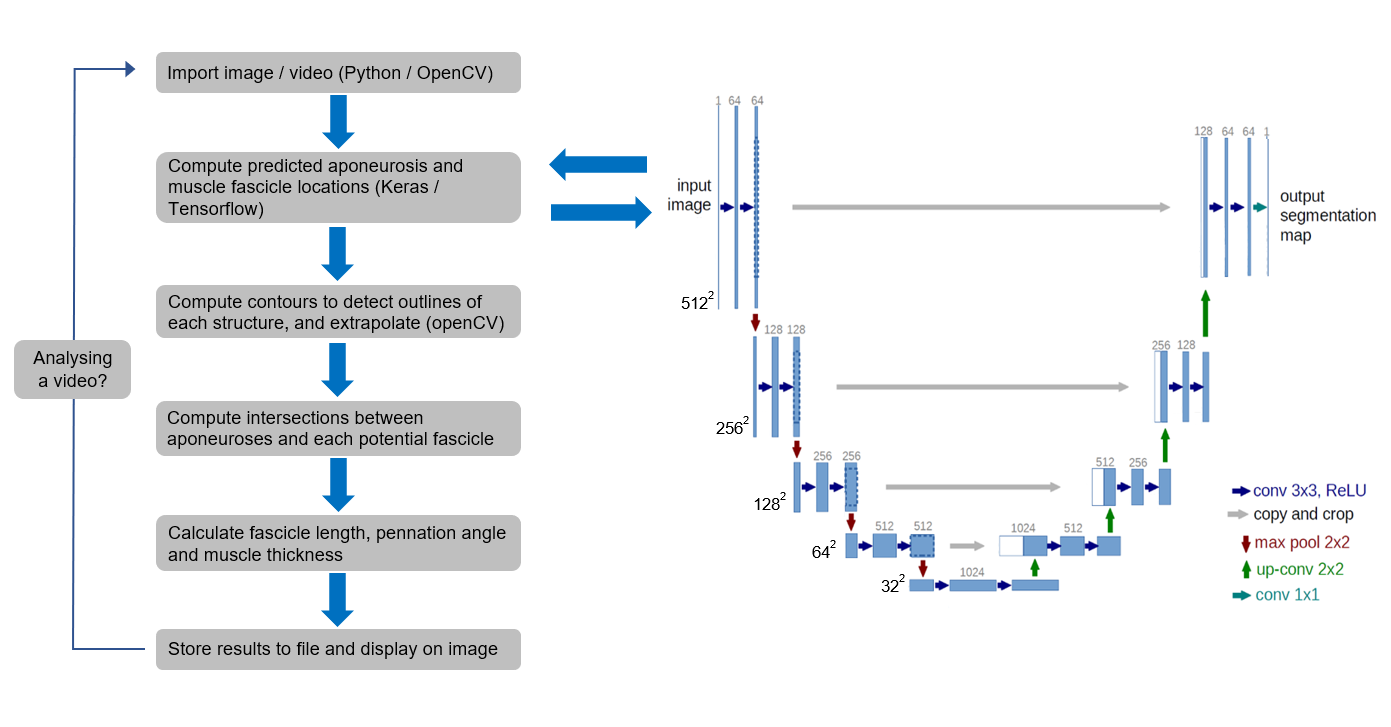
Figure 1: Schematic of the workflow, including details of the U-net model architecture.
Training was conducted using a GPU to accelerate the process, with models trained until overfitting was evident. The architecture's augmentation abilities are crucial to compensating for the limited size of biomedical datasets, allowing for robust model training despite relatively low image acquisition volumes.
Results
The system demonstrated proficient performance, producing results akin to both manual analyses and existing semi-automated methods in parameters such as fascicle length and pennation angle, with high concordance correlations and minimal mean differences.
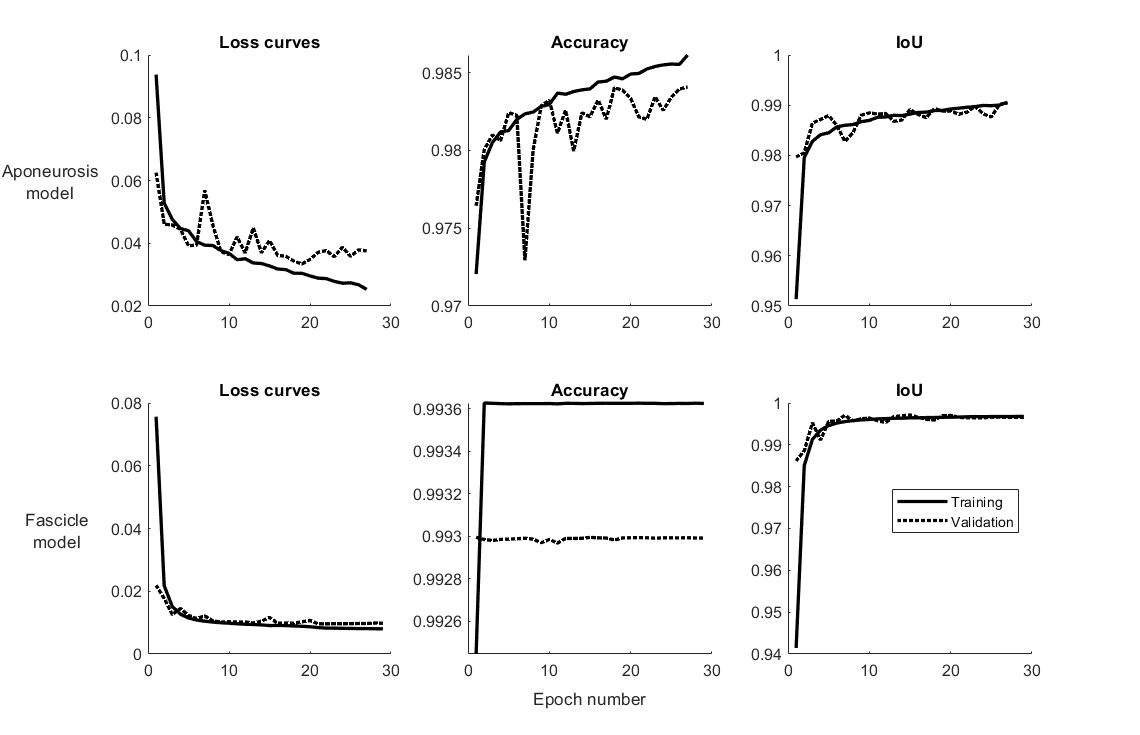
Figure 2: Loss function and intersection-over-union (IoU) training curves for the two trained models.
Moreover, the approach achieved a reduction in processing time, achieving inference on single images in approximately 0.7 seconds when utilizing a GPU. The system effectively identified multiple fascicles per image, offering a more comprehensive analysis than single-fascicle detection methodologies.
Comparative Analysis
Comparative analysis with traditional manual methods and the Ultratrack system, a prominent semi-automated approach, revealed that the deep learning model provided comparable insights into muscle architecture across dynamic tasks.
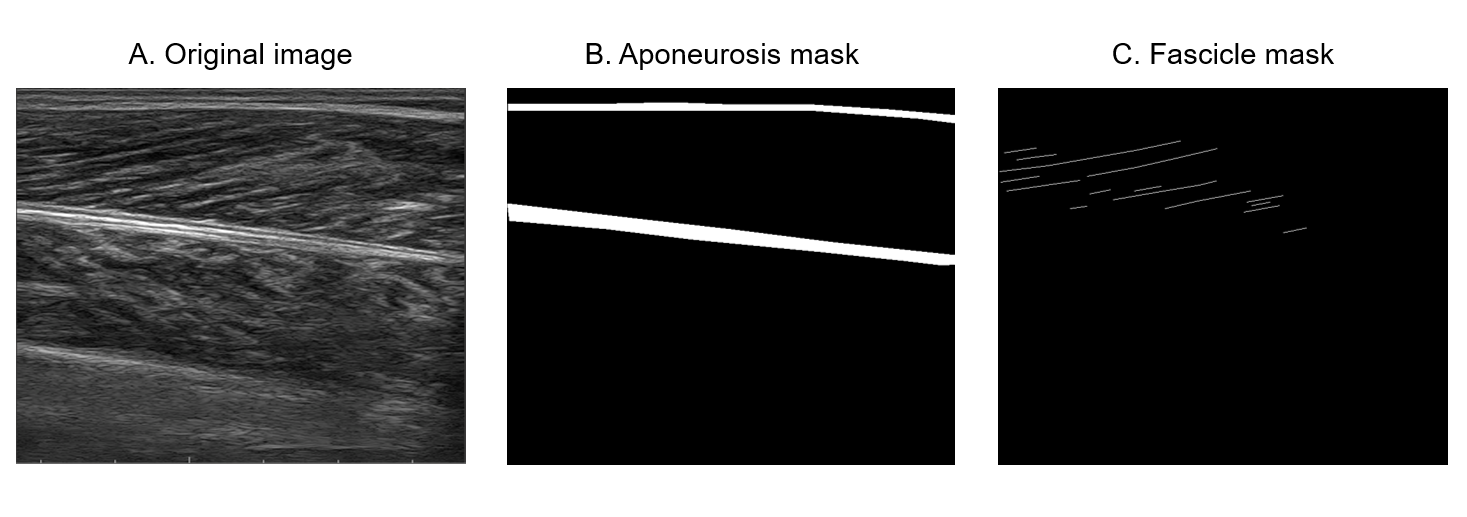

Figure 3: Examples of ultrasound images with annotations from the trained neural networks, illustrating fascicle length and pennation angle results.
Bland-Altman plots showcased minimal biases and close agreement between the deep learning predictions and manual measurements.
Discussion
The deep learning approach presents a scalable solution for muscle architecture analysis, eliminating the need for user intervention during the detection process. This advancement addresses prior limitations in automated systems, such as the requirement for manual feature delineation and susceptibility to variances in imaging parameters.
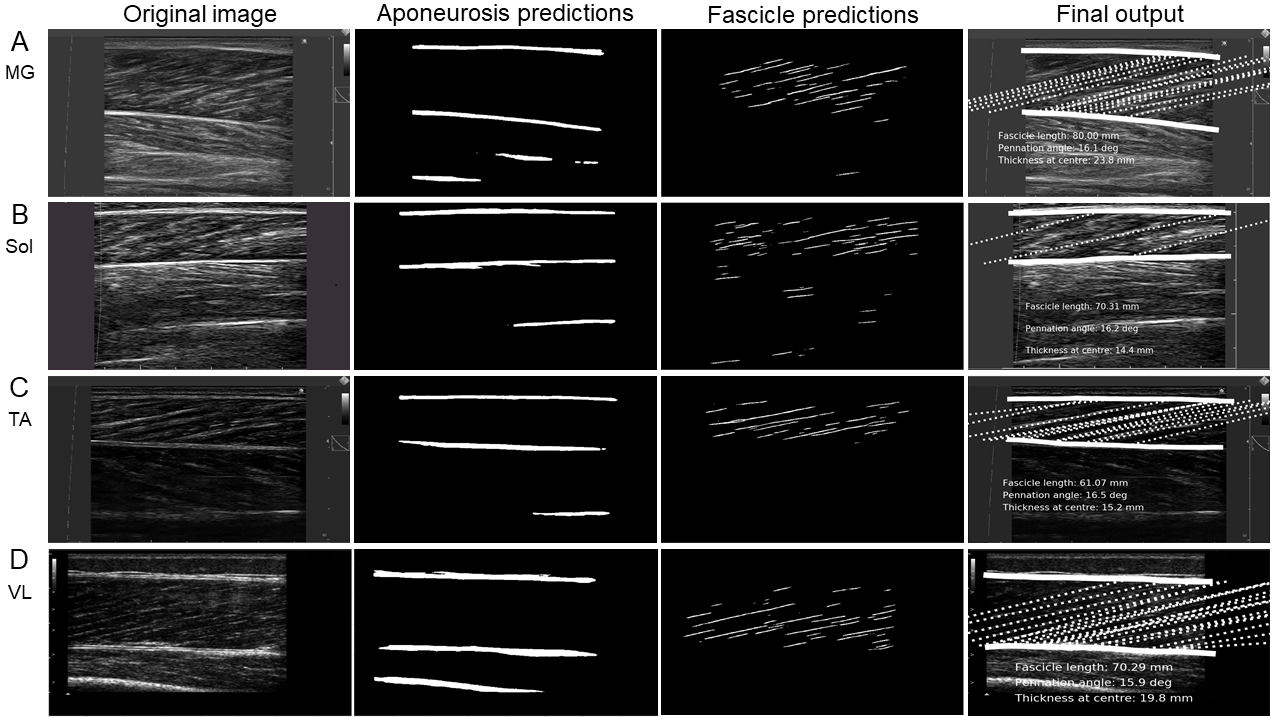
Figure 4: Bland-Altman plots showing results from the deep learning approach against manual analyses and the SMA method across various metrics.
Future research could focus on enhancing the temporal consistency of detected fascicle trajectories through recurrent network integration, potentially mitigating the observed frame-to-frame variability intrinsic to the current approach's independent frame analysis methodology.
Conclusion
The study successfully demonstrates a fully automated, open-source approach for ultrasound image analysis, capable of accurately determining critical muscle architecture parameters. With provision for both the necessary software and dataset access, this method stands as a valuable resource for future research developments, offering the potential for real-time analytics with adequate computational support.
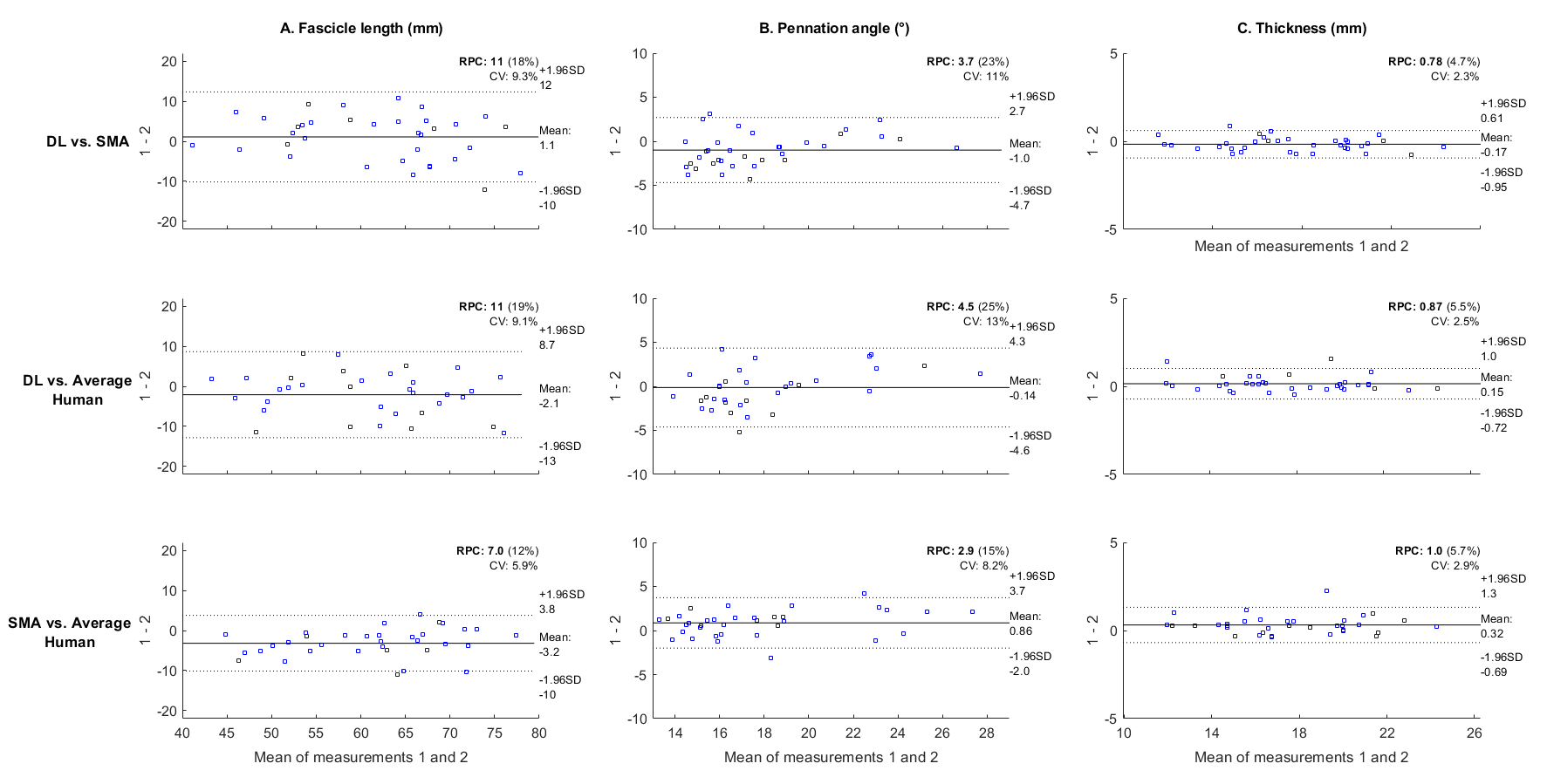
Figure 5: A comparison of the deep learning approach and UltraTrack for calculating muscle fascicle lengths and pennation angles across various dynamic tasks.
This methodology paves the way for reliable, automated muscle architecture assessments, essential for both clinical diagnostics and biomechanical research.






