- The paper introduces DDBMs that map between healthy and pathological brain MRIs, enabling accurate healthy counterfactual generation.
- It leverages a modified stochastic differential equation with Doob's transform and a 3D UNet architecture to condition the diffusion process.
- DDBMs achieve superior segmentation Dice scores and anomaly detection metrics, facilitating improved lesion localization in clinical imaging.
Denoising Diffusion Bridge Models for Healthy Counterfactual Generation in Medical Imaging
Introduction
The paper introduces a novel application of Denoising Diffusion Bridge Models (DDBMs) for generating healthy counterfactuals from pathological medical images, specifically brain MRI. The motivation is to enable the use of standard image analysis tools, which are typically designed for healthy scans, on pathological data by reconstructing plausible healthy versions of diseased images. This approach is particularly relevant for tasks such as anomaly detection and lesion localization, where the difference between the original and counterfactual images can be leveraged to identify abnormal regions.
Methodological Framework
Denoising Diffusion Probabilistic Models (DDPMs)
DDPMs model the data distribution by gradually adding noise to an image and then learning to reverse this process. In the context of healthy counterfactual generation, DDPMs are trained on healthy images, with the assumption that the denoising process will reconstruct healthy tissue when applied to partially noised pathological images. However, DDPMs are limited by their reliance on a Gaussian prior and the challenge of balancing anomaly inpainting with the preservation of subject-specific anatomical features.
Denoising Diffusion Bridge Models (DDBMs)
DDBMs extend DDPMs by conditioning the diffusion process on both the initial (healthy) and terminal (pathological) states, enabling direct mapping between healthy and diseased distributions. The forward process is defined by a modified SDE using Doob's h-transform, resulting in a tractable Gaussian bridge kernel. The reverse process leverages a bridge score function, approximated via a neural network trained with a denoising bridge score matching objective. This conditioning allows the model to selectively remove pathology while retaining individual anatomical characteristics.
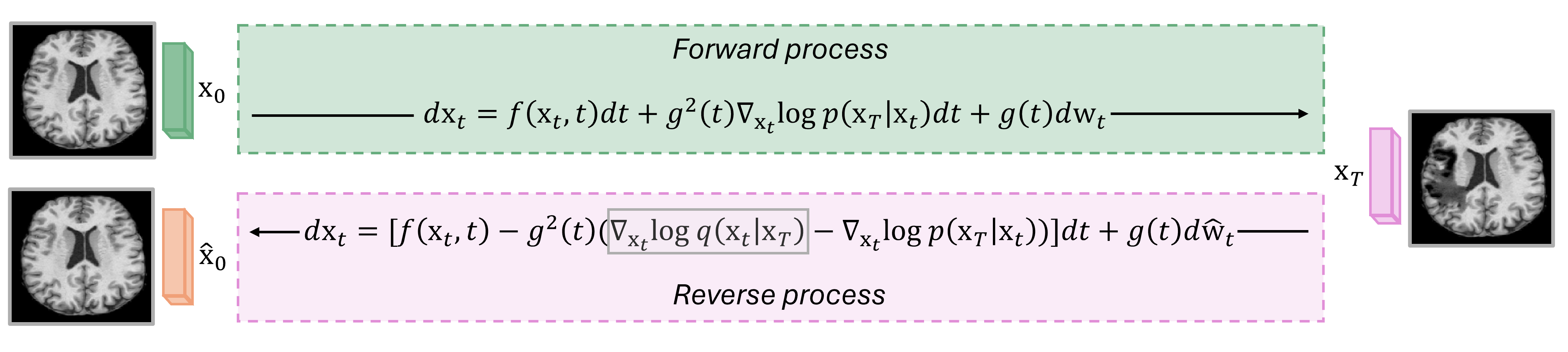
Figure 1: The DDBM framework maps between synthetic pseudo-pathology and real healthy volumes, with the denoising bridge score matching objective guiding the network to learn the bridge score function.
Synthetic Pathology Generation
Training DDBMs requires paired healthy and pathological images. The paper employs the UNA framework, which simulates pathology as an advection-diffusion process, generating realistic synthetic anomalies based on real lesion segmentations. These healthy–pseudo-pathology pairs are used to train the DDBM, with the healthy image as the start point and the synthetic pathology as the endpoint.
Implementation Details
The score network sθ is implemented as a 3D UNet, following the architecture of Song et al. (2021), and trained using the VP noise schedule. Sampling is performed using the DBIM procedure for efficient inference. Training is conducted on large-scale MRI datasets, with extensive preprocessing including skull-stripping, bias-field correction, normalization, and affine registration to MNI152 space.
Experimental Evaluation
Segmentation Consistency
The primary evaluation metric is the Dice score between segmentations of ground-truth healthy images and their generated counterfactuals, using SynthSeg. Across five datasets (ADNI, HCP, ADHD200, AIBL, OASIS3), DDBM achieves the highest Dice scores and average ranks for most brain regions, outperforming Latent Diffusion Models (LDM), conditional LDMs (cLDM), and the UNA baseline. This demonstrates superior preservation of anatomical structure in the generated counterfactuals.
Anomaly Detection
For real pathology, the model is evaluated on the ATLAS dataset for stroke lesion localization. Anomaly maps are constructed by comparing the original pathological scan with its healthy counterfactual. DDBM achieves the highest Dice, average precision, and lowest false positive rate, surpassing both unsupervised and supervised baselines.
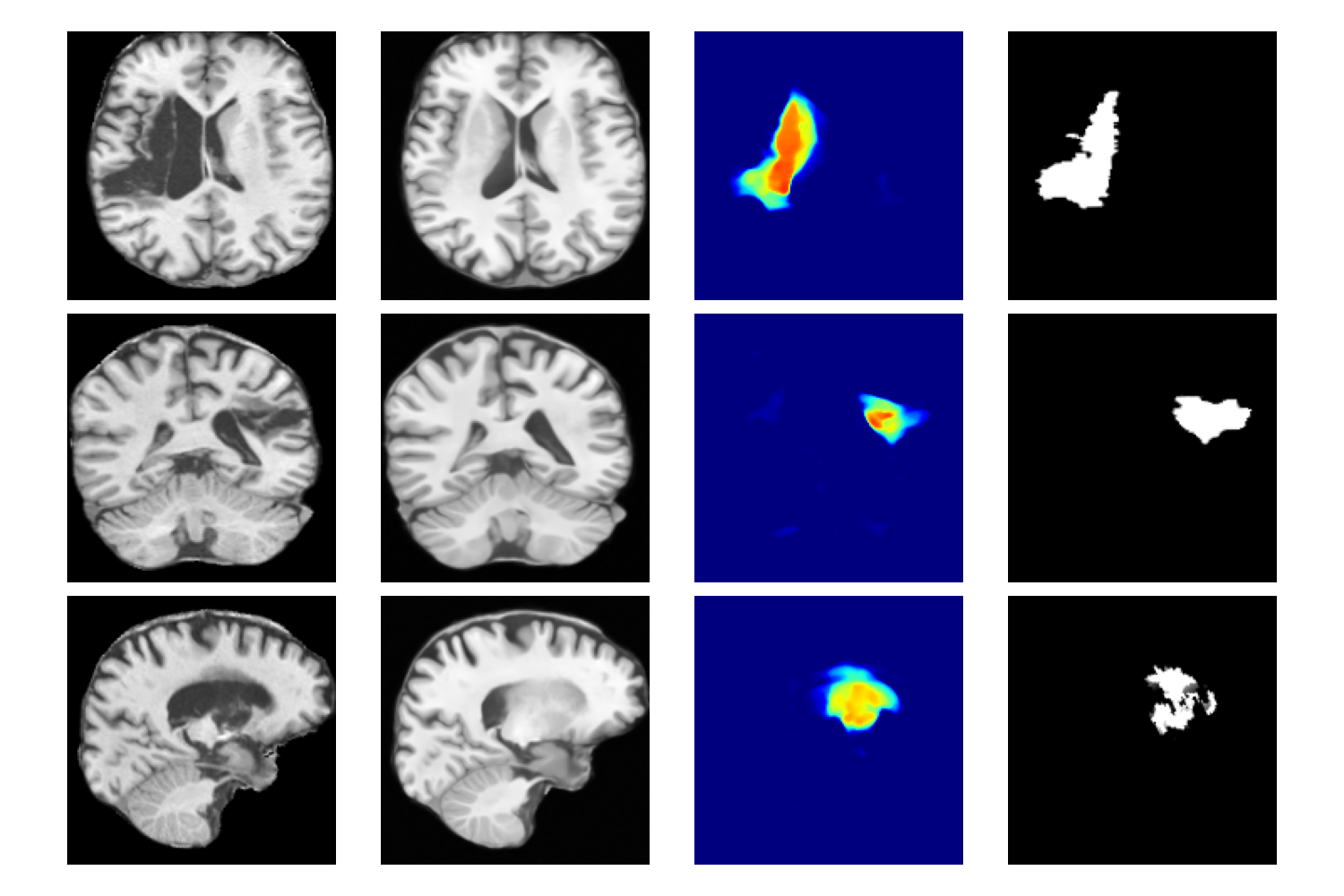
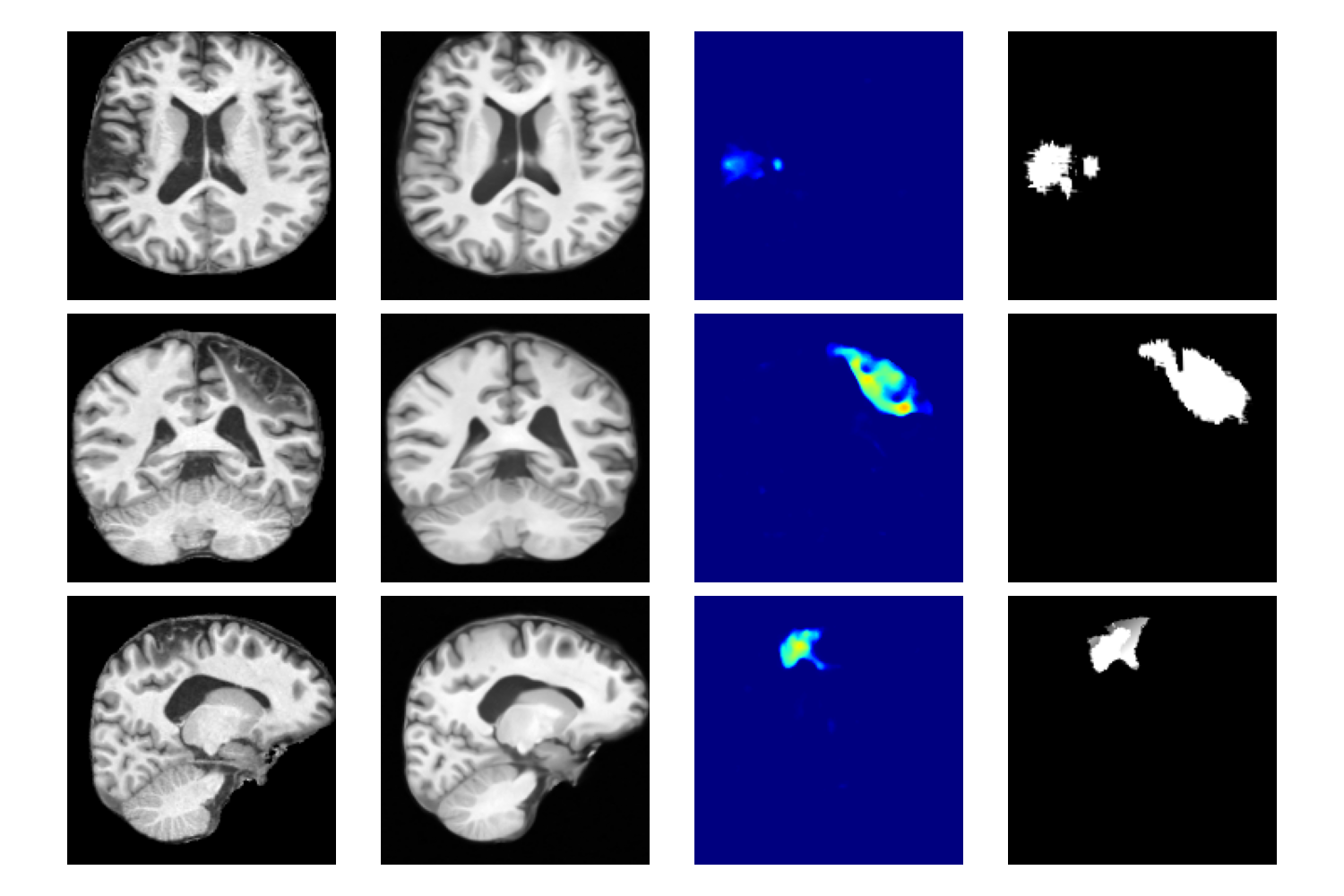
Figure 2: Qualitative example of anomaly detection on ATLAS test subjects, showing the original scan, DDBM-generated healthy counterfactual, anomaly map, and ground-truth segmentation.
Results and Analysis
- Segmentation Dice Scores: DDBM consistently ranks first across all datasets and regions, indicating robust anatomical fidelity in counterfactual generation.
- Anomaly Detection Metrics: DDBM achieves a Dice of 0.4774, AP of 0.4482, and FPR of 0.0026, outperforming UNA and other baselines.
- Qualitative Results: Anomaly maps generated by DDBM exhibit high contrast in abnormal regions and low contrast in healthy tissue, effectively localizing lesions while preserving surrounding anatomy.
Theoretical and Practical Implications
The DDBM framework provides a principled approach for distribution translation in medical imaging, leveraging structural correspondence between healthy and pathological scans. Conditioning on both endpoints enables selective inpainting of pathology without sacrificing individual anatomical features, addressing a key limitation of prior DDPM-based methods. The use of synthetic pathology generation further enhances training efficiency and realism.
Practically, DDBMs facilitate the application of normative analysis tools to pathological data, improve anomaly detection, and enable more accurate lesion localization. The approach is scalable to large datasets and compatible with existing segmentation pipelines.
Future Directions
Potential avenues for future research include:
- Incorporating anatomical priors or guidance to further enhance realism and structural fidelity.
- Extending the framework to other imaging modalities and disease types.
- Exploring integration with transformer-based architectures for improved scalability and expressivity.
- Investigating the use of DDBMs for longitudinal analysis and disease progression modeling.
Conclusion
The application of DDBMs to healthy counterfactual generation represents a significant methodological advance in medical image analysis. By conditioning on both healthy and pathological endpoints, DDBMs achieve superior performance in both tissue segmentation and anomaly detection, bridging the gap between normative and pathological anatomy. This framework opens new possibilities for precise, individualized medical image reconstruction and analysis.


