- The paper introduces CoNet, a U-Net adaptation that segments retinal images accurately with only 55 training images.
- It employs an ASPP block for multi-scale context integration while reducing convolutional depth to suit small datasets.
- It achieved an 88% mean Dice Score on the Duke DME dataset, outperforming state-of-the-art models in fluid segmentation.
Retinal Image Segmentation with Small Datasets
Introduction
Retinal imaging, particularly through Optical Coherence Tomography (OCT), plays a critical role in diagnosing and monitoring various eye diseases like Diabetic Macular Edema (DME), Age-related Macular Degeneration (AMD), and glaucoma, which can lead to irreversible vision loss. Despite advancements in deep learning (DL) techniques for automating these analyses, they typically require extensive datasets to train models effectively. The focus of this study is presenting a robust methodology to perform retinal image segmentation using a minimal amount of training data—only 55 images—without data augmentation.
Background
Retinal OCT is a high-resolution, non-invasive imaging technique providing detailed cross-sections (B-scans) of the retina, facilitating the analysis of structural changes indicative of diseases. Historically, segmentation methods in this domain have evolved from traditional techniques, like graph-based methods and active contours, to more sophisticated approaches involving CNNs and U-Net architectures. Such DL models have shown promise, particularly in segmenting complex anatomical structures and fluid accumulations, which are critical in disease progression analysis.
Methodology
The proposed method, named CoNet, is an adaptation of the U-Net architecture designed specifically for small datasets. It comprises several key components: an encoding path, bottleneck structure, Atrous Spatial Pyramid Pooling (ASPP) block, and a decoding path. Notably, the CoNet architecture modifies conventional practices by incorporating multi-scale context information through the ASPP block, enhancing segmentation accuracy even with limited data.
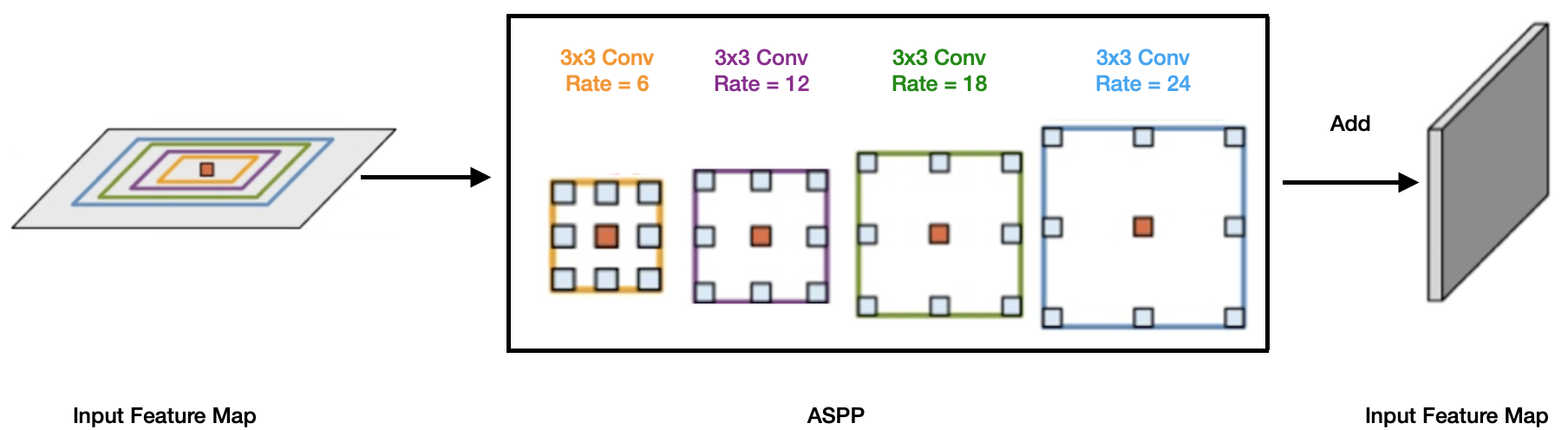
Figure 1: The ASPP block in the proposed model captures multi-scale information by applying multiple parallel filters with different frequencies.
CoNet utilizes a shallower network architecture, reducing from 5 to 3 convolutional blocks, optimizing for the small dataset size while maintaining the complexity needed to capture nuanced retinal features. The encoding path emphasizes spatial localization through convolutional layers with specific kernel sizes tailored to the B-scan's geometry, whereas the decoding path focuses on precision in pixel-level classification.
Experiments and Results
The study employed the Duke DME dataset, which contains 110 B-scans from patients with severe DME, annotated by experts to ensure reliable ground truth segmentation. Training was conducted using 55 B-scans, adhering to a systematic K-fold cross-validation strategy to ensure robust evaluation.

Figure 2: Annotation and labelling of the 10 segments (7 retinal layers, 2 backgrounds, and 1 fluid) in the Duke DME dataset.
CoNet demonstrated superior performance metrics. The model achieved a mean Dice Score of 88%, surpassing both human expert annotations and the state-of-the-art ReLayNet, particularly excelling in the segmentation of challenging fluid regions, which often exhibit significant variability.
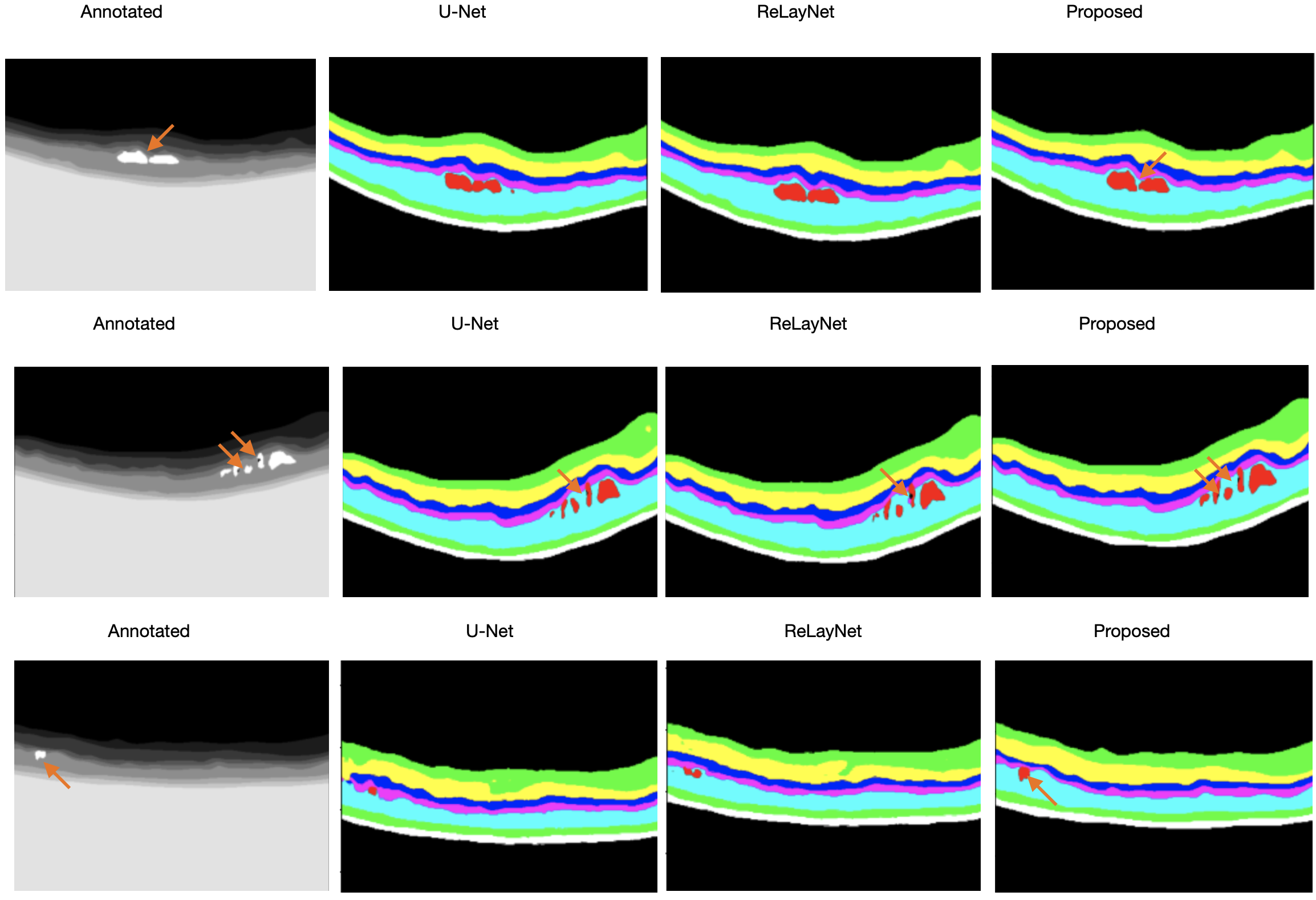

Figure 3: Examples to illustrate the visualization output of U-Net, ReLayNet, and the proposed CoNet, in order of the inputs, annotations, and outputs with orange arrows to demonstrate fine details picked up by the models.
Conclusion
The CoNet model exhibits significant potential in automated retinal image analysis, especially for applications constrained by limited data availability. Its architecture, featuring an ASPP block and a reduced convolutional depth, adapts well to the challenges of small datasets without compromising accuracy. This model sets a precedent for future exploration into similarly constrained medical imaging fields. Further advancements may involve extending CoNet to three-dimensional networks or applying it to diverse retinal pathologies to validate its applicability and resilience across varied diagnostic challenges.




